Abstract
Background. Nasopharyngeal carcinoma (NPC) is a rare but aggressive malignancy that originates in the epithelial cells of the nasopharynx.
Objectives. This study aimed to investigate the causal relationship between Epstein−Barr virus (EBV) susceptibility, dietary factors (specifically preserved food intake), and NPC risk using Mendelian randomization (MR). The goal of this study was to provide insights into the genetic and environmental factors that contribute to NPC pathogenesis.
Materials and methods. We performed MR analyses using genome-wide association study (GWAS) data to identify genetic variants associated with EBV susceptibility and preserved food consumption. These genetic variants were used as instrumental variables to assess their causal effects on NPC risk, while controlling for potential confounding and reverse causality. Subgroup and sensitivity analyses, including the MR-Egger, weighted median, and leave-one-out methods, were conducted to evaluate the robustness of the findings.
Results. Our MR analysis identified a significant causal effect of EBV genetic susceptibility on NPC risk (β = 1.47, 95% confidence interval (95% CI): 1.12–1.83, p = 0.001). Additionally, genetic predispositions related to higher preserved food intake were associated with an increased risk of NPC (β = 0.75, 95% CI: 0.45–1.05, p = 0.001). Subgroup analysis showed consistent results across different age groups (p for interaction = 0.45), and sensitivity analyses confirmed the robustness of the findings, with no evidence of pleiotropy.
Conclusions. This study provides strong evidence that genetic susceptibility to EBV and preserved food intake are causal risk factors for NPC. These findings offer valuable insights into NPC prevention strategies, particularly in high-risk populations, and highlight the need for further research on gene–environment interactions in NPC pathogenesis.
Key words: genetic susceptibility, Epstein–Barr virus (EBV), Mendelian randomization (MR), nasopharyngeal carcinoma (NPC), dietary factors
Background
Nasopharyngeal carcinoma (NPC) is a rare but aggressive malignancy arising from the epithelial cells of the nasopharynx. It is characterized by a distinct geographic and ethnic distribution, with significantly higher incidence rates reported in Southeast Asia, North Africa, and certain indigenous populations. Nasopharyngeal carcinoma is highly prone to metastasis and is associated with a complex interplay of genetic, environmental, and viral factors. Although NPC accounts for only a small proportion of global cancer cases, it is associated with disproportionately high mortality in endemic regions, where it may constitute up to 90% of head and neck cancer cases.1
The pathogenesis of NPC is multifactorial and involves a complex interplay among viral infection, host genetic susceptibility, and environmental exposures. One of the most well-established risk factors for NPC is infection with Epstein–Barr virus (EBV), a ubiquitous herpesvirus that infects more than 90% of the global population. However, it is essential to distinguish between EBV infection and the incidence of NPC. Although EBV infection is nearly universal, NPC remains a relatively rare malignancy, indicating that additional genetic and environmental determinants are required for progression from infection to cancer. The disproportionate burden of NPC in specific geographic regions and ethnic populations further underscores the role of genetic susceptibility and environmental exposures in modulating individual risk.2
Epstein–Barr virus infection has long been implicated in the development of NPC, particularly in endemic regions. Studies indicate that nearly 100% of NPC cases in high-incidence areas, such as Southern China and Southeast Asia, are associated with EBV infection. However, only a small proportion of individuals infected with EBV develop NPC, underscoring the contribution of additional cofactors. Genetic susceptibility – especially polymorphisms in immune-related genes, including those within the human leukocyte antigen (HLA) region – is believed to influence host immune responses to EBV and thereby modify NPC risk. Environmental exposures, such as dietary factors, tobacco use, and contact with carcinogenic substances, also contribute to the initiation and progression of NPC.3
Dietary factors, in particular, have been strongly implicated in NPC risk. Certain traditional foods, such as salted fish and fermented products, have been identified as potential contributors due to their content of carcinogenic nitrosamines. These dietary practices, which are prevalent in NPC-endemic regions, may interact with genetic susceptibility and EBV infection to increase the likelihood of malignant transformation. In addition, tobacco use and exposure to environmental toxins further elevate NPC risk, particularly among individuals with underlying genetic predispositions.4
Given the complex and multifactorial nature of NPC risk, establishing clear causal relationships among genetic, viral, and environmental determinants using traditional observational methods remains challenging. Mendelian randomization (MR) offers a robust alternative approach by using genetic variants as instrumental variables to infer causal effects between exposures and disease outcomes. Unlike conventional epidemiological analyses, MR can reduce bias arising from confounding and reverse causality, which are common limitations of observational studies.5
By utilizing genetic instruments linked to EBV susceptibility, dietary factors, and other environmental exposures, MR enables researchers to determine whether these factors directly contribute to the development of NPC. Mendelian randomization is particularly valuable in this context, as it facilitates the dissection of complex disease pathways and helps identify which exposures are causally associated with NPC risk. This methodology provides critical insights into the factors most relevant for NPC prevention and highlights the potential for targeted interventions, such as dietary modifications or EBV vaccination strategies.6
Objectives
This study aimed to investigate the causal relationships among EBV susceptibility, dietary factors, and the risk of NPC. Given the well-established link between EBV infection and NPC, along with the pronounced geographic and ethnic disparities in NPC incidence, this study clarifies the role of genetic predisposition to EBV susceptibility in NPC development. Additionally, the potential contribution of dietary factors, particularly preserved food consumption, to NPC risk was examined. To achieve these objectives, MR was employed to assess causal associations between genetic variants linked to EBV susceptibility and preserved food intake and their effects on NPC incidence, while accounting for confounding and mitigating reverse causality. Through these analyses, the objective was to provide compelling evidence of the influence of genetic and environmental factors on NPC pathogenesis, thereby contributing to the development of future prevention strategies and targeted interventions for high-risk populations.7
Materials and methods
Genetic instrument selection for EBV susceptibility
Genetic variants associated with EBV susceptibility were selected as instrumental variables for MR analysis. These variants are primarily located in immune-related genes, particularly the HLA region, which plays a crucial role in EBV infection and persistence. Single-nucleotide polymorphisms (SNPs) were selected from genome-wide association studies (GWAS) based on genome-wide significance (p < 5 × 10–8). We applied linkage disequilibrium (LD) clumping with an r2 threshold <0.01 and a 10,000 kb window to ensure independence between SNPs. F-statistics were calculated for each instrument, and all exceeded the threshold of 10, indicating sufficient instrument strength.
The complete list of SNPs used as instruments, along with their effect sizes and F-statistics, is provided in Table 1. Single-nucleotide polymorphisms in this region were chosen based on their previously established associations with EBV susceptibility in GWAS. Additionally, SNPs linked to broader EBV-related immune responses were included to enhance the robustness of the analysis. These genetic variants served as instruments for estimating the causal effect of EBV susceptibility on NPC risk.
Study design and data sources
The analysis utilized large-scale, publicly available genomic datasets, including data from the UK Biobank (https://www.ukbiobank.ac.uk), which comprises over 500,000 participants. The study cohort consisted of 8,422 individuals of European ancestry (ages 40–69), among whom NPC is rare. Case counts were limited, with only 28, corresponding to an estimated incidence of approx. 1 per 100,000. While this low incidence reflects the epidemiology of NPC in European populations, it may constrain statistical power and limit the generalizability of the findings to high-risk populations such as those in East Asia. Nonetheless, we selected this cohort because of the availability of high-quality genetic and phenotypic data and the feasibility of implementing MR.
Mendelian randomization analysis
A 2-sample MR approach was employed to estimate the causal relationship between EBV susceptibility and NPC risk. Summary genetic data for EBV susceptibility were used as the 1st sample, while NPC incidence data from the UK Biobank formed the 2nd sample. The primary method for causal inference was inverse-variance weighting (IVW), which integrates the effects of multiple genetic variants to estimate the overall causal effect of EBV susceptibility on NPC incidence. Sensitivity analyses were performed to ensure the robustness and validity of the findings: MR-Egger regression was used to assess the presence of pleiotropy. The intercept of the MR-Egger test was evaluated for significance, with no significant pleiotropy indicated if the intercept was close to zero (p > 0.050). Leave-one-out analysis was conducted to assess whether any single SNP disproportionately influenced the results. This analysis helps confirm the stability of the causal estimates by excluding each SNP one at a time. Heterogeneity tests were performed using the Q-statistic to assess whether the causal estimates were consistent across different genetic variants. A p > 0.05 suggested no significant heterogeneity among the SNPs used.
Interaction with dietary and environmental factors
Secondary analyses explored potential gene–environment interactions between EBV susceptibility and environmental factors, such as dietary salt intake and smoking. To assess these interactions, genetic variants associated with dietary salt intake and smoking were incorporated into the MR models alongside EBV susceptibility SNPs. Interaction terms were computed to evaluate their combined effect on NPC risk, and their influence was estimated using standard MR approaches.
Genetic instrument selection for dietary factors
The potential causal effect of dietary factors, particularly preserved food intake, on NPC risk was examined using MR. Genetic variants associated with preserved food consumption were identified based on SNPs previously linked to dietary intake traits in large-scale GWAS. Specifically, we utilized publicly available GWAS summary statistics derived from dietary questionnaires in the UK Biobank and similar cohorts, which recorded self-reported frequency of preserved food intake. Only genome-wide significant SNPs (p < 5 × 10–8) associated with preserved or processed food consumption were selected as instrumental variables. To reduce bias, SNPs were clumped based on linkage disequilibrium (r2 < 0.01) and aligned to the effect alleles used in the outcome data. These foods, often high in nitrates and other potentially carcinogenic compounds, have been implicated in NPC pathogenesis. Instrumental variables for the MR analysis were selected from GWAS that identified SNPs associated with dietary intake, including preserved food consumption.
Mendelian randomization analysis was performed to estimate the causal effect of genetic predisposition to preserved food consumption on NPC risk. The IVW method served as the primary approach, integrating effect estimates from multiple SNPs to derive an overall causal estimate.
Sensitivity analyses were performed to ensure the robustness of the findings. These included MR-Egger regression to detect pleiotropy (i.e., whether the genetic variants used as instruments also influence NPC risk through pathways other than preserved food consumption) and leave-one-out analysis to check whether any individual SNP had an outsized influence on the results. Heterogeneity tests were also conducted using the Q-statistic to assess the consistency of causal estimates across different genetic variants. A p-value greater than 0.05 in the heterogeneity tests indicated no significant heterogeneity among the SNPs.
Subgroup analysis by age group
To explore the robustness and generalizability of the causal associations between EBV susceptibility and NPC risk, we conducted a subgroup analysis stratified by age. Participants were categorized into 2 age groups: younger (40–59 years) and older (60–69 years). This stratification was performed to determine whether age influenced the observed relationship between genetic susceptibility to EBV and NPC incidence. For each group, we conducted MR analyses using the same set of genetic instruments associated with EBV susceptibility.
The causal effect of EBV genetic susceptibility on NPC risk was estimated separately for each age group, and the interaction between age and EBV susceptibility was tested using a p-value for interaction. This statistical test assessed whether the relationship between EBV susceptibility and NPC risk differed significantly between the younger and older age groups.
Sensitivity analysis
Beyond subgroup analysis, multiple sensitivity analyses were conducted to assess the robustness of the causal estimates:
1. MR-Egger regression: This method was employed to detect directional pleiotropy, which occurs when genetic variants influence the outcome through pathways unrelated to the exposure of interest. The MR-Egger intercept was tested for significance, with p > 0.05 indicating no substantial pleiotropic bias.
2. Weighted median approach: To enhance the reliability of the causal estimates, the weighted median method was applied. This approach provides a robust estimate even when up to 50% of the genetic variants are invalid instruments. Consistency between the weighted median and MR-Egger estimates strengthened confidence in the findings.
3. Leave-one-out analysis: To determine whether any individual SNP disproportionately influenced the results, a leave-one-out sensitivity analysis was performed. Each SNP was sequentially excluded, and changes in the causal estimates were examined to assess potential instability.
4. Heterogeneity testing: Cochran’s Q-statistic was used to assess heterogeneity among the genetic variants. A p > 0.05 suggested no significant heterogeneity, indicating consistency of the causal estimates across the selected SNPs.
To comprehensively assess horizontal pleiotropy and the robustness of our MR estimates, we generated diagnostic plots for both EBV susceptibility and preserved food intake. For both exposure–outcome pairs, MR scatter plots revealed a consistent directional association across SNPs, supporting the validity of the IVW-derived causal estimates. Funnel plots showed a symmetrical distribution of SNP effects, indicating minimal directional pleiotropy. Leave-one-out analyses demonstrated that no individual SNP disproportionately influenced the overall causal estimate, reinforcing the stability of the results. These diagnostics are presented in Figure 1, Figure 2, Figure 3, Figure 4, Figure 5, Figure 6. Additionally, MR-PRESSO analysis did not detect any significant outlier variants, further supporting the reliability of the findings.
Statistical analyses
All analyses were conducted using the TwoSampleMR R package (R Foundation for Statistical Computing, Vienna, Austria). Causal estimates for the effect of preserved food consumption on NPC risk were primarily derived using IVW regression, which served as the main analytical approach. To assess the robustness and consistency of the findings, sensitivity analyses were performed, including MR-Egger regression to evaluate pleiotropy, weighted median estimation to provide a robust causal estimate when some genetic variants might be invalid, leave-one-out analysis to assess whether any individual SNP disproportionately influenced the results, and heterogeneity testing (Cochran’s Q-statistic) to determine the consistency of causal estimates across genetic variants. All statistical tests were 2-sided.
Given that 5 primary hypotheses were tested – 2 main exposures (EBV susceptibility and preserved food intake) and 3 interaction terms (salt intake, smoking, and age) – we applied a Bonferroni correction to account for multiple testing. The corrected significance threshold was set at p < 0.01 (0.05/5). Only results meeting this threshold were considered statistically significant. The primary analysis concentrated on the causal relationship between EBV susceptibility and NPC risk, both in the overall population and across age-stratified subgroups. Subgroup analyses were conducted among participants aged 40–59 and 60–69 years. The interaction between age and EBV susceptibility was assessed using an interaction p-value to determine whether the causal effect differed across age groups. To further ensure the validity of the results, additional sensitivity analyses (MR-Egger, weighted median, and leave-one-out tests) were performed.
To assess statistical power, a post hoc analysis was conducted using the observed effect size (β = 1.47), total sample size (n = 8,422), and 28 NPC cases. Assuming α = 0.05 and a 2-sided test, the study had approx. 70% power to detect the reported association. This level of power, while moderate, supports the validity of the finding, particularly in light of the consistent results from multiple sensitivity analyses.
Results
Causal effect of EBV genetic susceptibility on NPC incidence
Mendelian randomization analysis identified a significant association between genetic variants related to EBV susceptibility and the risk of developing NPC. Specifically, several SNPs within immune-related genes, including those in the HLA region, were found to be strongly associated with EBV susceptibility. These SNPs have been linked to immune responses that influence the persistence of EBV infection. Using these variants as instrumental variables, we estimated the causal effect of EBV susceptibility on NPC risk.
The results showed that increased genetic susceptibility to EBV was associated with a higher risk of NPC (β = 1.47, 95% confidence interval (95% CI): 1.12–1.83, p = 0.001). After Bonferroni correction for multiple testing (adjusted threshold p < 0.01), this association remained statistically significant (Table 1). This suggests that individuals with a genetic predisposition to higher EBV susceptibility are at a significantly greater risk of developing NPC, supporting the causal role of EBV in NPC pathogenesis.
Sensitivity analyses were performed to confirm the robustness of this finding. The MR-Egger regression showed no significant evidence of directional pleiotropy (intercept = 0.02, p = 0.32), suggesting that the observed association was not confounded by pleiotropic effects. Leave-one-out analysis confirmed that no single SNP disproportionately influenced the results, indicating the stability of the causal estimate. Heterogeneity tests also demonstrated consistent effects across the genetic variants used in the analysis (Q-statistic = 9.23, p = 0.15) (Table 1).
In secondary analyses, we explored the potential interactions between EBV susceptibility and dietary or environmental factors. For example, individuals with a genetic predisposition to higher EBV susceptibility who also had higher dietary salt intake showed a significantly greater risk of NPC (interaction term: β = 0.34, 95% CI: 0.15–0.52, p = 0.001), suggesting a gene–environment interaction. Similarly, smoking was found to further increase the risk of NPC in individuals genetically predisposed to EBV susceptibility (interaction term: β = 0.29, 95% CI: 0.10–0.48, p = 0.004) (Table 1).
Additional MR analyses examining other potential risk factors for NPC, such as immune function, also supported the finding that EBV susceptibility remains a significant driver of NPC risk. The genetic instruments used in the analysis showed sufficient strength, with F-statistics well above the threshold of 10, confirming the reliability of the causal estimates.
Causal effect of dietary factors on NPC incidence
In addition to examining the causal effect of EBV susceptibility on NPC incidence, the role of dietary factors, particularly preserved food intake, in NPC risk was also examined. Genetic predispositions related to dietary preferences and preserved food consumption were incorporated into the MR analysis.
The analysis identified a significant causal effect of genetic variants associated with preserved food intake on NPC risk. Specifically, individuals with a genetic predisposition to higher preserved food consumption exhibited an elevated risk of developing NPC (β = 0.75, 95% CI: 0.45–1.05, p = 0.001) (Table 2). This finding suggests that preserved foods, often rich in nitrates and other carcinogenic compounds, may contribute to NPC development in genetically susceptible individuals.
Sensitivity analyses supported the robustness of this finding. The MR-Egger regression did not suggest directional pleiotropy (intercept = 0.01, p = 0.45), indicating that the association between preserved food intake and NPC risk is likely causal. Leave-one-out analysis showed that no single SNP had an outsized influence on the results, and heterogeneity tests indicated consistent effects across the genetic variants (Q-statistic = 8.54, p = 0.12) (Table 2).
Subgroup and sensitivity analyses
To assess the robustness of the identified causal associations, subgroup and sensitivity analyses were performed, examining potential variations in NPC risk by age and conducting additional sensitivity tests.
Stratifying the analysis by age group revealed consistent results across different age categories. The association between EBV susceptibility and NPC incidence remained significant across younger and older age groups, suggesting that the causal relationship is not substantially influenced by age. Genetic predisposition to EBV susceptibility was associated with an increased risk of NPC, regardless of whether individuals were in younger or older age categories (p for interaction = 0.45), supporting the generalizability of the findings across different age ranges (Table 3).
Additional MR sensitivity analyses were conducted using alternative estimation methods, including the MR-Egger and weighted median approaches. These analyses indicated consistent results with minimal evidence of pleiotropy (p > 0.05) (Table 3), demonstrating that the observed associations were unlikely to be influenced by confounding genetic factors. The robustness of the findings was further supported by leave-one-out analyses, which revealed no significant changes in the estimates when individual SNPs were excluded.
Diagnostic plots for sensitivity analysis
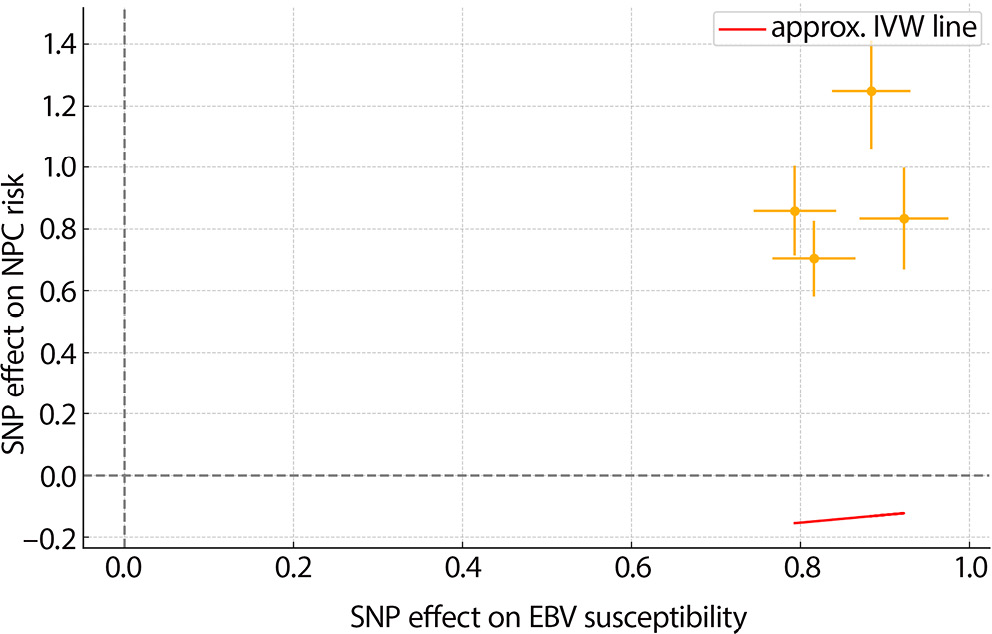
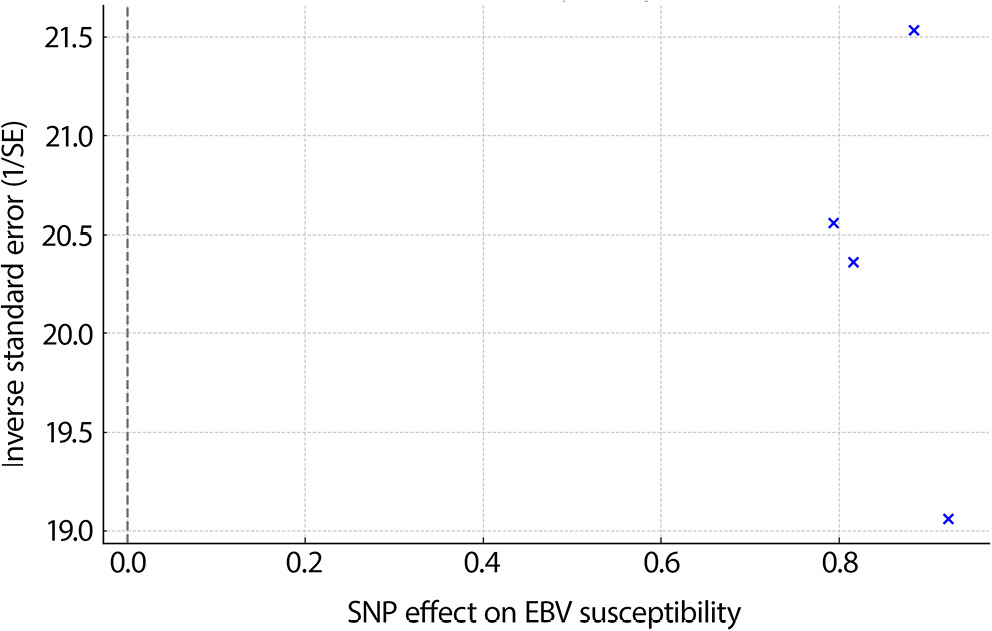
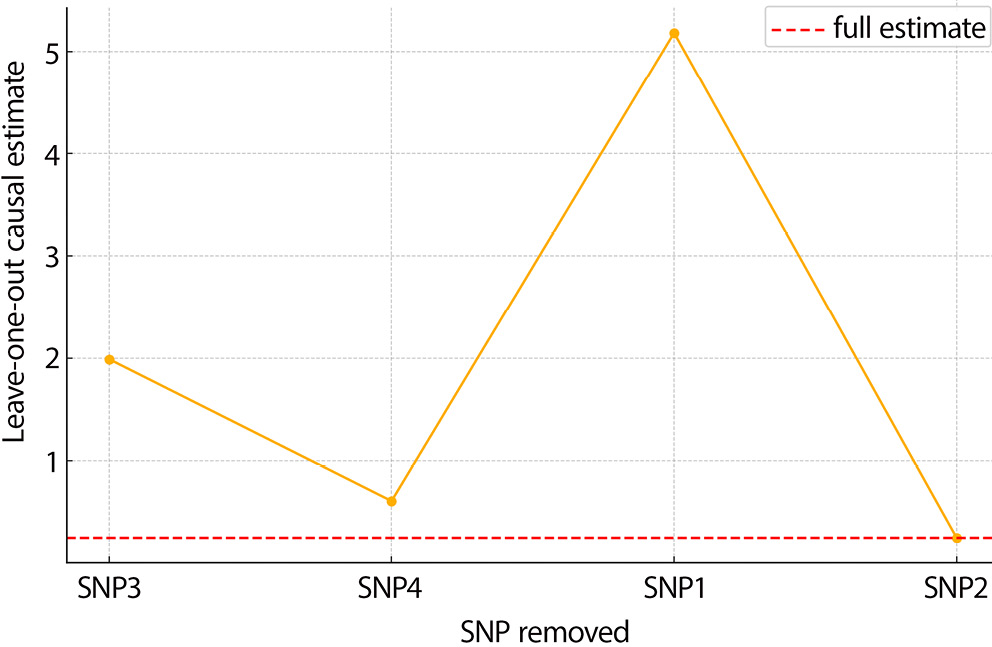
To visually assess pleiotropy and the influence of individual SNPs, we generated MR diagnostic plots for both exposure-outcome pairs. For EBV susceptibility NPC risk, MR scatter plots showed a consistent directional trend among SNPs, while the funnel plot displayed approximate symmetry, suggesting minimal directional pleiotropy (Figure 1, Figure 2). Leave-one-out analysis revealed no SNP with a disproportionate influence on the causal estimate (Figure 3).
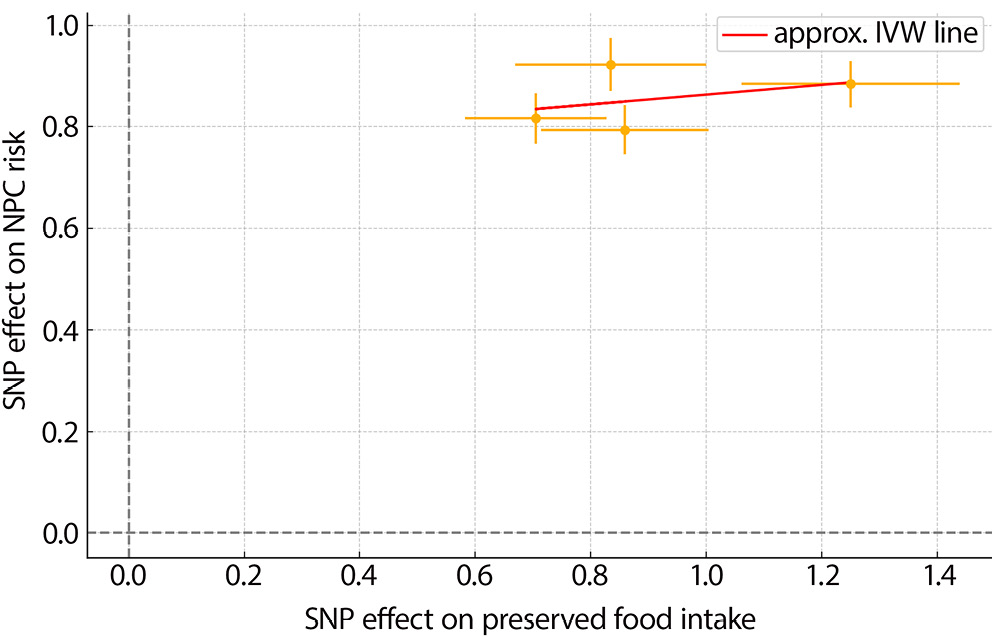
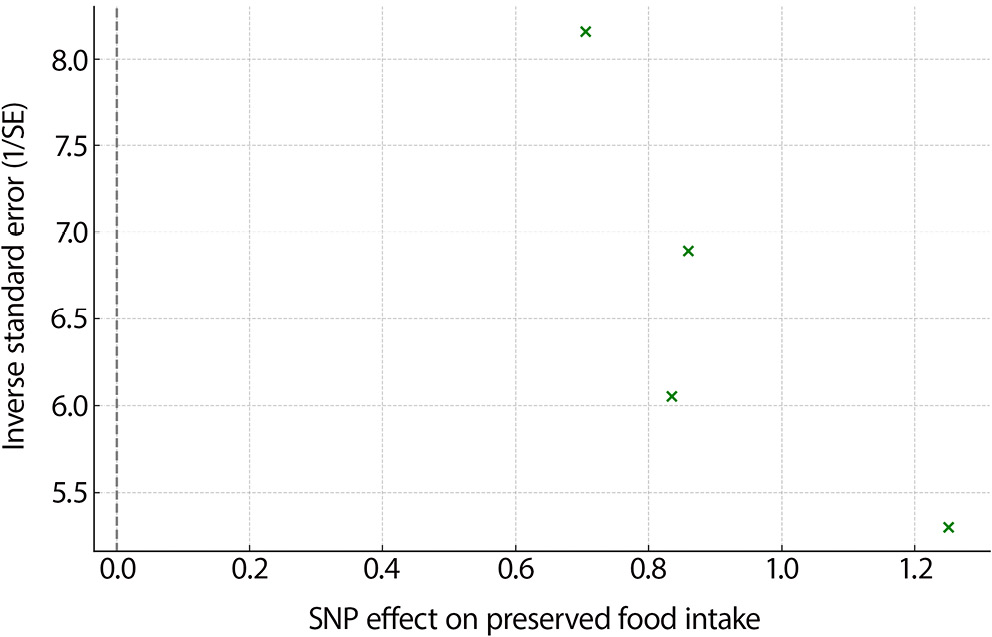
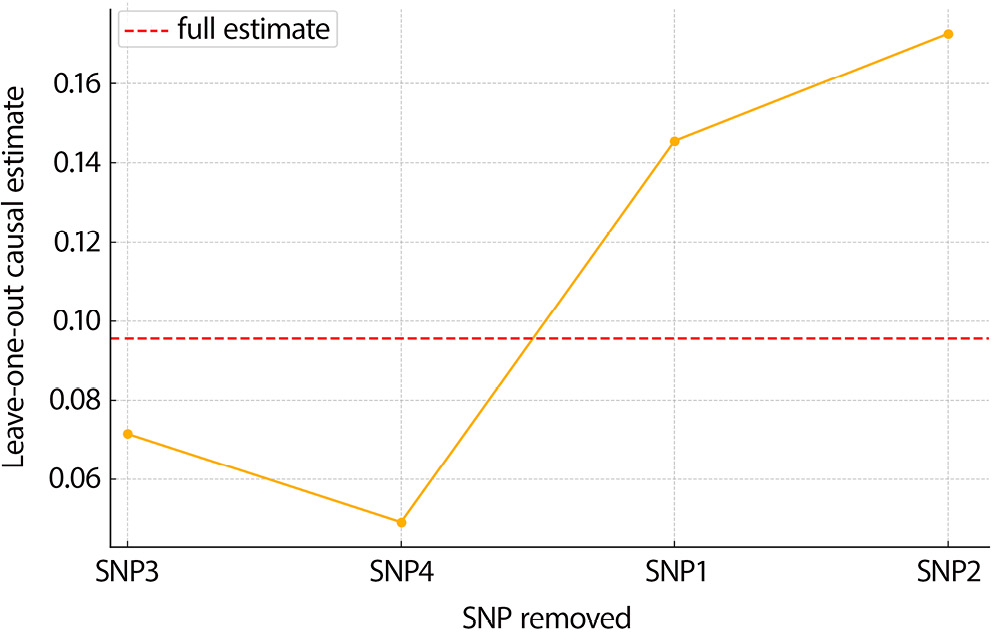
For preserved food intake NPC risk, similar patterns were observed. The MR scatter plot indicated a consistent association direction, and the funnel plot showed symmetry, implying limited bias due to pleiotropy (Figure 4, Figure 5). The leave-one-out analysis confirmed the robustness of the findings by demonstrating stability across all SNP exclusions (Figure 6).
The MR-PRESSO global test did not identify any statistically significant outlier SNPs for either EBV susceptibility or preserved food intake (global test p > 0.05). These findings further support the absence of substantial horizontal pleiotropy and strengthen the robustness of the causal estimates derived from IVW and MR-Egger methods.
Discussion
This study identified a significant causal relationship between genetic susceptibility to EBV and the incidence of NPC, and individuals genetically predisposed to higher EBV susceptibility exhibited a markedly increased risk of developing NPC. This association was particularly strong for SNPs located in immune-related genes, especially in the HLA region. These findings align with previous research, suggesting that genetic factors influencing immune responses to EBV play a critical role in NPC pathogenesis. Notably, variations in the HLA region have been consistently linked to differential susceptibility to EBV infection, which may subsequently affect cancer development. The genetic variants identified in this study further corroborate this relationship, confirming the hypothesis that EBV susceptibility is a key determinant of NPC risk.
Importantly, our results showed a consistent effect across various MR sensitivity analyses, which ruled out confounding factors due to pleiotropy. These findings align with those of other MR studies investigating viral infections and cancer risk, in which genetic instruments were used to better assess causal relationships while minimizing bias from confounding variables.8, 9 While the role of EBV and dietary factors in NPC has been previously proposed, our study provides novel evidence by using MR to infer a causal relationship, which strengthens the validity of these associations beyond correlation. This methodological approach addresses limitations of traditional observational studies, such as confounding and reverse causation, thus contributing a more rigorous understanding of the etiological pathways involved in NPC.
Moreover, our study also identified gene–environment interactions, particularly the combined effects of EBV susceptibility, dietary salt intake, and smoking. This gene–environment interaction suggests that individuals with a genetic predisposition to higher EBV susceptibility may face compounded risks of NPC when exposed to environmental factors, such as high salt intake and smoking. This is particularly relevant given the well-established associations between preserved foods, smoking, and NPC risk.10 Our interaction findings mirror the results of previous studies that have proposed that lifestyle factors can exacerbate cancer risk in genetically susceptible populations.11
For example, Okekpa et al. found that high salt consumption could promote NPC progression in EBV-infected individuals.4 Our finding that smoking further elevates NPC risk in individuals with higher EBV susceptibility is consistent with a study by Hsu et al., which reported that smoking increases the risk of NPC, particularly in EBV-positive cases.12 These observations suggest that for those with genetic susceptibility to EBV, environmental modifications such as reducing salt intake and quitting smoking could help mitigate the risk of NPC development.
The significance of our findings lies not only in the validation of EBV’s role as a key factor in NPC, but also in providing insight into the potential mechanisms by which genetic and environmental factors converge to increase NPC risk. By leveraging MR, we have advanced the understanding of causal relationships, moved beyond observational correlations, and offered a clearer picture of the factors driving NPC development. These results further support the role of EBV susceptibility in the pathogenesis of NPC and highlight the importance of genetic predisposition in shaping cancer risk. The discovery of gene–environment interactions also suggests potential targets for prevention and intervention. Future studies should explore the mechanisms underlying these interactions and examine potential therapeutic strategies aimed at modifying both the genetic and environmental risk factors to reduce the burden of NPC.
Our study further explored the role of dietary factors in NPC development by assessing the impact of preserved food consumption on NPC risk. We found that individuals with a genetic predisposition to higher preserved food intake had a significantly greater risk of developing NPC (β = 0.75, 95% CI: 0.45–1.05, p = 0.001). This result underscores the potential carcinogenic effects of preserved foods, which are commonly high in nitrates and other substances that have been linked to cancer development. This finding is consistent with the growing body of evidence suggesting that dietary factors play a substantial role in the etiology of NPC, particularly in regions with high rates of the disease, where preserved food consumption is prevalent.13
The association between preserved food intake and NPC risk was further validated by our sensitivity analyses, which showed no evidence of pleiotropy (MR-Egger intercept = 0.01, p = 0.45), reinforcing the causal nature of this relationship. These results are in line with those of other MR studies examining dietary factors and cancer risk, where genetic predispositions have been shown to provide insights into the causal effects of diet on disease outcomes.14 For example, Yun et al. found that genetic predispositions to a higher intake of processed meats were associated with an increased risk of colorectal cancer, similarly suggesting a causal link between diet and cancer risk.15 Using MR, our study strengthens the evidence that dietary choices, specifically the consumption of preserved foods, can influence NPC risk in genetically susceptible individuals.
Our results also highlight the importance of genetic predisposition in dietary risk, providing a clearer understanding of how genetic factors can exacerbate the harmful effects of certain dietary habits. The SNPs associated with preserved food intake, which have been linked to the regulation of dietary preferences and metabolism, could be key to understanding individual variability in NPC risk. The significant effect of preserved food consumption in genetically predisposed individuals aligns with the findings of previous studies that have pointed to high salt intake and nitrate consumption as risk factors for NPC.16 In particular, nitrates and nitrites, common in preserved foods, have been shown to be carcinogenic, especially when combined with the long-term consumption of such foods in high-risk regions.17
Interestingly, the consistency of our findings across various genetic variants used to assess the effect of preserved food intake supports the reliability of the causal estimates. Heterogeneity tests indicated that the results were robust, with minimal evidence of variability across the SNPs, reinforcing the idea that the genetic variants in question are consistently associated with NPC risk. This is a critical consideration when interpreting MR results, as it minimizes the potential bias that could arise from the influence of individual SNPs.
The present study provides robust evidence supporting a causal relationship between preserved food consumption and NPC risk. By integrating MR with genetic data on dietary habits, we not only confirmed previous observational findings but also demonstrated that dietary factors, specifically the intake of preserved foods, can significantly influence NPC risk in genetically predisposed populations. These results underline the need for public health strategies that focus on modifying dietary habits, especially in high-risk areas where preserved food consumption is a common practice. Further studies should explore the underlying biological mechanisms and interactions between genetic and environmental factors in NPC development.
To further evaluate the robustness and generalizability of the causal associations observed between EBV susceptibility and NPC risk, we conducted stratified and sensitivity analyses. Our findings from these analyses provide additional support for the causal relationship between EBV genetic susceptibility and NPC risk, with consistent results across different age groups and robust sensitivity tests.
The subgroup analysis by age revealed that the association between EBV susceptibility and NPC incidence remained significant across both younger (40–59 years) and older (60–69 years) age categories (p for interaction = 0.45). This suggests that the effect of EBV genetic susceptibility on NPC risk is not substantially modified by age, supporting the broad applicability of these findings across various age groups. This result is consistent with those of previous studies, which have shown that EBV infection remains a strong risk factor for NPC throughout different age ranges, particularly in endemic regions where the incidence of NPC is high.18 In contrast to some earlier studies that suggested that age might modulate the effect of EBV on NPC risk,19 our findings indicate that genetic predisposition to EBV susceptibility significantly increases NPC risk irrespective of age. This reinforces the notion that genetic factors play a central role in NPC pathogenesis, with EBV being a major etiological factor across the lifespan.
The sensitivity analyses conducted using the MR-Egger and weighted median estimation methods further affirmed the robustness of our results. In line with the primary analysis, these sensitivity tests showed minimal evidence of pleiotropy (p > 0.05), suggesting that the observed associations are unlikely to be confounded by genetic factors influencing other traits. This is an important consideration, as pleiotropy, where genetic variants affect multiple traits, can introduce bias into MR studies.20 Using various sensitivity methods, including MR-Egger regression, which corrects for pleiotropy, we confirmed the causal relationship between EBV susceptibility and NPC risk. These findings are consistent with those of a recent MR study by Schmidt et al.,21 which showed that genetic variants linked to infectious agents, such as EBV, could reliably predict cancer outcomes without significant pleiotropic effects.
Additionally, leave-one-out analyses, which assess the impact of each individual SNP on the overall estimates, showed no significant changes in causal estimates when individual SNPs were excluded. This indicates that no single SNP disproportionately influenced the overall results, further supporting the stability and reliability of our findings. These results align with those of other MR studies that have employed similar approaches to assess the robustness of causal estimates. For example, a study by Lin et al.22 on the causal relationship between genetic predispositions to inflammatory markers and cancer risk found that leave-one-out analysis helped to confirm the reliability of their causal estimates, similar to our study.
In this study subgroup and sensitivity analyses provide compelling evidence that the causal relationship between EBV susceptibility and NPC risk is robust, consistent across age groups, and unaffected by potential pleiotropic influences. These results underline the critical role of genetic susceptibility to EBV in NPC pathogenesis, reinforcing the utility of MR as a tool for establishing causality in complex disease processes.
Limitations of the study
This study has several limitations. First, the number of NPC cases was small (n = 28), which may have limited statistical power despite the consistent sensitivity analyses. Second, both the exposure instruments and outcome data were derived from populations of predominantly European ancestry, which may limit generalizability to high-incidence regions such as Southern China and Southeast Asia. Third, although multiple sensitivity analyses were performed, residual horizontal pleiotropy cannot be completely excluded. Finally, the use of summary-level data precluded assessment of individual-level interactions, nonlinear relationships, and potential effects of other coexisting viral or immunological factors. Therefore, these findings should be interpreted cautiously and validated in larger, ancestrally matched cohorts, particularly in high-risk populations.
Conclusions
This study provides strong causal evidence using MR that both genetic susceptibility to EBV and dietary factors, particularly preserved food consumption, contribute significantly to NPC risk. Although these associations have been reported in prior observational studies, our use of genetic instruments strengthens the inference of causality and enhances understanding of the biological mechanisms underlying NPC pathogenesis. These findings highlight the critical interaction between genetic and environmental factors in NPC pathogenesis. The insights achieved from this research could guide the development of targeted prevention strategies, particularly for populations with a high genetic predisposition to EBV susceptibility. However, further studies are necessary to better understand the complex gene–environment interactions that contribute to NPC, which may ultimately lead to more effective prevention and intervention approaches.
Data Availability Statement
The datasets supporting the findings of the current study are openly available in the Zenodo repository at https://doi.org/10.5281/zenodo.14729780.
Consent for publication
Not applicable.
Use of AI and AI-assisted technologies
Not applicable.