Abstract
Alzheimer’s disease (AD) is the leading cause of dementia in the aging population. Pathogenic processes related to the accumulation of amyloid plaques (Aβ) and intracellular neurofibrillary tangles (NFTs) begin during the asymptomatic stage long before the onset of deterioration in cognitive functions and neurodegeneration, which makes rapid diagnosis and treatment difficult. Although biochemical diagnostic markers isolated from the body fluids of AD patients are currently used, scientists are engaged in research into molecular biomarkers that will significantly accelerate the diagnosis long before the first clinical symptoms appear. The research presented here focused on microRNAs (miRNAs), small, non-coding RNA molecules that are involved in the regulation of the post-transcriptional expression of many genes. A review of the literature revealed that miRNAs play an important role in regulating the expression of genes involved in the pathophysiological mechanisms of AD. Changes in the levels of miRNAs in a patient’s body fluids can be used for rapid diagnosis.
Original scientific articles published between 2014 and 2023 describing clinical and experimental studies on the role and expression levels of various miRNAs were selected from scientific databases such as PubMed, NCBI, Science Direct, and Google Scholar. The selected miRNAs were divided into 2 groups based on their expression level in AD: those with increased expression and those with decreased expression. A review of the latest scientific reports confirms that miRNAs may be a promising source of non-invasive and widely available biomarkers. Additionally, their modulation may prove to be an effective therapeutic strategy in AD.
Key words: Alzheimer’s disease, microRNA, expression, downregulated, upregulated
Introduction
Alzheimer’s disease (AD) is a progressive and incurable neurodegenerative disease that accounts for approx. 70% of dementia cases among the aging population worldwide.1, 2 Individuals with AD may experience behavioral changes, visual disturbances, as well as visuospatial and perceptual problems. They also experience a progressive loss of recent memory, while older memories remain intact.3
The incidence of dementia increases with age, with up to a 15-fold increase observed between the ages of 60 and 85.4 Additionally, gender plays a role, with 2/3 of AD patients being women due to their longer life expectancy and greater genetic risk. Even 1 copy of the APOE-ε4 allele is enough to increase the risk of AD in women.5
Alzheimer’s disease can be classified as familial early-onset AD (EOAD, approx. 5%), which has a genetic basis associated with autosomal mutations in the APP, PSEN1 and PSEN2 genes.6, 7 Around 95% of cases of AD are late-onset AD (LOAD), which is sporadic, idiopathic and of unknown origin. It is associated with a polymorphism in the APOE gene, which encodes apolipoprotein E (APOE).8 The presence of a single APOE ε4 allele increases the likelihood of AD in women by 4 times, while the APOE ε4 homozygote increases the risk by several times in both women and men.5 The clinical pathogenesis of AD is caused by the accumulation of extracellular neuritic plaques (amyloid) in the brain due to the accumulation of amyloid β (Aβ) peptides and intracellular neurofibrillary tangles (NFTs), mainly containing highly phosphorylated tau proteins.9
The Aβ peptides are produced by the proteolysis of the amyloid precursor protein (APP), which plays an important role in brain homeostasis. However, in individuals with AD, the Aβ peptide is produced through the amyloidogenic pathway, which is mediated by β-secretase (BACE1) and γ-secretase (presenilin 1 and presenilin 2). This pathway produces 2 peptides: one consisting of 40 amino acids (Aβ 1–40) and another consisting of 42 amino acids (Aβ 1–42).10, 11
Hyperphosphorylation of tau proteins leads to the development of Alzheimer’s-type neuronal fibrous degeneration (NFT). This reduces the number of synaptic connections between nerve cells, leading to neurodegeneration. There is a correlation between synaptic density and cognitive abilities at the different stages of AD.12
Identifying patients with mild cognitive impairment (MCI) who may progress to AD is crucial. Between 12% and 20% of MCI patients develop dementia annually, while others remain stable for many years or even recover from MCI.13 Therefore, it is important to implement therapy in the initial phases of the disease to increase the likelihood of successful treatment and preservation of cognitive functions. However, detecting individuals in the prodromal stage of AD is challenging as current diagnostic techniques cannot accurately differentiate between the early stages of AD and age-related cognitive deficits and are impractical for routine use.14
Currently, positron emission tomography (PET) and cerebrospinal fluid (CSF) biochemical markers are used to diagnose AD. However, due to their invasive nature and cost, these tests are not routine. Furthermore, they are unable to detect the disease before clinical symptoms appear. Therefore, scientists are researching molecular biomarkers that could diagnose AD at an early stage.15
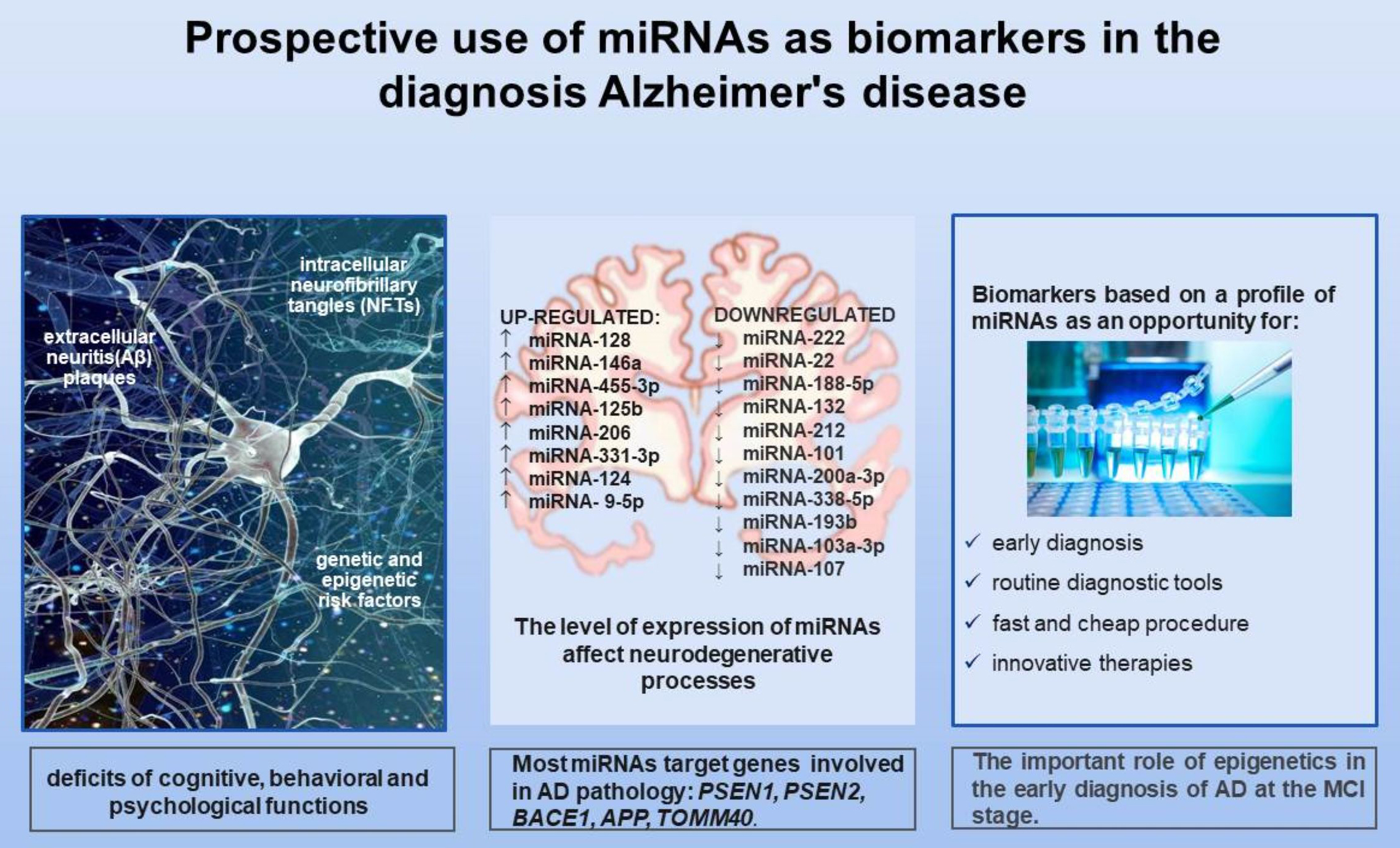
The expression of most genes is regulated at the post-transcriptional level by small, non-coding RNA microRNAs (miRNAs). Approximately 70% of known miRNAs are expressed in the brain. Therefore, their alteration and abnormal regulation may be involved in the etiopathology of AD. Numerous studies have shown that miRNAs not only affect the expression of genes with key importance in AD, such as APP, BACE 1, PSEN 1, PSEN 2, MAPT, and APOE, but also proteins associated with synapses, whose functional changes lead to synaptic dysfunction.4, 16 This article describes a selection of miRNAs with altered expression levels in AD (Table 1). 7, 15, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 45, 46, 47, 48, 49, 50 The research presented aimed to elucidate the role of miRNAs in the mechanisms involved in neurodegenerative processes and their potential use as biomarkers in AD. The scientific databases PubMed, NCBI, Science Direct, and Google Scholar were used to search the literature. Articles were identified through a keyword search using terms such as “Alzheimer’s disease AND miRNAs’’, “microRNAs AND biomarkers”, “β-amyloid AND miRNAs”, “tau protein AND miRNAs”, “synaptic connections AND miRNAs”, and “AD diagnostics AND miRNAs”. The selection process involved the removal of duplicates, the elimination of articles based on their titles, and a review of the abstracts. The next limiting factor was the publication date and type of article. The authors selected only original articles published between 2014 and 2023. The selected miRNAs were tested on transgenic mice, cell lines, and human clinical material, including brain tissue, CSF, and the peripheral blood of patients with AD. The literature was also analyzed in terms of the regulation of genes related to Aβ metabolism, tau proteins, neuronal plasticity, synaptic deficits, and anti-inflammatory signaling by miRNA. A total of 84 articles met the adopted criteria. As a result of a comprehensive analysis, 20 miRNAs were selected and described, the research results of which were presented in at least 2 original articles. The selected miRNAs were classified into 2 groups according to their level of expression, namely those that downregulated and upregulated expression.
Objectives
The objective of this review was to demonstrate the significant role of miRNAs in the diagnosis of AD based on changes in their expression levels. The article presents potential biomarkers based on selected miRNAs involved in the pathogenesis of AD.
Diagnostic markers in AD
The diagnosis of AD is primarily based on neuropsychological evaluation, CSF analysis or PET to determine pathological biomarkers.25 Biomarkers of body fluids, such as Aβ, total tau (t-tau) and phosphorylated tau (p-tau), as well as PET imaging, may indicate synaptic dysfunction in the brain. Biochemical markers are frequently used in research. However, CSF and PET examinations are not commonly used due to their invasive nature and high cost.51 Therefore, researchers have been studying the use of blood biomarkers for many years.
Biomarkers can be classified into 3 categories: those related to amyloid pathology in the brain (Aβ1–40 and Aβ1–42), those determining the degree of neurodegeneration (total tau protein (t-tau) and neurofilament light chain (NfL) and those indicating tau protein pathology (p-tau181, p-tau 231 and p-tau 217).52 The above blood biomarkers for AD are summarized in Table 2.52, 53, 54, 55, 56
Blood marker tests show promise for diagnosing AD; however, the problem of early diagnosis persists. It is important to note that elevated levels of p-tau181, p-tau217 or p-tau231 are correlated with amyloid pathologies and NFT formation in the brain. The most optimal biomarkers are those involved in many regulatory brain processes and neuropathological signaling. These biomarkers can diagnose AD even before any neuropathology develops.
MiRNAs are potential biomarkers that can reflect changes in cellular homeostasis and indicate the presence of multiple pathologies.57 Research has shown that levels of individual miRNAs vary during the development and differentiation of neurons in the human brain and in the aging of the central nervous system (CNS), with these changes being more prominent in AD than in MCI.3, 58
In their meta-analysis, Moradifard et al. examined and analyzed miRNAs involved in AD, their target genes, mRNA-miRNA interactions between them, and signaling pathways. They compared a microarray dataset with expression profiles of miRNAs, from different areas of the brain. Moradifard et al. found that in AD, numerous miRNAs show differential expression, either decreased or increased. Furthermore, they identified associations between individual genes and AD-related pathways, including interactions with the ECM receptor pathway and CAM cell adhesion molecules. These pathways are crucial for neuronal development, synaptic activity, synaptic connection formation, and blood–brain barrier integrity.9
A family of miRNAs implicated in the pathology of dementia outlines the molecular genetics and epigenetics of AD. The mirRNAs have great potential due to their stability in extracellular environments, reactivity and ease of quantification using techniques such as real-time quantitative polymerase chain reaction (RT-qPCR). They are also resistant to the effects of thawing and freezing cycles, making them ideal for use in biomarker studies.15 The significant changes in miRNA expression during diseases offer great potential for clinical diagnosis using miRNA signatures. Efforts are being made to create miRNA signatures that can diagnose different diseases, identify cancers, and predict therapeutic benefits and drug sensitivity.59
Levels of miRNAs may differ in patients at different stages of AD. Therefore, miRNAs have great potential as diagnostic and prognostic biomarkers. They have an advantage over the currently used biochemical markers Aβ1–40, Aβ1–42, t-tau, p-tau181, p-tau231, p-tau217, and NfL because they allow the detection of the disease at a very early stage, even before the appearance of pathological changes in the brain and clinical symptoms in various body fluids such as blood and serum. They can be obtained in a non-invasive way, making the procedure safer and cheaper than collecting CSF.
miRNAs and their role in gene expression
The miRNAs are small non-coding RNA molecules (snRNA) of 21–25 nucleotides that are involved in the post-transcriptional regulation of gene expression in eukaryotes.13 These single-stranded nucleic acids are classified as either RNA polymerase II or III (RNA Pol II, RNA Pol III). The mechanism by which miRNAs regulate gene expression is base pairing with the 3’ untranslated region (3’-UTR). The degree of complementarity within the RNA-induced silencing complex (RISC) determines whether it will lead to inhibition of the target mRNA translation or its degradation. Therefore, overexpression of mammalian miRNAs mainly works to reduce target mRNA levels and lower the miRNA expression to increase mRNA expression levels.22, 60
The miRNAs originate from long transcripts with a double-stranded structure known as primary miRNAs (pri-miRNAs). Pri-miRNAs are transcribed by RNA polymerase II, to which they have a special affinity. In the cell’s nucleus, pri-miRNA is recognized by the Pasha protein, a partner of the Drosha type III RNAase.11 Pri-miRNA undergoes 2 stages of endonuclease processing before it becomes a mature, active miRNA. The first stage involves the Drosha enzyme, whose activity depends on the presence of the RNA-binding protein DGCR8.59
Next, the pre-miRNA is exported from the cell nucleus to the cytoplasm by Exportin 5, which interacts with the RAN protein. In the cytoplasm, RNase III (Dicer) cleaves the pre-miRNA, which is associated with the RNA-binding protein TRBP, to produce mature miRNA. Following cleavage, 1 strand of the miRNA molecule is incorporated into the RISC, while the other strand is degraded.10, 61
Mature miRNAs that are embedded in RISC complexes bind to the 3’-UTR of the target gene RNA. The composition of this protein complex is not yet fully understood. However, it contains the key protein Argonaute, which binds mature miRNAs and searches for target mRNAs, causing translational repression. This results in a decrease in the expression of the gene encoded by the mRNA.10, 59, 61
Since the identification of the first miRNA in 1993, it has been estimated that at least 1% of the human genome is encoded by miRNAs. Each miRNA is capable of regulating up to 200 mRNAs.62 One miRNA can target many genes, and 1 gene can be regulated by many different miRNAs. The miRNAs are a potential tool to study multifactorial diseases, e.g., neurodegenerative diseases, including AD.11
These molecules are involved in various biological processes, including growth, differentiation, regulation of the cell cycle, and metabolic cycles at the cellular level. In the nervous system, miRNAs regulate the proliferation, differentiation and apoptosis of nerve cells at various stages of development and also play a crucial role in memory formation.31 MiRNAs found in neurons are associated with polyribosomes that affect protein expression, suggesting that they play a role in neural tissues and brain growth and development. These findings indicate that the dysregulation of miRNA expression may be associated with several neurodegenerative processes.63
MiRNAs involved in AD pathogenesis
The miRNAs regulate the expression of genes associated with AD development. Research is underway on many miRNAs that regulate the expression of genes such as APP, BACE1, PSEN 1, PSEN 2, ROCK1, PTPN1, and others, which are responsible for neurodegenerative processes such as the accumulation of Aβ in the brain, excessive phosphorylation of the tau protein, and induction of inflammation or apoptosis. Research is also being carried out on miRNAs that regulate genes involved in synapse function (e.g., CPEB1 and BDNF).
Several miRNAs participate in the amyloidogenic pathway that results in the creation of harmful forms of insoluble Aβ. Many miRNAs target the 3’-UTR of APP mRNA. Overexpression of APP stimulates the production and accumulation of Aβ, which causes impaired neuronal activity, synaptic disorders, and ultimately, dementia.10 Patel et al. found that overexpression of the miR-106a/520c reduced APP levels.14 In addition, low levels of miRNA-101a-3p were shown in AD, and transfection of HEK-293 T cells with miRNA-101a-3p mimics and regulates autophagy in AD pathogenesis by targeting the MAPK1 pathway. These studies suggest that miRNA-101a-3p may play an important role in future therapeutic activities.64 Hébert et al. confirmed that overexpression of miRNA-106, miRNA-20a and miRNA-17-5a inhibits APP expression, which is consistent with similar research findings.65 Liu et al. showed overexpression of miRNA-200b, miRNA-135a and miRNA-429 in their studies on APP/PSEN122 transgenic mice. An additional experiment was performed in primary hippocampal neurons and SH-SY5Y cells to confirm the effect of miRNAs on APP and BACE1 expression. The study confirmed that APP expression was inhibited in cells transfected with miRNA-200b and miRNA-429, while BACE1 expression was suppressed by the overexpression of miRNA-135a.66
BACE1 is involved in Aβ formation, and its expression is regulated by several miRNAs. Among them, overexpression of miRNA-29 leads to reduced BACE1 protein levels.67 Similarly, miRNA-135b and miRNA-195a have shown a negative correlation with BACE1 protein levels, which results in lower Aβ levels. Zhang et al. conducted a study in which they transfected mouse hippocampal cells with miRNA-135b inhibitors and miNA-135b mimickers. The study found that overexpression of miRNA-135b improved learning in the maze test, confirming its protective role.68 Rat neurons were transfected with a miRNA-195 mimetic to study its effect on the expression of APP and BACE1. The results showed that miRNA-195 effectively inhibited the expression of both APP and BACE1 by almost 60% compared to the control group.69
MiRNA-34a is involved in the clearance of Aβ, which accumulates in the brain and leads to the formation of amyloid plaques. Studies have shown that miRNA-34a is overexpressed in microglial cells of people with sporadic AD. This microRNA targets the amyloid receptor TREM2 on the surface of microglial cells, which is responsible for Aβ phagocytosis in the CNS. Significant TREM2 deficiencies have been reported in the inflammatory neurodegeneration of AD.70
The process of tau protein phosphorylation and dephosphorylation is regulated by miRNAs. In a study by Wang et al., miRNA-138 was found to be overexpressed in AD and to promote tau phosphorylation in a mouse cell line model. The RARA gene is a direct target of miRNA-138. Downregulation of miRNA-138 increases RARA expression and attenuates the glycogen synthase-3β kinase activity of GSK-3b, thereby inhibiting tau protein phosphorylation.71
MiRNA-146a affects tau phosphorylation by post-transcriptional repression of ROCK1. Inhibition of miRNA-146a expression using antagomir reverses phosphorylation of key signaling pathway components (ROCK1-PTEN-Tau) in the brain.18
Several studies suggest that miRNAs are present in both neuronal axons and dendrites, indicating their potential role in synaptic functions. Synaptic miRNAs play important roles in various aspects of synaptic activity, including synaptic development. These include miRNA-134, miRNA-214, miRNA-188-5p, miRNA-138, miRNA-153, miRNA-124, miRNA-9-3p, miRNA-34, miRNA-125b, and miRNA-132, which are involved in synaptic plasticity, synaptogenesis, synaptic morphology, and synaptic excitability.10, 11, 72
miRNAs upregulated in AD
The changes in the levels of miRNAs in body fluids of individuals with AD and the greater stability of miRNAs compared to mRNA have increased the interest of scientists in these molecules as potential diagnostic biomarkers. Many miRNAs target several genes directly associated with neurodegenerative processes in AD, including PSEN1, PSEN2, BACE-1, APP, TOMM40, and BDNF. Some miRNAs are overexpressed, while others are suppressed due to their protective or inducing roles in AD.62 It is currently believed that during the progression from MCI to AD, there is a change in gene expression direction, from predominantly overexpression in MCI to predominantly underexpression in AD, and this change in gene expression is an important factor in AD development.73
The following are the results of scientific research on 8 miRNAs, whose expression levels increased during AD.
miRNA-128
It was observed that some miRNAs show an increased expression in patients compared to control groups. Although earlier studies on miR-128 focused mainly on its role in tumors of the CNS, Zhang et al. directed their interest to miRNA-128 and the role of this molecule in the AD. The results of his research indicate a significantly elevated level of miRNA-128 in the serum of patients with AD when contrasted with the control group. As part of the study, they analyzed the expression of miRNA-128 and determined its diagnostic utility. A positive correlation was demonstrated between serum miRNA-128 levels and inflammatory factors interleukin (IL)-1β and tumor necrosis factor alpha (TNF-α).17
miRNA-146a
Increased expression of miRNA-146a was demonstrated in areas of the brain most impacted by tau protein pathology, specifically the hippocampus and temporal cortex. In AD patients, miRNA-146a is involved in tau hyperphosphorylation and the pathogenesis of AD. MiRNA146a regulates ROCK1 mRNA in neural cells and directly targets the protein kinase ROCK1. Inhibition of ROCK1 may contribute to lower levels of PTEN phosphatase phosphorylation and induce aberrant tau phosphorylation. Moreover, in a transgenic mouse model of AD called 5xFAD, 3 mutations (2 in the PSEN1 gene and 1 in the APP gene) inhibit miRNA-146a and led to increased ROCK1 protein levels and repression of tau protein hyperphosphorylation, partially restoring memory functions. The overexpression of miRNA-146a in the SH-SY5Y human neuroblastoma cell line was found to significantly increase tau phosphorylation and inhibit ROCK1 protein translation, leading to the disruption of neuronal microtubules and the cytoskeleton. PTEN phosphatase is known to dephosphorylate tau, and the loss of PTEN function has been associated with neurodegeneration mediated by tau hyperphosphorylation and neurofibrillary tangle formation.18
Huang et al.’s research indicates significantly higher levels of miRNA-146a in the peripheral blood of AD patients compared to a control group of healthy individuals. However, a negative correlation was found between the level of miRNA-146a and the concentration of Aβ1–42, as well as the results of the MMSE test in AD patients. Additionally, tau protein levels were positively correlated with miRNA-146a.19 Maffioletti et al. obtained different results in their comparative studies. No significant differences were found when comparing the level of miRNA-146a in the plasma of both AD patients and the control group. However, studies have shown that the level of miRNA-146a increases with age. Additionally, a gender-based relationship was detected, with lower levels of miRNA-146a observed in women compared to men.74
miRNA-455-3p
Although numerous studies on miRNA-455-3p indicate that it is involved in various human diseases, especially considering colon, prostate, liver, gastric, and small cell lung cancers, miRNA-455-3p may also play a role of a potential biomarker in neurodegenerative diseases. Kumar et al. showed a higher expression of miRNA-455-3p in the serum of patients with familial and sporadic AD compared to healthy control. They conducted their study on post-mortem brains of individuals with AD and healthy individuals. They also extended their research by analyzing the fibroblasts and B cells of patients with familial and sporadic AD, as well as an age-matched control group. The levels of miRNA-455-3p were found to be increased in fibroblasts from patients with familial AD and in fibroblasts and B cells from patients with sporadic AD when compared to healthy controls. MiRNA-455-3p expression has been shown to increase in serum samples, cell lines, post-mortem brain models, and mouse models of AD.20
In a separate analysis, Kumar et al. investigated the molecular targets of miRNA-455-3p that affect AD development. They identified a correlation between miRNA-455-3p and multiple signaling pathways and their corresponding genes. The pathogenesis of AD is directly related to several signaling pathways and their respective genes, including extracellular matrix (ECM)-receptor interactions, adherens junctions, transforming growth factor beta (TGF-β) signaling pathways, and actin cytoskeleton regulation. These pathways involve genes such as THBS1, COL3A1, HSPG2, COL6A1, RUXN1, MYC, Smad2, PLK1, and TNC.75
Xiao et al. demonstrated that miRNA-455-5p targets the 3’UTR of the CPEB1 gene, which plays a crucial role in protein synthesis in neurons. Higher levels of miRNA-455-5p inhibit the translation of CPEB1, leading to a decline in synaptic plasticity and memory.21
miRNA-125b
Upon analysis of the CSF of AD patients, it was demonstrated that miRNA-125b was overexpressed compared to the control group of healthy individuals. In addition, in vitro studies were conducted on the Neuro2a APPSwe/Δ9 cell line, where miRNA-125b mimetics were transfected, resulting in the overexpression of inflammatory factors TNF-α, IL-1β, IL-6, and IL-10, induction of apoptosis, and inhibition of cell proliferation. Transfected cells demonstrated increased levels of APP proteins, Aβ peptides and p-ERK proteins. The overexpression of miR-125b significantly suppressed the expression of sphingosine kinase 1 (SphK1) proteins, which affects cell death and survival by maintaining the sphingosine-1-phosphate (S1)/ceramide balance.23
Nuclear factor kappa-light-chain-enhancer of activated B cells (NF-kB) is overexpressed in AD tissues and strongly activates pro-inflammatory genes. Pro-inflammatory cytokines and peptides, including IL-1β, TNFα, Aβ42, as well as HSV-1 and aluminum, activate both NF-kB and possibly miRNAs. NF-kB has been shown to activate the transcription of several miRNAs, including miRNA-125b. Overexpression of miRNA-125b silences brain genes related to phagocytosis and neurotropism. Therefore, any biophysical or physiological stressor that activates NF-kB in a cell can also activate miRNA-125b, which is strongly transcriptionally controlled by NF-kB. MiRNA-125 regulates the levels of the enzymes 15-lipoxygenase-15-LOX and neuroprotectin D1 (neuroprotectin-D1-NPD1) derived from docosahexaenoic acid (DHA). A reduction in their levels impairs homeostasis as well as anti-apoptotic and neuroprotective effects of genes that are typically responsible for anti-inflammatory and neuroprotective signaling in brain cells.22
While studies have found elevated miRNA-125b levels in AD cell culture models and patient tissue samples; however, research on miRNA-125b levels in patient plasma is limited. In their study on changes in the expression of several miRNAs in the plasma of AD patients, including miRNA-125b, Vergallo et al. did not find any significant differences in the level of miRNA-125b compared to the control group of healthy individuals.76
miRNA-206
Another miRNA whose increased expression has been observed in the course of AD is miRNA-206. It was tested in the serum using RT-qPCR. In a group of MCI patients at higher risk of developing dementia and with worsening outcomes over 4 years displayed significantly higher levels of miRNA-206 expression in comparison to AD patients. Increased miRNA-206 expression was significantly associated with memory impairment and cognitive decline based on the Mini-Mental State Examination (MMSE). The mechanisms behind increased miRNA-206 expression, and its function in the CNS are not yet understood. However, the miRNA-206 targeting of brain-derived neurotrophic factor (BDNF), histone deacetylase 4 (HDAC4) and Jun D transcription factor has profound effects on the CNS. BDNF is a particularly important target of miRNA-206 due to its functions in maintaining CNS homeostasis, regulating neurotransmission, synaptic plasticity, dendritic branching, and neuronal survival. Individuals with MCI and AD show a decrease in BDNF mRNA and protein levels in their brains. Studies in mouse models of AD, with overexpression of the mutant human APP protein, have confirmed the silencing of BDNF by miRNA-206.15
Moon et al. confirmed the role of miRNA-206 in AD. The study analyzed a group of patients with early dementia and examined the level of miRNA-206 in the olfactory membrane of MCI patients. The results showed that the expression level of miRNA-206 was higher in MCI patients compared to the control group.24
miRNA-331-3p and miRNA-9-5p
Chen et al. conducted an experiment on a mouse model to demonstrate changes in the expression of miRNA-331-3p and miRNA-9-5p during the AD process. The study found that miRNA-331-3p and miRNA-9-5p targeted sequestosome 1 (Sqstm1) and optineurin (Optn) receptors, which influence the selective autophagy pathway. Autophagic activity increased in the early stages of AD during initial Aβ deposition but decreased in the later stages of the disease. Following treatment of SH-S5Y5 cells with inhibitors of miRNA-331-3p or miRNA-9-5p, the levels of Sqstm1 and Optn increased. Therefore, the use of miRNA-331-3p and miRNA-9-5p antagonists may prevent memory loss and mobility impairment in AD. It is possible to distinguish between early and late stages of AD by assessing the expression of miRNA-331-3p and miRNA-9-5p, autophagic activity, and Aβ accumulation.25
To confirm the diagnostic value of miRNA-331-3p, studies were conducted in both AD patients and SH-SY5Y cells. The expression of miRNA-331-3p was significantly reduced in both the patients and the cell line. Correlations were found between miRNA-331-3p expression levels in the patient’s serum and their MMSE scores, as well as pro-inflammatory cytokines. The studies on SH-SY5Y cells confirmed the results, showing that overexpression of miR-331-3p increased cell viability and inhibited inflammatory responses.26
miRNA-124
In their experiment on transgenic mice, Hou et al. demonstrated that the overexpression of miRNA-124 in the hippocampus resulted in reduced levels of non-receptor protein phosphatase 1 (PTPN1), hyperphosphorylation of tau and the formation of insoluble tau protein. To understand the regulatory mechanisms of the miRNA-124/PTPN1 pathway, the researchers examined levels of the R1 transcription factor (REST) under the influence of the stress factor Aβ1–42 oligomer. R1 transcription factor mediates the expression of various classes of ncRNA, including miRNA-124. High levels of Aβ1–42 oligomer in AD patients cause REST silencing in both cortical and hippocampal neurons in the CA1, CA3 and CA4 fields, as well as overexpression of miRNA-124. Therefore, the overexpression of REST suppresses miRNA-124 expression and reverses the decrease in PTPN1.27
Another group of researchers obtained contrasting results regarding miRNA-124 in the brains of patients with sporadic AD. Their research indicates that the level of miRNA-124 is lower compared to individuals from the control group. An et al. demonstrated in their studies on the human neuroblastoma cell line SH-SY5Y that high levels of miRNA-124 inhibit the activity of BACE1, which plays a crucial role in Aβ production. Therefore, miRNA-124 can be used as a BACE1 inhibitor.30
miRNAs downregulated in AD
Many studies on miRNAs as potential biomarkers in AD show differences in the expression of individual classes involved in neurodegenerative processes. Not only does miRNA overexpression indicate the development of the disease, but reduced expression levels also influence changes in the brain and the risk of progressive dementia.
This section describes 12 miRNAs that are downregulated in AD, based on animal studies, in vitro studies and in body fluids from AD patients.
miRNA-222
The group of miRNAs with reduced expression in AD includes miRNA-222. The expression of miRNA-222 was significantly decreased in mild and moderate AD groups compared to the healthy group. Moreover, miRNA-222 expression in the moderate AD group was considerably lower than in the mild AD group.31
Wang et al. demonstrated that miRNA-222 contributes to the development of AD by influencing cell cycle dysfunction and the expression of cyclin-dependent kinase inhibitor 1B (p27kip1). Normally, the expression of p27Kip1 proteins inhibits the phosphorylation of the retinoblastoma protein (pRb), which halts cell proliferation in the G1 phase of the cell cycle. Studies have shown that aberrant expression of cell cycle markers, such as p27Kip1, contributes to the pathogenesis of AD.32
miRNA-22
Han et al. confirmed the important role of miRNA-22 in inhibiting inflammation of microglia characteristic of AD. Compared to healthy controls, AD patients showed a significant decrease in circulating miRNA-22 levels, while the expression levels of pro-inflammatory factors, namely IL-18, IL-1β and TNF-α, were significantly increased in AD patients. The correlation analysis revealed a negative correlation between miRNA-22 and the inflammatory factors IL-18, IL-1β and TNF-α. This suggests that miRNA-22 plays a key role in the development of AD. It prevents pyroptosis and the release of inflammatory factors by regulating the mRNA expression of gasdermin D (GSDMD), a substrate of inflammatory caspases.33
Zhai et al. confirmed the protective role of miRNA-22 in APP/PS1 mice. The study induced inflammation and cell pyroptosis using Aβ, followed by transfection of cells with miRNA-22 mimetic using exosomes. The study demonstrated that miRNA-22 targets the 3’-UTR of the GSDMD gene mRNA, inhibiting GSDMD expression. This leads to a reduction in caspase-1 and inflammatory factors such as IL-1β, IL-6 and TNF-α. Research suggests that exosomes loaded with miRNA-22 can enhance cognitive performance in mice with AD more effectively by inhibiting inflammation.34
miRNA-188-5p
This study found that oligomeric Aβ1–42 reduces the expression of miRNA-188-5p. The researchers examined the expression of miRNA-188-5p in brain tissue of patients with AD and age-matched control subjects. The results showed significantly lower miRNA-188-5p expression in the cortex and hippocampus of AD patients. Furthermore, there was a significant increase in immunoreactivity against the protein neuropilin-2 (Nrp-2), which is the target of miRNA-188-5p. The elevated expression of miRNA-188-5p in 5xFAD transgenic mice restored synaptic transmission and dendritic spine density that was inhibited by oligomeric Aβ1–42.35
miRNA-132 and miRNA-212
Hernandez-Rappi et al. demonstrated that a deficiency of miRNA-132 and miRNA-212 in mice promotes Aβ pathology. Both miRNA-132 and miRNA-212 play an important role in synaptic plasticity, neurite outgrowth and memory formation. The study found that levels of miRNA-132 and miRNA-212 are reduced in individuals with MCI and AD compared to healthy controls. This reduction is associated with memory impairment. The research group demonstrated the effects of miRNA-132/212 deletion on increased tau phosphorylation and accumulation, as well as the production and removal of Aβ, using miRNA-132/212 knockout mice. Additionally, using a transgenic mouse model (3xTg-AD), it was proven that miRNA-132 targets SIRT1, MAPK1/ERK2 and MAPT, the levels of which were increased in 3xTg-AD mice, resulting in increased tau pathology and memory deficits. Reintroducing miRNA-132 and miRNA-212 into the body can partially halt this process.36
Furthermore, Deng et al. conducted studies in rats and found that miRNA132 inhibits the expression of the MAPK1 gene, which is responsible for mitogen-activated protein kinase 1. The MAPK1 serine-threonine kinase pathway is involved in the p38 signaling pathway that enhances inflammation and apoptosis during oxidative stress. Therefore, low expression of miRNA-132 in AD leads to an increase in MAPK levels, resulting in inflammation and neuronal death.37
miRNA-101
To investigate the role of miRNAs in the regulation of APP gene expression, APP protein levels were measured in hippocampal cells from rats with reduced Ago2 expression. Western blot analysis showed that APP levels were significantly higher in neurons with reduced Ago2 levels. Among the miRNAs that potentially target the APP 3’-UTR, miRNA-101 plays a key role. In this cell model, miRNA-101 showed a negative correlation with APP proteins. To determine the role of miRNA-101 in modulating the Ago2/miRNA pathway, hippocampal cells were transfected with specific microRNA inhibitors. The results showed that inhibition of miRNA-101 expression significantly increased APP protein levels, suggesting that APP expression in the hippocampus is regulated by miRNA-101.77
To confirm the regulatory role of miRNA-101 on endogenous APP in human cells, Long et al. performed a study in human HeLa cell line. In addition, the inhibitory effect of miRNA-101 on APP levels was compared not only in the human HeLa cell line but also in the human astrocyte cell lines and rodent neurons. For this purpose, miRNA-101 mimetics were used. These studies showed the strongest expression of miRNA-101 in model CNS neurons.39
miRNA-200a-3p
MiRNA-200a-3p, a member of the miRNA-200 family, is known to play a significant role in human cancers. Recent studies suggest that it may also have an impact on the pathology of AD. Studies in mice (APP/PS1) have shown that miRNA-200a-3p levels are decreased in AD. Similarly, significantly lower levels of miRNA-200a-3p were observed in the plasma of AD patients compared to the healthy control group. Experiments have shown that overexpression of miRNA-200a-3p inhibits the production of Aβ1–42 in APPswe-transfected cells, while a reduction of miRNA-200a-3p leads to overproduction of the toxic form of Aβ. The target of miRNA-200a-3p has been identified as BACE1 mRNA, and the expression level is negatively correlated with miRNA-200a-3p. Additionally, miRNA-200a-3p targets the PRKACB gene, which encodes one of the catalytic subunits of PKA that increases the level of phosphorylated tau. Studies have confirmed that miRNA-200a-3p mimetics significantly decreases the expression of both PRKACB mRNA and protein.40
Another research group suggests that miRNA-200a-3p is involved in Aβ-induced neuronal apoptosis. In studies conducted in the hippocampus of APPswe/PS mice, overexpression of miRNA-200a-3p and reduced levels of SIRT1 were observed. SIRT1 is an anti-apoptotic protein that inhibits neuronal apoptosis. Reporter gene assay confirmed the miRNA-200a-3p binding sites in the 3’-UTR of SIRT1 mRNA. Suppression of miRNA-200a-3p inhibits apoptosis by targeting SIRT1. Therefore, miR-200a-3p may be a potential therapeutic target for the treatment of AD.41
miRNA-338-5p
MiRNA-338-5p is another miRNA that may serve as a biomarker for AD due to its reduced levels. It targets γ-secretase, a key enzyme involved in Aβ1–42 production. To determine the impact of miR-338-5p on AD, its concentration was evaluated in individuals diagnosed with AD and a cohort of healthy participants. Studies have shown a significant decrease in miRNA-338-5p levels in people with AD. As expected, the overexpression of miRNA-338-5p suppresses BACE1 gene expression. Neuroinflammation in the CNS is linked to an increased accumulation of Aβ in AD and plays a crucial role in the development of neuropathology. It results in a reduction of synaptic connections and cognitive abilities. Therefore, overexpression of miRNA-338-5p may alleviate nervous system inflammation and restore synaptic function.42
Li et al. demonstrated the protective effect of miRNA-338-5p on cognitive dysfunction in APP/PS1 transgenic mice, achieved by reducing brain amyloid plaque deposition and delaying apoptotic neuronal loss caused by Aβ40 accumulation.43
miRNA-193b
In the case of miRNA-193b, decreased expression was found in hippocampal neurons of APP/PS1 transgenic mice, while the total level of miRNA-193b was overexpressed in exosomes. Research has indicated that a reduction in miRNA-193b levels in the hippocampus results in an increase in miRNA-193b found in exosomes labeled with a protein from the ATP-binding cassette transporter A1 (ATP-ABCA1). The difference in miRNA-193b expression observed in AD may be due to the body’s effort to compensate for the loss of APP. The precise biological functions of miRNA-193b in AD have yet to be investigated. It is unclear whether ABCA1-tagged exosome signaling has any effects.45 Inhibitory oligonucleotides capable of specifically binding and inhibiting miRNA-193b activity were used to demonstrate that miRNA-193b binding sites are located in the 3’UTR sequence of the APP gene transcript. The miRNA-193b inhibitor significantly increased APP expression compared to the control group.44
miRNA-137
Nong et al. investigated the protective role of miRNA-137 in neurotoxicity of DIV14 rat primary cortical neurons. They confirmed Aβ-induced neurotoxicity by testing for caspase-3 activity, which is closely related to cell apoptosis. Using miRNA-137 mimetics, it was found that the overexpression of miRNA-137 significantly alleviated the neurotoxicity caused by Aβ. Furthermore, they explained the molecular mechanisms responsible for the neuroprotective effect of miRNA-137. The study used a luciferase assay to detect a complementary binding site between miRNA-137 and extracellular signal-regulated kinase 1/2 (ERK1/2). Furthermore, the study found that miRNA-137 overexpression negatively regulates ERK1/2 expression. These findings reveal the crucial role of miRNA-137/ERK1/2 signaling in AD.46
Experimental models of AD, including N2a cells and mouse cortical neurons, have been instrumental in elucidating the molecular mechanisms underlying the influence of miR-137 on Aβ-induced neurotoxicity. Reduced miRNA-137 levels have been observed to increase neurotoxicity, with this effect being attributed to the direct targeting of TNFAIP1 and the subsequent suppression of its mRNA and protein levels. Conversely, elevated miR-137 expression has been demonstrated to mitigate neurotoxicity by targeting TNFAIP1, thereby inactivating the NF-κB pathway.47
miRNA-103a-3p and miRNA-107
Using microarray expression profiling in Gene Expression Omnibus (GEO), Chang et al. confirmed that miRNA-103a-3p and miRNA-107 suppressed cofilin translation. The decreased levels of these miRNAs lead to high levels of cofilin protein in AD. Furthermore, a negative correlation was found between the number of senile plaques and NFTs and the level of miRNA-107 expression in the gray matter of the cerebral cortex during the early stages of AD. The study found that the BACE1 gene is targeted by both miRNA-107 and miRNA-103a-3p. It also revealed that an increase in BACE1 levels in AD patients was accompanied by a decrease in miRNA-107 expression. Profiling has confirmed that miRNA-107 and miRNA-103a-3p significantly contribute to the development of AD by regulating the expression of BACE1, LRP1, CDK5R1, and DLG4.7
Wang et al. confirmed that cases with low levels of miRNA-107 were associated with more severe AD pathology. This is usually characterized by increased BACE1 levels compared to cases with high miRNA-107 levels.48
In their experiments on mouse brains, Shu et al. showed that as Aβ levels increase, miRNA-107 expression decreases. In their study, miRNA-107 mimetics were used to prevent the negative processes induced by increased levels of Aβ. MiRNA-107 protects cells against death in the CA1 region, impairment of synaptic transmission, memory loss, increases in Aβ1–42 and p-tau levels, and depression of the BDNF-TrkB pathway.49
The role of miRNA-107 as an AD biomarker was confirmed through in vivo studies, which examined its level in the plasma of AD patients and a control group. The results showed reduced expression in people with AD. Furthermore, a positive correlation was found between miRNA-107 levels and MMSE test results, and a negative correlation was found with the level of dementia in AD patients.50
Limitations
The primary limitation of this review is the number of miRNAs described. Of the numerous miRNAs involved in the pathogenesis of AD, the authors selected only 20 (8 with increased expression and 12 with decreased expression in AD), a comprehensive listing of which would exceed the scope of this publication. The study is also limited by a lack of knowledge about the mechanism of action of all miRNAs regulating AD-related genes. It is important to remember that AD is a polygenic disease. Experimental models of AD, as well as human post-mortem and in vivo studies, indicate that dysregulation of several miRNAs may influence the pathophysiologic mechanisms of AD, including the Aβ pathway, tau pathology, the brain immune response, and inflammation, including regulation of oxidative stresses.
To ensure objectivity, patient selection must be based on several factors, including age, gender, comorbidities, medications, and stressors. It is also important to remember to select uniform biological materials, use consistent analytical techniques in research, adhere to standardized test methodology, and employ appropriate statistical analyses. Long-term studies are essential to accurately delineate the alterations in the expression levels of individual miRNAs at each stage of the disease.
Conclusions
Recent studies have confirmed that several miRNAs play a key role in the neurodegenerative processes associated with AD. These miRNAs can be identified in the CSF or peripheral blood of patients. The expression levels of miRNA-128, miRNA-455-3p, miRNA-206, miRNA-22, and miRNA-107 have been observed to vary in both experimental and clinical studies in AD patients. The use of different biological materials and analytical techniques may result in inconsistencies in miRNA expression, as evidenced by studies on miRNA-146a and miRNA-125b. Nevertheless, the majority of evidence indicates that miRNAs may be of significant value in the diagnosis and treatment of AD. Currently, there are no comprehensive miRNA profiles available for practical use. A major challenge is to standardize research methods and determine the expression ranges of all AD-related miRNAs. This will allow proper interpretation of blood test results in patients. We hope that miRNA-based research will become a viable method for diagnosing, preventing, delaying, and treating AD.