Abstract
Background. Many papers revealed the abnormal expression of circular RNA (circRNA), a kind of non-coding RNA, in mammals. However, the potential functional mechanisms are still unknown.
Objectives. In this paper, we aimed to elucidate the function and mechanisms of hsa-circ-0000098 in hepatocellular carcinoma (HCC).
Materials and methods. Bioinformatics was used to analyze the Gene Expression Omnibus (GEO) database (GSE97332) and predict the targeted gene site of miR-136-5p. The starBase online database was utilized to predict that MMP2 is the downstream target gene of miR-136-5p. The expression of hsa_circ_0000098, miR-136-5p and matrix metalloproteinase 2 (MMP2) in HCC tissues or cells was detected using quantitative real-time polymerase chain reaction (qRT-PCR) method. The migration and invasion abilities of processing cells were measured with transwell assay. The luciferase reporter assay was carried out to verify the targets of hsa_circ_0000098, MMP2 and miR-136-5p. The western blot assay was performed to detect the expression of MMP2, MMP9, E-cadherin, and N-cadherin.
Results. According to the analysis of GEO database of GSE97332, hsa_circ_0000098 had a prominent expression in HCC tissues. A continued analysis of relevant patients has verified that the high expression of hsa_circ_0000098 is present in HCC tissues with relative to poor prognosis. We also proved that the migration and invasion abilities of HCC cell lines can be inhibited by silencing hsa_circ_0000098. In view of the above findings, we continued to study the hsa_circ_0000098 mechanism of action in HCC. The study revealed that hsa_circ_0000098 can sponge miR-136-5p and then regulate MMP2, which is a downstream target gene of miR-136-5p, in order to promote HCC metastasis by regulation of miR-136-5p/MMP2 axis.
Conclusions. Our data showed that has_circ_0000098 facilitates the migration, invasion and malignant progression of HCC. On the other hand, we demonstrated that the mechanism of action of hsa_circ_0000098 in HCC might be due to the regulation of miR-136-5p/MMP2 axis.
Key words: hepatocellular carcinoma, MMP2, miR-136, hsa_circ_0000098
Background
Liver cancer is a malignant disease responsible for the majority of disease-induced deaths worldwide.1 As a kind of hepatic malignancy, hepatocellular carcinoma (HCC) has become the 3rd leading cause of mortality and the 6th most common malignancy.2 In the initial stage, liver cancer has hidden characteristics so that most people ignore the disease and get a belated diagnosis. Gene expression therapy or transplant is the common therapeutic method.3 Despite tremendous efforts invested in the past few decades, surgical techniques and chemoradiotherapy regimens that are used to cure HCC result in a limited prognostic improvement. The 5-year survival rate of HCC patients is not satisfactory.4, 5 Although some reports indicate that many genes are related to the carcinogenicity of HCC,6 there is no convincing evidence to expound the exact molecular mechanism. Therefore, it is important for researchers to explore the pathogenic process and molecular mechanism of HCC.
Circular RNAs (circRNAs), an important class of non-coding RNAs (ncRNAs), can participate in numerous physiological processes,7 including encoding proteins,8 sponging miRNA9 and regulating gene transcription.10 Gene expression can be regulated by ncRNAs at the transcriptional and post-transcriptional level, thus various ncRNAs play fundamental roles in the metastasis and tumorigenesis of HCC.11 On the other hand, different from usual linear RNAs, circRNAs have covalently closed loop structures without the polyadenylation tail, which are more stable when interacting with RNA exonuclease.12, 13 Circular RNAs are broadly expressed in mammals and their expression is related to various human diseases, especially cancer.14 They are correlated with metastasis, drug resistance and tumorigenesis.15 In the past few years, it has been reported that lots of circRNAs (hsa_cir_0007874,16 hsa_ccirc_000084717) have been reported to be involved in HCC and regulate the progression of the disease.14 Furthermore, there are studies indicating that circRNAs (hsa_circ_002059, hsa_circ_000009618, 19) can govern the migration and proliferation of gastric cancer cells according to the modulation of gene expression of matrix metalloproteinase-2 (MMP2), MMP9 and E-cadherin. However, functional mechanisms of circRNAs need further research.
Objectives
In this study, we performed an integrated analysis of circ_0000098, a kind of circRNA that has not been mentioned in the previous studies, to figure out the molecular and functional mechanisms for the diagnosis and therapy of HCC.
Materials and methods
Cell culture and samples
Four HCC cell lines (Hep3B, Huh7, MHCC97L, and SK-hep1) which are representative cell lines in HCC research, and normal human liver cell line (THLE-21) were purchased from American Type Culture Collection (ATCC, Manassas, USA). The cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Thermo Fisher Scientific, Waltham, USA) containing 10% fetal bovine serum (FBS; Gibco, Waltham, USA), 100 units/mL of penicillin, 100 µg/mL of streptomycin (Invitrogen, Carlsbad, USA), and 2 mM of L-glutamine in a humidified atmosphere containing 5% CO2 at 37°C. The tumor and adjacent noncancerous specimens were surgically dissected from 64 patients with HCC at Xiantao First People’s Hospital Affiliated to Yangtze University, Jingzhou, China. All the procedures were performed according to the Declaration of Helsinki. The protocols were discussed with the patients and written informed consent was obtained before the surgical collection of the samples. Samples were immediately stored at −80°C until further use. The study was approved by the ethical review committee of Xiantao First People’s Hospital Affiliated to Yangtze University (approval No. KY-E-2020-8-10).
Cell transfections
The vector responsible for lentiviral overexpression in circ_0000098 was transfected into Hep3B and SK-hep1 cell lines by adding puromycin into the stable cells. We performed circRNA knockdown using validated Stealth RNAi siRNA against circ_0000098 (Invitrogen), according to the manufacturer’s instructions. In brief, 4×105 cells were seeded in 60-millimeter plates, and siRNA treatment was performed the next day. A total of 100 μL transfection reagent mixture Lipofectamine™ 2000 (Thermo Fisher Scientific) was obtained after mixing 20 nM of transfection solutions with HiPerFect Transfection Reagent (Qiagen, Valencia, USA) and serum free media. The amount used and serum free media were used according to the instruction manual of the HiPerFect Transfection Reagent. Then, the transfection complexes were fed by incubating the mixture solution for 15 min. Subsequently, cells were inoculated with 10 μL solutions, the mixture solution was incubated at 37°C for 48 h and cells were collected 72 h after transfection for subsequent experimentation. The procedures were followed by gene silencing.
Nuclear/cytoplasmic fractionation
and quantitative real-time polymerase chain reaction assay
The extraction of nuclear and cytoplasmic RNA from tumor samples was performed using the extraction reagent (Pierce™ 660nm Protein Assay Reagent; Thermo Fisher Scientific). The amount of used liver cancer cells was equal to 1×107 and the principal method complied with the manufacturer’s instructions. RiboLock RNase inhibitors were introduced and the isolation of total RNA was performed using whole-cell lysates of RNase R achieved using RNAesay mini kits (Qiagen). The circumstantial experimental method was performed as following. A total of 250 μL of each fraction were instilled with 750 μL of TRIzol Reagent for RNA extraction. The cDNA synthesis was performed using 1 μg total RNA in 20 μL reaction volumes by means of All-In-One 5X RT MasterMix (Takara, Shiga, Japan). The U6 was used as a reference gene of miR-136-5p, and GAPDH was used as a reference gene of hsa_circ_0000098 and MMP2. Relative gene expression was measured using the 2−ΔΔCt method. The primer sequences were as following:
hsa_circ_0000098 forward:
5’-GGTGTAATTGCTTCTGCCATCA-3’,
reverse: 5’-GTCCAGCCAAAATGGCAGTG-3’;
miR-136-5p forward:
5’-ACACTCCAGCTGGGACTCCATTTGTTTT-3’,
reverse: 5’-CCAGTGCAGGGTCCGAGGT-3’;
MMP2 forward: 5’-GTGAAGTATGGGAACGCCG-3’,
reverse: 5’-GCCGTACTTGCCATCCTTCT-3’;
GAPDH forward: 5’-GAAAGCCTGCCGGTGACTAA-3’,
reverse: 5’-GCGCCCAATACGACCAAATC-3’;
U6 forward: 5’-CTCGCTTCGGCAGCACA-3’,
reverse: 5’-AACGCTTCACGAATTTGCGT-3’.
RNA pull-down assay
For RNA pull-down assay, biotinylated miR-136-5p probe was obtained from RiboBio (Guangzhou, China). The C-1 magnetic beads (Thermo Fisher Scientific) were used to incubate with the probe in order to acquire probe-coated beads. Then, they were incubated with cell lysates Hep3B and SK-hep1 at 4°C. After the washing of RNA complexes, quantitative real-time polymerase chain reaction (qRT-PCR) assay was performed to determine the expression of miR-136-5p and MMP2.
RNA immunoprecipitation assay
The RNA immunoprecipitation assay (RIPA) was performed using EZMagna RIP kit obtained from EMD Millipore (Burlington, USA). The relevant experiments were carried out in accordance with the manufacturer’s guidelines. Briefly, cells (Hep3B, SK-hep1) were lysed with RIPA lysis buffer. The supernatant was harvested after the centrifugation and cultured with magnetic beads conjugated to human anti-Ago2 antibody (Abcam, Cambridge, USA) or anti-IgG antibody (Abcam). Finally, the expression of hsa_circ_0000098 and miR-136-5p was detected using qRT-PCR assay.
Transwell assay
Transwell chambers (Corning Company, Corning, USA) were used to evaluate the migration and invasion abilities of HCC cells. In brief, liver cancer cells (about 1×104 cells) were resuspended and planted into upper chambers with or without the addition of Matrigel (BD Bioscience, Waltham, USA). At the same time, 800 μL medium with 30% FBS was added to the lower chambers. All chambers were incubated for 24 h at 37°C. Subsequently, transmigrated cells were set in the chambers. Then, crystal violet (Sigma-Aldrich, St. Louis, USA) was utilized to stain the cells. Finally, Nikon Diaphot inverted microscope (Nikon Corp., Tokyo, Japan) was utilized to count and photograph the migrated or invaded cells.
Western blot
Cells were lysed with RIPA lysis buffer, and BCA Protein Assay Kit (Beyotime Biotechnology, Beijing, China) was utilized to quantify them. Total proteins (40 μg) were charged to sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel and transferred onto a polyvinylidene difluoride (PVDF) membrane (EMD Millipore). The membrane was blocked with 5% skim milk and incubated with MMP2 (cat. No. AF1420), MMP9 (cat. No. AF5234), E-cadherin (cat. No. AF6759), and N-cadherin (cat. No. AF0243) antibodies (1:1000; Beyotime Biotechnology). After incubation, the secondary antibody horseradish peroxidase (HRP)-labeled Goat Anti-Rabbit IgG (H+L) (cat. No. A0208, 1:1000; Beyotime Biotechnology) was utilized to incubate the membrane. The membrane was then visualized with an electrochemiluminescence (ECL) reagent (Beyotime Biotechnology). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH, cat. No. AF1186; Beyotime Biotechnology) was used as an internal control.
Dual-luciferase reporter assay
The hsa_circ_0000098 and MMP2 wild type (hsa_circ_0000098 wt and MMP2 wt) and mutant type (hsa_circ_0000098 mut and MMP2 mut) reporter vectors were set by Beijing TransGen Biotech Co. Ltd. (Beijing, China). The reporter plasmids were co-transfected with either miR-136-5p mimics or negative mimics control into Hep3B cells. After transfection for 48 h, the cells were lysed and the relevant luciferase activities were detected using a dual-luciferase reporter assay system (Promega, Madison, USA).
Statistical analyses
The IBM SPSS v. 20.0 (IBM Corp., Armonk, USA) was used to perform the statistical analyses. The Student’s t test and one-way analysis of variance (ANOVA) followed by the Tukey’s post hoc test were utilized to analyze 2 or multiple groups, respectively. The survival curves were assessed by the Kaplan–Meier (K–M) analysis. The log-rank test was used to analyze the difference between K–M curves. The χ2 test was used to examine the correlation between hsa-circ-0000098 expression and clinical data. The normality of the data and the homogeneity of variance between the groups were tested using the Shapiro–Wilk test and the Levene’s test, respectively. The value of p > 0.05 indicated that the assumption of normality of data and homogeneity of variance was consistent, and further parameter testing could be performed. The number of independent replications in all the experiments amounted to 3. The data are shown as mean ± standard deviation (M ±SD). The value of p < 0.05 was considered statistically significant. The results of the statistical analyses are presented in Table 1.
Results
High expression of hsa_circ_0000098 in HCC gives rise to worse prognosis
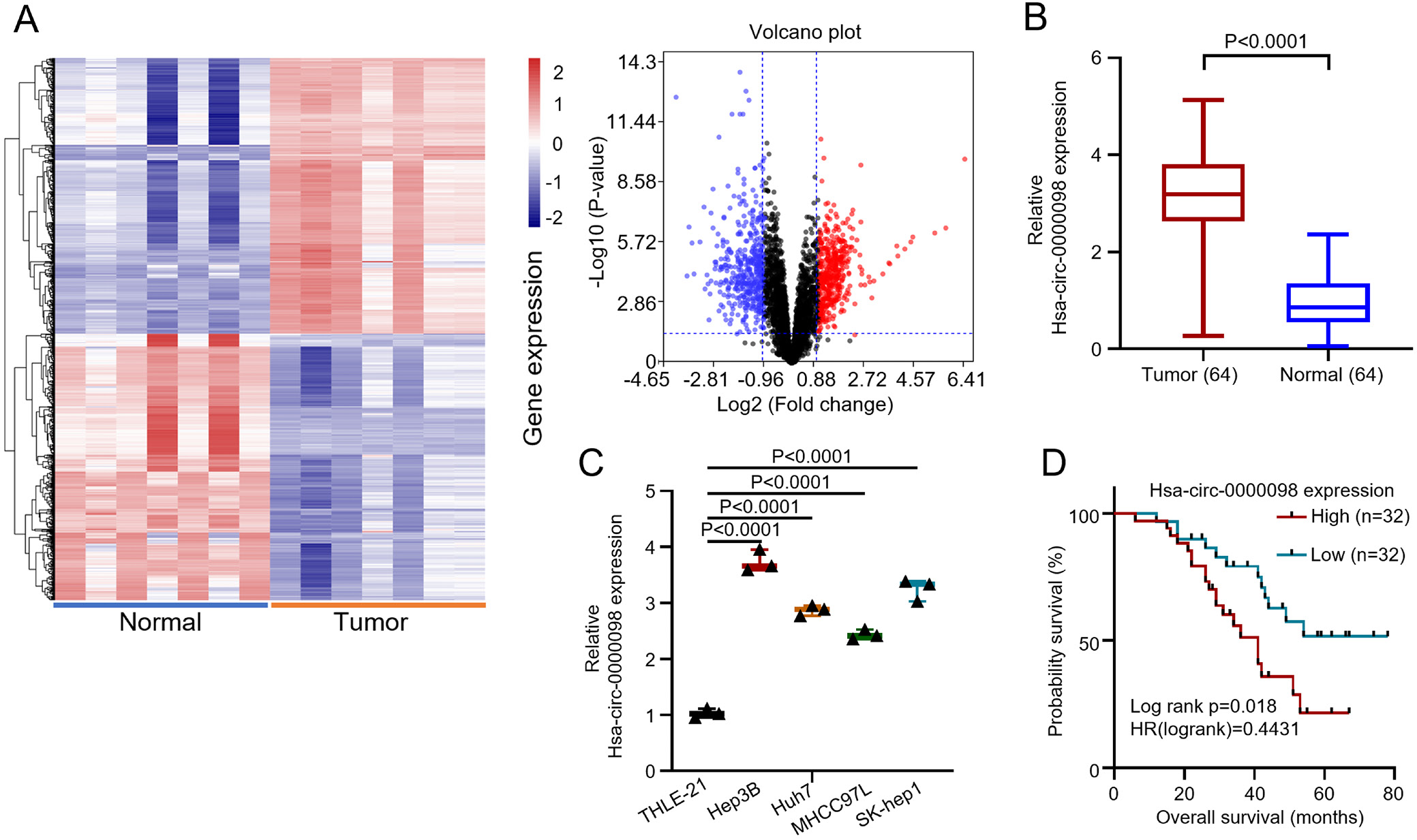
In order to investigate the impact of hsa_circ_0000098 on the development of HCC, 64 HCC patients have been chosen for this study. Their biopsied HCC and para-carcinoma tissues were collected and analyzed. To investigate the hsa_circ_0000098 expression profiles in HCC, we analyzed the microarray data from Gene Expression Omnibus (GEO) database GSE97332. Compared with normal tissues, hsa_circ_0000098 was highly expressed in HCC tumor tissues, and the heat map and volcano plot both indicated the diversity of circRNAs (Figure 1A). The qRT-PCR indicated that hsa_circ_0000098 was highly expressed in HCC tissues (Figure 1B; t = 14.88, degrees of freedom (df) = 126, p < 0.001). Furthermore, in order to examine the difference in the hsa_circ_0000098 expression between HCC cells and normal cells, several HCC cell lines (Hep3B, Huh7, MHCC97L, SK-hep1) were compared with normal human liver cell (THLE-21). The results show that hsa_circ_0000098 has a high expression in all HCC cell lines, while the normal liver cell, THLE-21, shows an inferior expression (F(4,10) = 161.7, p < 0.001; Figure 1C). As for Figure 1D, we selected the median expression value of the HCC tissues in Figure 1B as the cutoff value, and patients with HCC were divided into hsa_circ_0000098 high expression group (n = 32) and low expression group (n = 32). The K–M survivorship curve indicated that the prognostic survival rate of hsa_circ_0000098 high expression group is distinctly lower than that of the low expression group. Overall, the above data show that hsa_circ_0000098 is highly expressed in HCC, which gives rise to worse prognosis.
The relevant clinicopathological characteristics of patients are presented in Table 2, according to the degree of hsa_circ_0000098 expression after the χ2 testing. The results revealed that the expression of hsa_circ_0000098 is closely related to tumor size (p = 0.024), tumor-node-metastasis (TNM) stage (p = 0.024), differentiation (p = 0.035), and lymph node metastasis (p = 0.045). Inversely, the expression of hsa_circ_0000098 does not correlate with patient’s age and gender.
Silence of hsa_circ_0000098 inhibits
cell migration and invasion in vitro
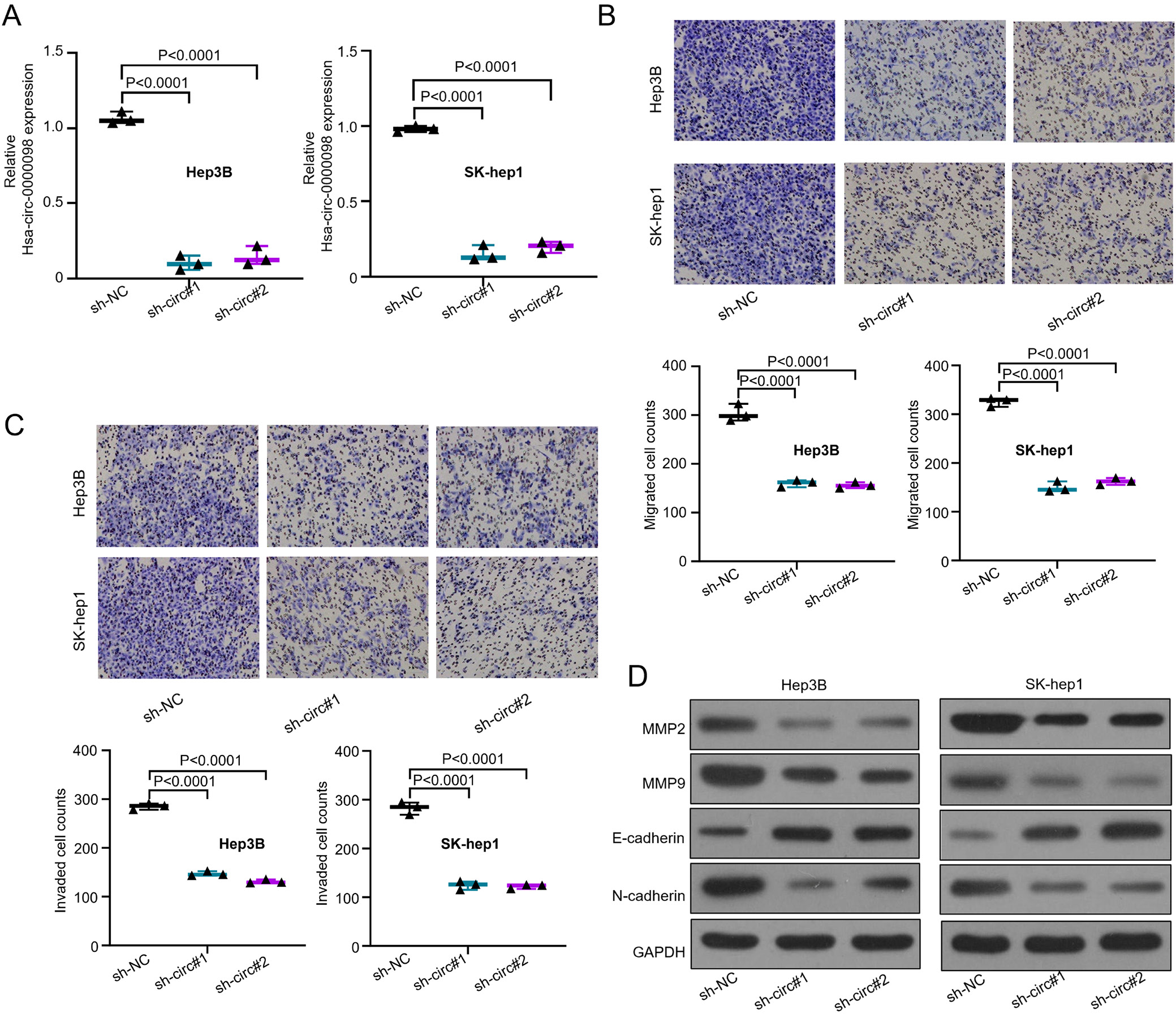
In order to explore the relationship between severe HCC and hsa_circ_0000098, an established knockdown method was conducted in order to silence the hsa_circ_0000098 expression. First, Hep3B and SK-hep1 with the highest expression of hsa_circ_0000098 (Figure 1C) were chosen as experimental targets to be silenced by 2 designed shRNAs (sh-circ#1 and sh-circ#2). Figure 2A (F(2,6) = 346.3, p < 0.001, and F(2,6) = 442.1, p < 0.001) indicates that compared with short hairpin RNA negative control (sh-NC), sh-circ#1 and sh-circ#2 can effectively knock out more than 70% of hsa_circ_0000098 from Hep3B and SK-hep1 cells. The hsa_circ_0000098 was successfully knocked down. Subsequently, the transwell assay with or without pre-coated Matrigel was used to detect the migration and invasion abilities of Hep3B and SK-hep1 cells. The results proved that after the knockdown of hsa_circ_0000098, the migration and invasion abilities of Hep3B and SK-hep1 cells decreased (F(2,6) = 159.4, p < 0.001, and F(2,6) = 349.4, p < 0.001, Figure 2B; F(2,6) = 813.6, p < 0.001, and F(2,6) = 300.9, p < 0.001; Figure 2C).
On the other hand, matrix metalloproteinases (like MMP2 and MMP9) can degrade almost all protein compositions in the extracellular matrix (ECM) and destroy the histological barriers of tumor cell invasion, playing a vital role in the tumor migration and invasion. Additionally, the tumor progression and migration are linked to E-cadherin and N-cadherin. Hence, through western blot assay, we detected the expression of MMP2, MMP9, E-cadherin, and N-cadherin in Hep3B and SK-hep1 cells transfected with sh-NC, sh-circ#1 and sh-circ#2. The expression of MMP2, MMP9 and N-cadherin shows a distinct decrease after the knockdown of hsa_circ_0000098 both in Hep3B and SK-hep1 cells (Figure 2D). The expression of E-cadherin shows a distinct increase. The results suggest that hsa_circ_0000098 has an important role in HCC migration and invasion.
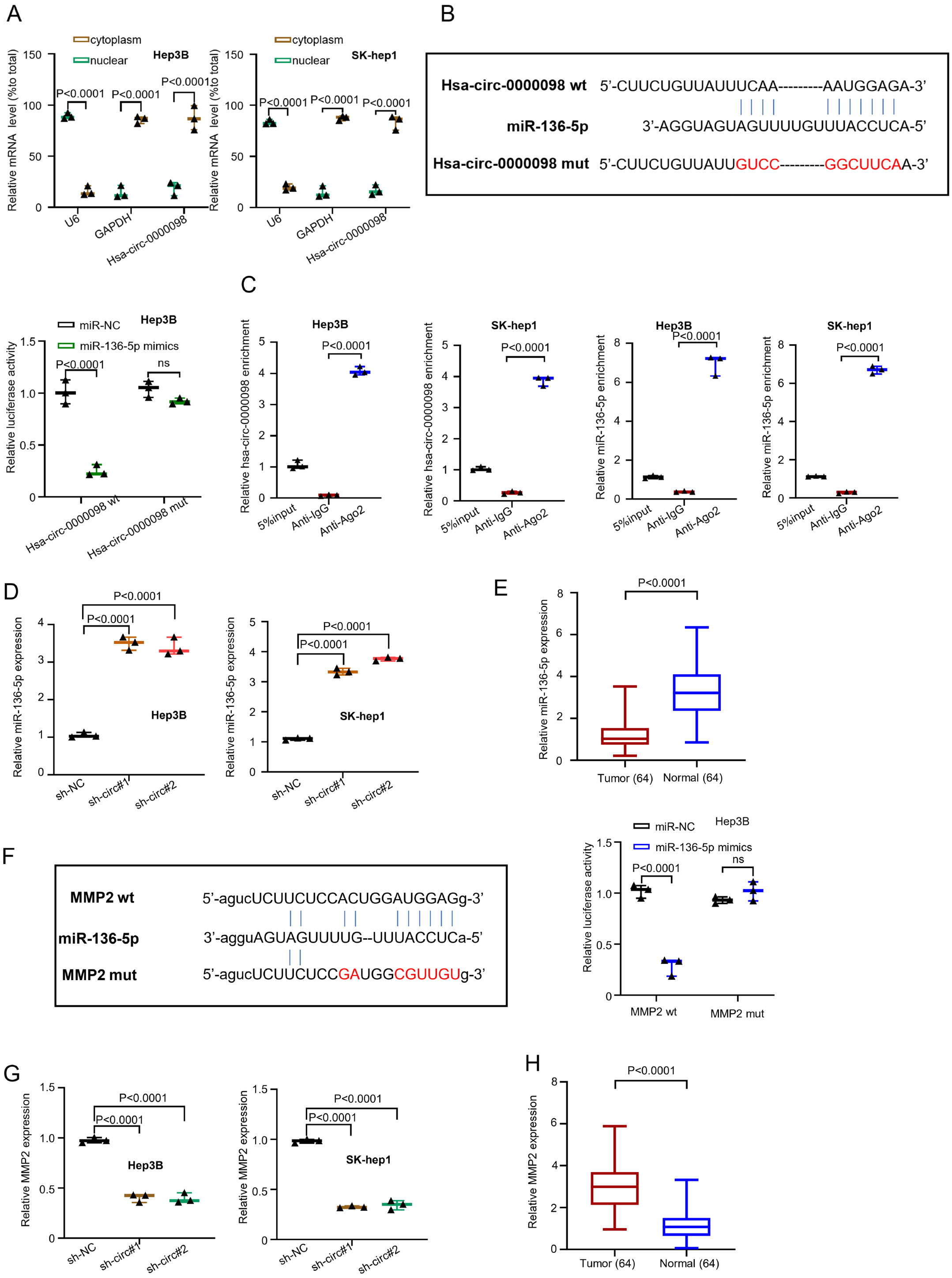
Hsa-circ-0000098 is expressed in cytoplasm and functions as ceRNA through sponge mir-136-5p/MMP2 signal axis
The circRNA has the potential to act as a competitive endogenous RNA (ceRNA) to sponge miRNAs and mitigate the inhibitory effect on the targeted mRNA expression.20 In order to explore the hsa_circ_0000098 mechanism of action in HCC, we performed a series of experiments. First, the location of the highest hsa_circ_0000098 expression was estimated using qRT-PCR through nuclear/cytoplasmic fractionation. The graph presented in Figure 3A (F(2,12) = 234.9, p < 0.001, and F(2,12) = 440.3, p < 0.001, and F(1,12) = 50.04, p < 0.001, and F(1,12) = 142.6, p < 0.001) indicates that most of hsa_circ_0000098 can be detected in the cytoplasm. However, the mature miRNAs are also present in the cytoplasm and there is the possibility of hsa_circ_0000098 to comb with miRNAs. The U6 and GAPDH were set as internal references of nucleus and cytoplasm, respectively. Then, we used a bioinformatics database (circRNA interactome: https://circinteractome.irp.nia.nih.gov/) to predict the potential target miRNA. We found that hsa_circ_0000098 can target miR-136-5p (Figure 3B). Subsequently, the luciferase assay was conducted with Hep3B cells to confirm the prediction (Figure 3C). The results indicate that compared with miRNA negative control (miR-NC), the overexpressed miR-136-5p can inhibit the luciferase activity of hsa_circ_0000098 wt in Hep3B cells. However, when we mutated circRNA, which was the binding site of predicted miR-136-5p, the corresponding inhibition disappeared (F(1,8) = 53.46, p = 0.0001; Figure 3B). Then, the RIPA-qRT-PCR method was performed on Hep3B and SK-hep1 cells (F(2,6) = 991.6, p = 0.0001; F(2,6) = 1099, p = 0.0001; F(2,6) = 402.4, p = 0.0001; F(2,6) = 2692, p = 0.0001; Figure 3C). The results show that hsa_circ_0000098 could bind to miR-136-5p, whereas the beads coupled with AGO2 pulled down significantly greater hsa_circ_0000098 and miR-136-5p than the IgG control. The results presented in Figure 3D (F(2,6) = 184.4, p < 0.001, and F(2,6) = 1048, p < 0.001) indicate that both sh-circ#1 and sh-circ#2 can increase the expression of miR-136-5p in Hep3B and SK-hep1 cells. The qRT-PCR indicated that miR-136-5p is downregulated in HCC tissues (t = 10.78, df = 126, p < 0.001; Figure 1E). Figure 3F presents the consequence of 3’-UTR of the MMP2 gene on the binding site of miR-136-5p. The analysis was predicted using starBase on-line database (http://starbase.sysu.edu.cn). On the other hand, the luciferase assay was undertaken with Hep3B (F(1,8) = 95.55, p < 0.001; Figure 3F). The results suggest that compared with miR-NC, the overexpressed miR-136-5p can inhibit the luciferase activity of MMP2 wt in Hep3B cells. However, when the binding site of predicted miR-136-5p was mutated, the corresponding inhibition disappeared. Finally, the variation of MMP2 expression in Hep3B and SK-hep1 cells was measured using qRT-PCR, after the knockdown of hsa_circ_0000098. The results indicate that in both Hep3B and SK-hep1 cells, sh-circ#1 and sh-circ#2 can decrease the expression of MMP2 (F(2,6) = 201, p < 0.001; F(2,6) = 494.8, p < 0.001; Figure 3G). Similar to hsa_circ_0000098, MMP2 was upregulated in HCC tissue (t = 11.16, df = 126, p = 0.0001; Figure 3H). The sh-NC was used to make comparisons with sh-circ#1 and sh-circ#2.
Hsa_circ_0000098 promotes
HCC migration and invasion
by regulating miR-136-5p/MMP2 axis
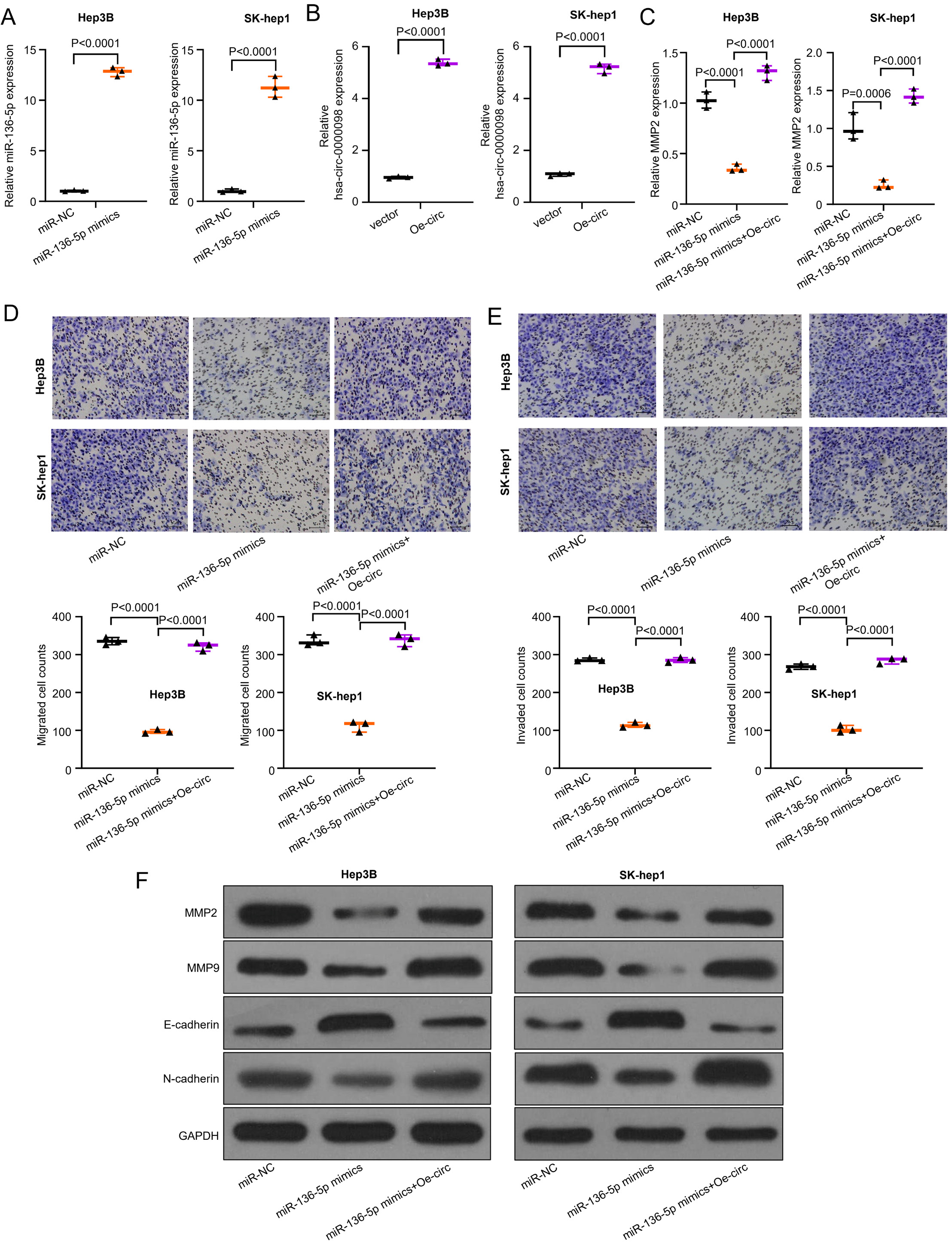
The described experiments proved that hsa_circ_0000098 plays a critical role in HCC cell migration and invasion. Yet, the exact regulatory mechanism is still unknown. In view of this and previous studies, we designed a series of experiments to examine the relevant mechanism. Synthesized miR-136-5p mimics were overexpressed in Hep3B and SK-hep1 cells. Thus, qRT-PCR method was used to evaluate the efficiency of overexpression. The used symbols and abbreviations are as follows: t1 – Student’s t-test results of miR-NC compared to miR-136-5p mimics group; t2 – t-test results of miR-136-5p mimics compared to miR-136-5p mimics+Oe-circ group; df1 – degrees of freedom of miR-NC compared to miR-136-5p mimics group; df2 – degrees of freedom of miR-136-5p mimics compared to miR-136-5p mimics+Oe-circ group (p < 0.05 was considered statistically significant); p1 – p-value of miR-NC compared to miR-136-5p mimics group; p2 – p-value of miR-136-5p mimics compared to miR-136-5p mimics+Oe-circ group.
The results indicate that miR-136-5p mimics, compared with miR-NC, can be overexpressed in both Hep3B and SK-hep1 cells (t1 = 45.51, df1 = 4, p1 < 0.001; t2 = 17, df2 = 4, p2 < 0.001; Figure 4A). Then, the overexpressed hsa_circ_0000098 (Oe-circ) was constructed by transfecting Hep3B and SK-hep1 cells. Figure 4B (t1 = 56.28, df1 = 4, p1 < 0.001; t1 = 36.25, df1 = 4, p1 < 0.001) displays the results of the evaluation of the efficiency of overexpression: compared with vector, Oe-circ can be effectively overexpressed with hsa_circ_0000098 in Hep3B and SK-hep1 cells. Thus, after the overexpression of miR-136-5p, we detected the expression variation of MMP2. The results revealed that the overexpressed miR-136-5p decreases the expression of MMP2 (F(2,6) = 163.7, p = 0.0001; F(2,6) = 72.01, p = 0.0001; Figure 4C). After the co-transfection of Oe-circ, the expression of MMP2 has increased. The transwell assay was used to detect the migration and invasion abilities of different groups (miR-NC, miR-136-5p mimics and miR-136-5p mimics+Oe-circ) in Hep3B and SK-hep1 cells. The similar results presented in Figure 4D (F(2,6) = 659, pSK = 0.3611; pL = 0.7111 (pSK – Shapiro–Wilk test; pL – Levene’s test); F(2,6) = 238.3, p < 0.001) and Figure 4E (F(2,6) = 839.9, p < 0.001, and F(2,6) = 462, p < 0.001) indicate that the overexpressed miR-136-5p can decrease the migration and invasion abilities of these 2 cell lines, while after the co-transfection of Oe-circ, the migration and invasion abilities of Hep3B and SK-hep1 cells increase (Figure 4D,E). Finally, the western blot assay was used to detect the expression of MMP2, MMP9, E-cadherin, and N-cadherin in different groups (miR-NC, miR-136-5p mimics, miR-136-5p mimics+Oe-circ) in Hep3B and SK-hep1 cells. The results show that miR-136-5p decreased the expression of MMP2, MMP9 and N-cadherin, and increased the expression of E-cadherin (Figure 4F); however, after the co-transfection of Oe-circ, the expression of MMP2, MMP9 and N-cadherin increased and the expression of E-cadherin decreased, which corresponds with our assumptions.
Discussion
Hepatocellular carcinoma is the 3rd leading cause of the cancer-related mortality worldwide. It is a multi-factorial disease caused by various risk factors such as cirrhosis, alcohol abuse, viral infections, and others.21, 22, 23, 24 The circRNAs have drawn attention as oncogenes or suppressors of cancer progression.16, 25 Since circRNAs are nonlinear reverse splicing structures, they are more stable even under the dispose of RNase R.19, 26, 27, 28 Until now, the studies have described circRNAs as ceRNAs which can sponge miRNA in order to regulate mRNA expression in HCC.27, 29, 30 For example, circRNA hsa_circRNA_104348 promotes HCC progression through modulating miR-187-3p/RTKN2 axis and activating Wnt/β-catenin pathway.31 The circRNA-5692 inhibits the progression of HCC by sponging miR-328-5p to improve the expression of DAB2IP.32 The circRNA-104718 acts as a competing endogenous RNA and promotes HCC progression through microRNA-218-5p/TXNDC5 signaling pathway.33 However, the specific function and mechanism of hsa_circ_0000098 associated with tumorigenesis and progression of HCC are still unclear. In this study, we designed a series of experiments to explore the specific role of hsa_circ_0000098 in HCC and confirmed that hsa_circ_0000098 is highly expressed in HCC. Afterwards, the in vitro studies confirmed that hsa_circ_0000098 is extremely involved in the HCC progression.
In view of the above conclusions, we continued to study the mechanism of action of hsa_circ_0000098 in HCC. CircInteractome online database predicted that hsa_circ_0000098 can sponge miR-136-5p. To the best of our knowledge, miR-136-5p was reported to have a low expression in liver cancer.3, 34 Additionally, miR-136-5p has an impact on carcinomatosis in various cancers (colorectal carcinoma,35 gallbladder carcinoma,36 human T-cell leukemia virus type 1 (HTLV-1),37 thyroid cancer,38 oral squamous cell carcinoma39). Remarkably, miR-136-5p has been identified to be downregulated in HCC. For instance, silencing of lncRNA LEF1-AS1 prevents the progression of HCC via the crosstalk with miR-136-5p/WNK1.40 The miR-136-5p is a target gene for circ_0027089, thereby modulating NACC1 to aggravate HCC is associated with hepatitis B virus.41 In our study, miR-136-5p was downregulated in HCC tissues, which is consistent with the literature on the subject.
The downstream target gene of miR-136-5p is predicted to be MMP2, which is one of the prominent members of MMP family. The MMP2 and MMP9 are usually highly expressed in malignant tumors and take part in the degradation of tumor ECM and base, assisting tumor cells in breaking the metastasis barriers.42 Dorandish et al. revealed that MMP2 is the predominant regulator of cytotoxic effects brought about by Aβ in lung cancer cells,43 especially in HCC.44 Hence, MMP2 and MMP9 play crucial roles in the migration and invasion of tumor cells.
Epithelial–mesenchymal transition (EMT) is a biological process in which epithelial cells transform and exhibit a mesenchymal phenotype via a specific program.45 It has been shown that EMT exhibits a significant effect on malignant tumors, which could transform epithelial cells to obtain migration and invasion abilities associated with malignancy.46 Thereby, it is important to investigate the molecular mechanism underlying the EMT process by N-cadherin and E-cadherin expression. In the present study, the results revealed that N-cadherin decreased and E-cadherin increased after the co-transfection of sh-circ, which means that the knockdown could suppress the ability to migrate and invade.
In conclusion, hsa_circ_0000098, highly expressed in HCC, can mediate the migration, invasion and malignant progression of liver cancer. This might be due to the regulation of miR-136-5p/MMP2 axis. This study has also provided inspiration for relieving pain of HCC patients. For the first time, we made efforts in exploring the relationship and mechanism of action between hsa_circ_0000098 and HCC, which will pave the way for the subsequent study.
Limitations
The study did not verify the signal pathways studied in in vivo experiments. Subsequent research should focus on in vivo experiments to provide more reliable data and strategic treatment for future clinical studies.
Conclusions
Our data demonstrated that hsa_circ_0000098 is highly expressed in HCC, which facilitates the migration, invasion and malignant progression of cancer cells. On the other hand, we also proved that the mechanism of action of hsa_circ_0000098 in HCC might be due to the regulation of miR-136-5p/MMP2 axis.
Data availability
All supporting data are available from the corresponding author upon request.