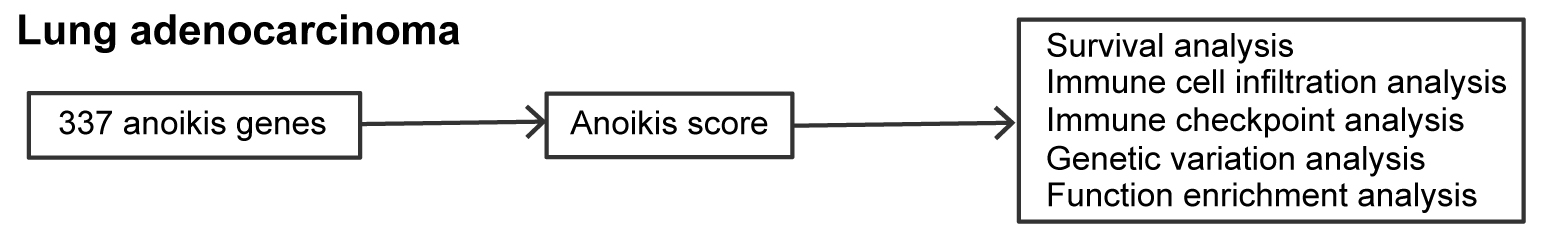
Abstract
Background. Lung adenocarcinoma (LUAD) is the most common subtype of lung cancer, and while advancements in diagnosis, surgery, radiotherapy, and molecular therapy have improved clinical prognosis, the long-term survival rate and quality of life of patients remain unsatisfactory. Therefore, identifying new prognostic biomarkers and potential therapeutic targets is crucial.
Objectives. This study aimed to analyze the role of anoikis-related gene characteristics in LUAD.
Materials and methods. The anoikis-related genes were obtained from the GeneCards database. Genetic expression data and clinical characteristic information were collected from The Cancer Genome Atlas (TCGA)-LUAD, and the Gene Expression Omnibus (GEO) GSE31210, GSE37745, and GSE68465 datasets. Random survival forest and least absolute shrinkage and selection operator (LASSO) models were applied to construct the risk model. An analysis of immune cell infiltration and function was performed with the scores.
Results. Four prognosis-related genes (TLE1, GLI2, PLK1, and BAK1) were obtained and used to construct the anoikis score. We found that the patient survival rate was higher in the low-anoikis score (LAS) group. Moreover, both the stromal and immune scores were negatively correlated with the anoikis score. With the increase of the anoikis score, the levels of natural killer cells, regulatory T cells, CD4+ T cells, CD8+ T cells, B cells, and macrophages decreased. The anoikis score had a negative regulatory relationship with the immune response, natural killer cell activation and T cell activation. The TP53 mutation was significant in LUAD patients and was present in 56% of the high-anoikis score (HAS) group and in 40% of the LAS group.
Conclusions. The anoikis score was associated with poor prognosis in LUAD patients. Anoikis-related genes were associated with tumor immune dysregulation and TP53 mutation in LUAD. This study opens a new perspective for LUAD therapy.
Key words: prognosis, immune, mutation, lung adenocarcinoma (LUAD), anoikis
Background
Lung cancer remains one of the leading causes of cancer-related deaths worldwide1 and its incidence continues to increase.2 The most common subtype of lung cancer is lung adenocarcinoma (LUAD), which accounts for approx. 40% of cases.3 While progress in diagnosis, surgery, radiotherapy, and molecular therapy has resulted in improved clinical outcomes for LUAD patients,4 the overall quality of life and long-term survival rates remain unsatisfactory.5, 6, 7 Therefore, there is a need to identify new prognostic biomarkers and potential therapeutic targets.
Anoikis refers to the type of cell death that is triggered when a cell loses its connection to both surrounding cells and the extracellular matrix (ECM).8 This type of cell death plays a crucial role in maintaining tissue homeostasis and regulating normal development.9 Anoikis is primarily triggered by the interaction between mitochondrial interference and cell surface death receptor activation.10, 11 However, during tumorigenesis and development, cells can develop resistance to anoikis, which is believed to be a key factor in promoting cancer cell metastasis12 and survival under unfavorable conditions.13 Several studies have demonstrated the prognostic significance of anoikis in various diseases, such as glioblastoma (GBM),14 head and neck squamous cell carcinoma (HNSCC)15 and clear cell renal cell carcinoma (ccRCC).16 Additionally, the management of anoikis resistance is a critical factor in cancer development.17, 18 To gain more insight into the role of anoikis in LUAD, we analyzed the genetic signatures related to this biological process.
The rapid advancement of publicly available biological databases has facilitated easy access to vast amounts of bioinformatics data samples related to lung cancer. This makes bioinformatics results with large sample sizes more representative. In addition, with the widespread application of databases and network tools, and the integration of bioinformatics methods, medical research is trending towards utilizing these advancements.19 The least absolute shrinkage and selection operator (LASSO) model is a common machine learning analysis method that has been widely used to select genetic features, due to its ability to effectively select important feature values with non-zero coefficients.20 Multiple studies have utilized bioinformatics analysis, including LASSO analysis, to predict the prognostic value of various genes in a wide array of diseases.21, 22, 23 These studies pave the way for further data mining and validation of related biomolecular targets.
Objectives
Survival curves, immunity and mutation analysis were used to identify the role of anoikis-related genes in LUAD and their potential as prognostic markers.
Materials and methods
Data acquisition
We collected 337 genes related to anoikis from the GeneCards database and standardized them based on a relevance score greater than 1.24 Data related to genetic expression associated with anoikis and clinical characteristics were obtained from The Cancer Genome Atlas (TCGA)-LUAD dataset (502 cases) and the Gene Expression Omnibus (GEO) datasets, including GSE31210 (226 cases), GSE37745 (106 cases) and GSE68465 (442 cases). The TCGA-LUAD data were used as a training set, while the GSE31210, GSE37745 and GSE68465 data served as the test set.
Construction of the risk model
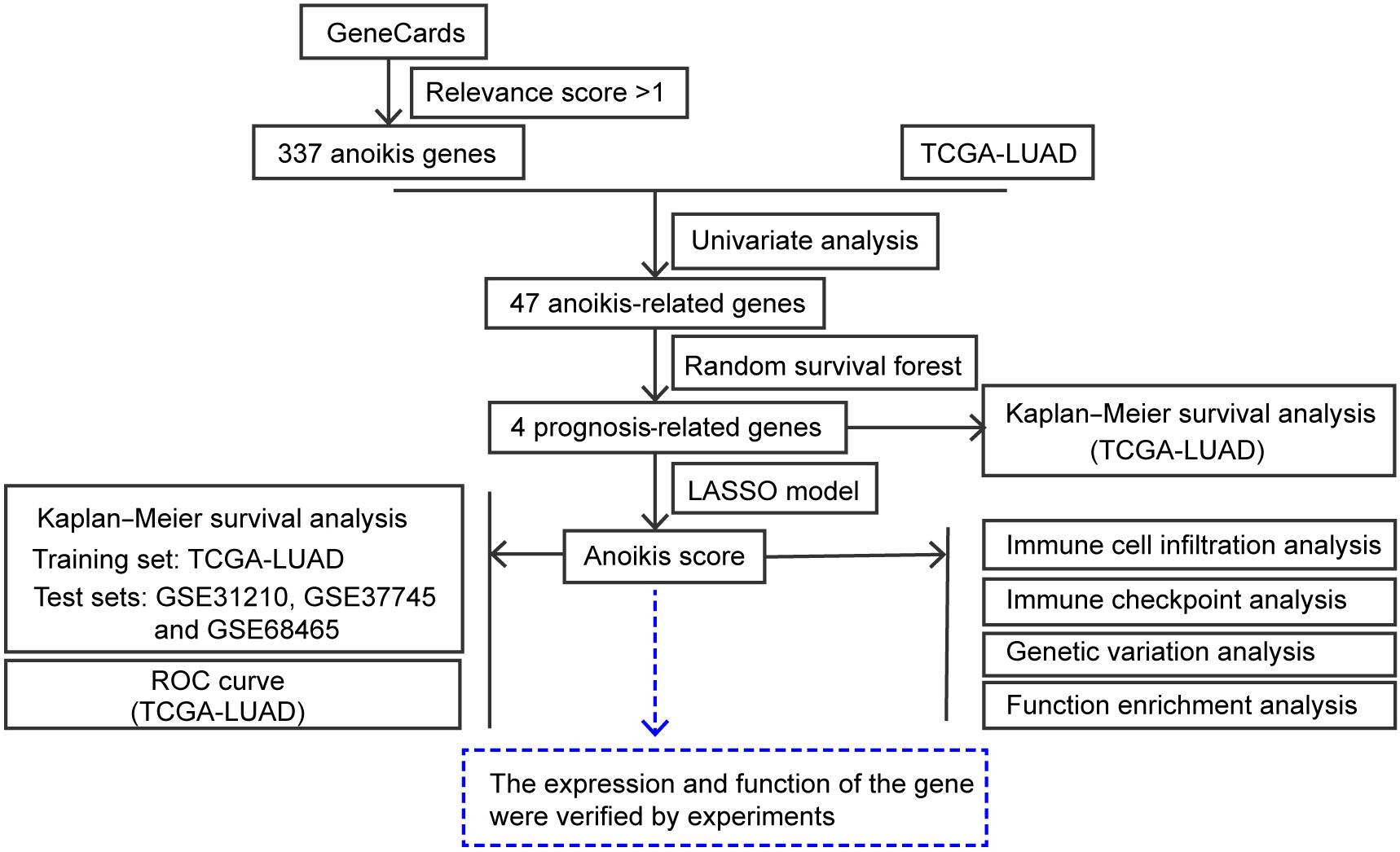
The study involved conducting a univariate analysis between the anoikis genes and TCGA datasets. Afterwards, the anoikis genes were subjected to a random survival forest analysis to identify prognosis-related genes. Further analysis using the LASSO model was performed to construct the prognosis score, also known as the anoikis score (Equation 1):
 (1)
(1)
A diagrammatic representation of the study workflow is shown in Figure 1. The LASSO regression minimized the loss function to obtain coefficient estimates for each explanatory variable. Then, the number of non-zero coefficients was observed to determine the number of variables selected for the risk model.
Immune cell infiltration
and function analysis
The infiltrating immune cells in LUAD patients were assessed using multiple algorithms, including Estimation of STromal and Immune cells in MAlignant Tumours using Expression data (ESTIMATE), Microenvironment Cell Populations-counter (MCPcounter), single sample Gene Set Enrichment Analysis (ssGSEA), and Tumor Immune Estimation Resource (TIMER), to determine the prognostic index. Based on anoikis scoring, the tumor infiltration of the immune cell subpopulations and the expression levels of immune checkpoints in LUAD patients were evaluated using Pearson’s correlation analysis. To gain further insight into the influence of the anoikis score on functional pathways in LUAD, the Gene Ontology (GO) enrichment analysis was performed using GSEA with the clusterProfiler.
Mutation and CNV analysis
The mutation and copy number variation (CNV) analyses were conducted using data from TCGA. The R package maftools were used for mutation analysis. The CNV landscape and the copy number gains or losses at the peaks were evaluated using the GISTIC 2.0 (https://gatk.broadinstitute.org).
Statistical analyses
Data were visualized using the R package “ggplot2”. The patient survival curve was generated using the Kaplan–Meier method. The receiver operating characteristic (ROC) curve was estimated using the R package “pROC”. The likelihood ratio test (univariate analysis), the log-rank test (Kaplan–Meier survival analysis), Pearson’s correlation test (Pearson’s correlation analysis), and Fisher’s exact test (χ2 test) were also performed. For Pearson’s correlation test, the t-statistic was calculated based on the Pearson’s correlation coefficient (cor) transformation in order to determine the p-value according to the degrees of freedom (df) and the level of significance. The Benjamini–Hochberg (BH) method was performed to correct p-values. The value of p < 0.05 was considered statistically significant.
Results
Construction of the anoikis score in LUAD
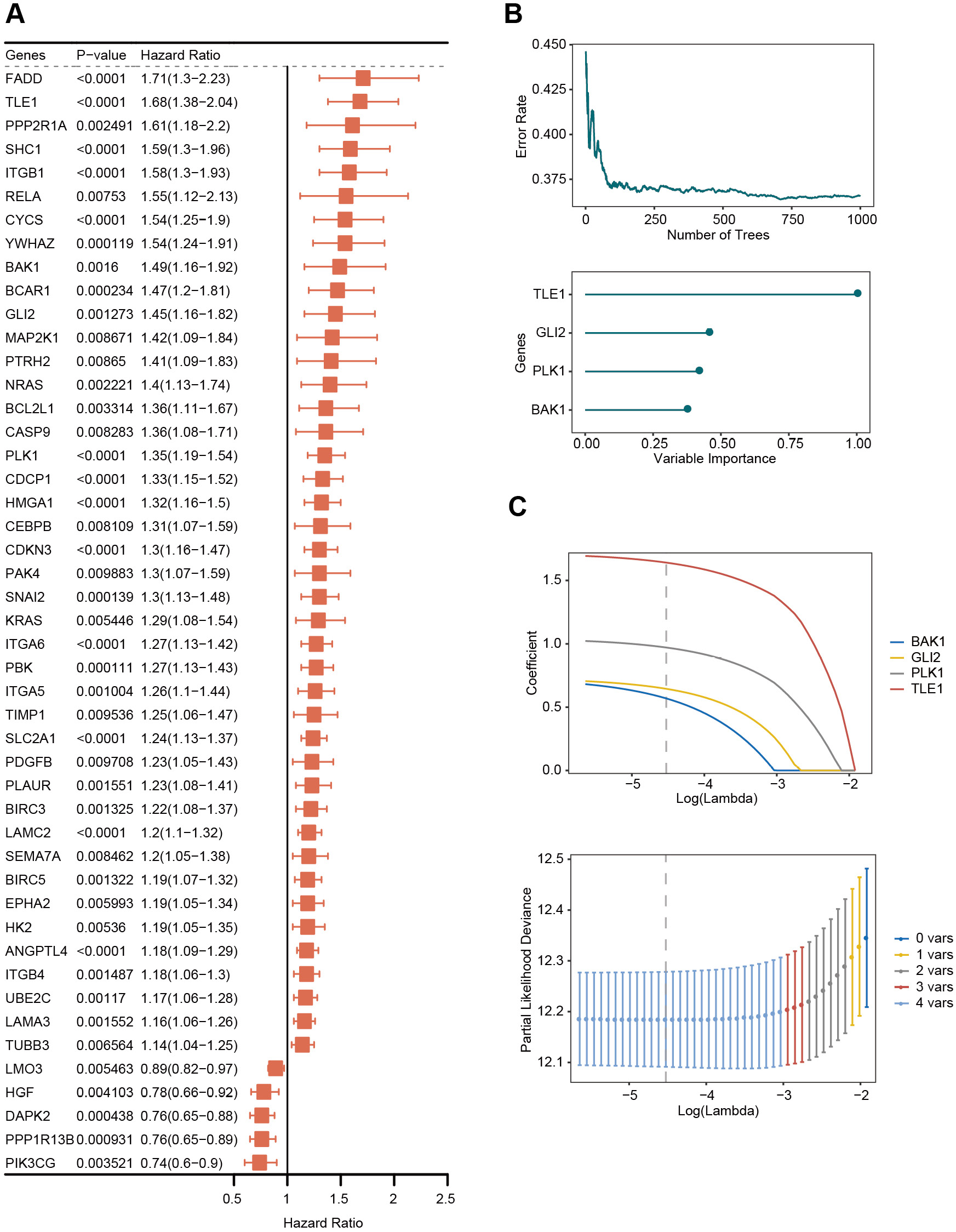
Anoikis is a type of programmed cell death that occurs when cells are detached from their ECM. Using data from the GeneCards database with a relevance score >1, 337 genes associated with anoikis were identified. A univariate analysis was performed on gene expression data from the TCGA-LUAD dataset, resulting in the identification of 47 anoikis-related genes (Figure 2A and Supplementary Table 1). Among these genes, 42 were classified as high-risk genes (hazard ratio (HR) > 1), including FADD, TLE1, PPP2R1A, SHC1, ITGB1, and RELA, and 5 genes (LMO3, HGF, DAPK2, PPP1R13B, and PIK3CG) were classified as low-risk genes (HR < 1). Further analysis using the random survival forest algorithm revealed 4 prognosis-related genes, including TLE1, GLI2, PLK1, and BAK1 (Figure 2B). Together, these genes were used to construct an anoikis score with the LASSO model, which assigned weights to each gene based on its predictive value. The anoikis score is displayed in Figure 2C.
Survival analysis
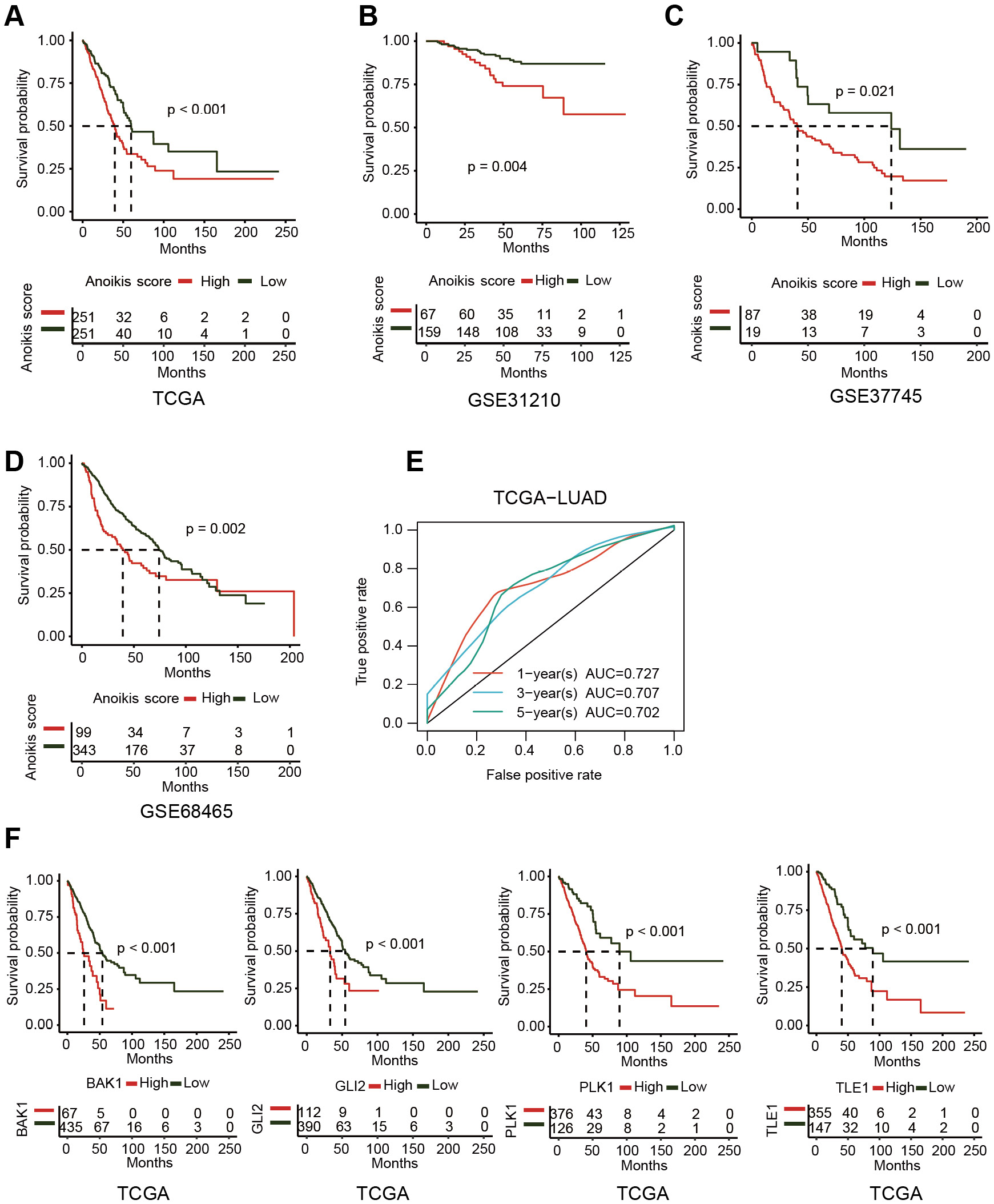
We categorized patients into 2 subgroups based on their high or low anoikis score, and the survival of LUAD patients was analyzed by comparing high-anoikis score (HAS) and low-anoikis score (LAS) groups. Based on the TCGA-LUAD training dataset, the Kaplan–Meier survival plots were created, revealing that patients in the LAS group had higher survival rate than those in the HAS group (p < 0.001, cut-off point = 2.29 and log-rank test = 13.56) (Figure 3A). Similar results were obtained from the validation dataset GSE31210 (p = 0.004, cut-off point = 1.68 and log-rank test = 8.23), GSE37745 (p = 0.021, cut-off point = 1.17 and log-rank test = 5.30) and GSE68465 (p = 0.002, cut-off point = 1.90 and log-rank test = 9.30) (Figure 3B–D and Supplementary Fig. 1). The effectiveness of the model was assessed using a ROC curve (Figure 3E). The 1-year area under the ROC curve (AUC) was 0.727, the 3-year AUC was 0.707 and the 5-year AUC was 0.702. Patients with high expression of the 4 prognosis-related genes: TLE1 (p < 0.001, cut-off point = 3.90 and log-rank test = 22.15), GLI2 (p < 0.001, cut-off point = 1.40 and log-rank test = 13.35), PLK1 (p < 0.001, cut-off point = 2.85 and log-rank test = 20.93), and BAK1 (p < 0.001, cut-off point = 5.33 and log-rank test = 24.78) had lower survival rates than those with low expression of the above genes (Figure 3F). These findings suggested that a high anoikis score predicts poor outcomes in LUAD patients.
Immune cell infiltration
and immune checkpoint analysis
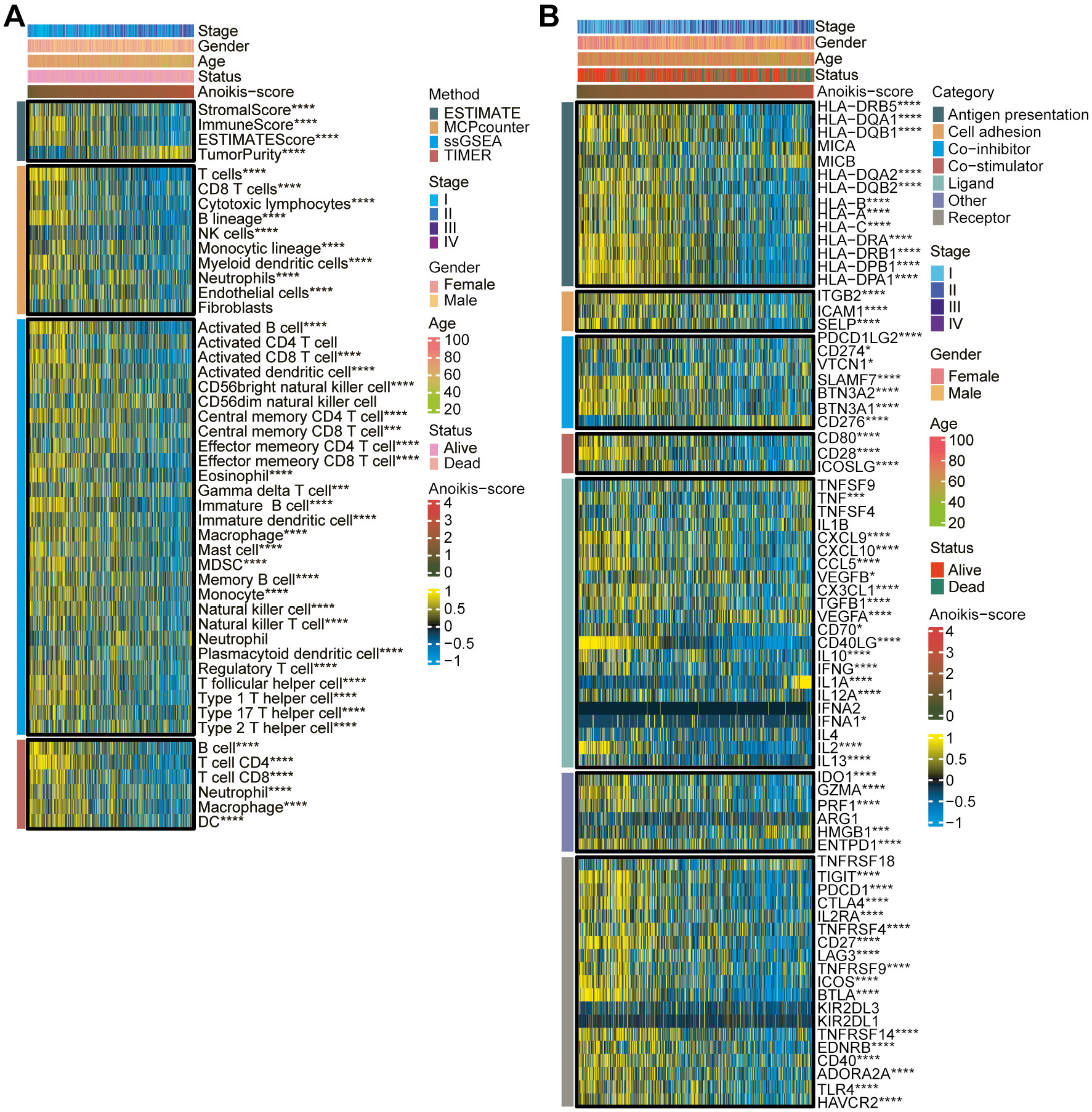
The anoikis score was correlated with immune cell infiltration using ESTIMATE, MCPCounter, ssGSEA, and TIMER algorithms (Figure 4A and Supplementary Table 2). Our results showed a negative correlation between the anoikis score and stromal score (p < 0.001, Pearson’s correlation) and immune score (p < 0.001, Pearson’s correlation). As the anoikis score increased, there was a significant decrease in the infiltration levels of eosinophils, immature B cells, mast cells, monocytes, natural killer cells, regulatory T cells, CD4+ T cells, CD8+ T cells, B cells, and macrophages.
In addition, we analyzed the expression of immune checkpoints for the anoikis score (Figure 4B and Supplementary Table 3). We found that the levels of the checkpoint for antigen presentation (HLA-DQA1 and HLA-DPB1), cell adhesion (ITGB2 and SELP), co-inhibitor (SLAMF7 and BTN3A2), co-stimulator CD28, ligand (CXCL9, CCL5, CD40LG, and IL2), and receptor (TIGIT, PDCD1, CTLA4, CD27, and TLR4) were negatively correlated with the anoikis score. However, co-inhibitor CD276 and ligands (VEGFA and IL1A) were positively correlated with the anoikis score.
Functional enrichment analysis by GSEA
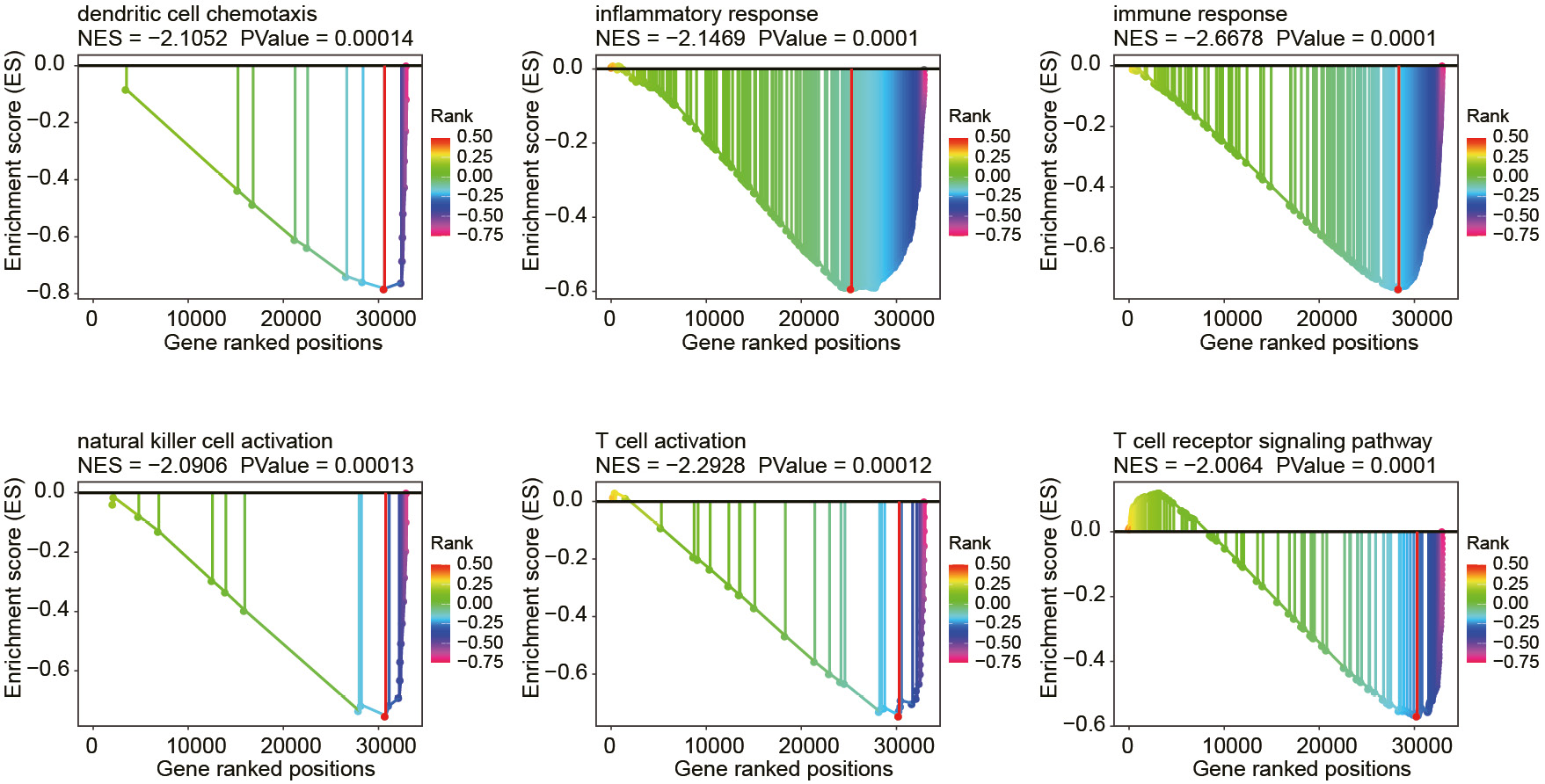
After analyzing the immune cell infiltration and immune checkpoints, we examined the functional pathways associated with the anoikis score through GSEA. We found that several functional pathways were negatively correlated with the anoikis score (Figure 5). Specifically, these pathways included the dendritic cell chemotaxis (normalized enrichment score (NES) = −2.1052, p < 0.001), inflammatory response (NES = −2.1469, p < 0.001), immune response (NES = −2.6678, p < 0.001), natural killer cell activation (NES = −2.0906, p < 0.001), T cell activation (NES = −2.2928, p < 0.001), and T cell receptor signaling pathway (NES = −2.0064, p < 0.001).
Genetic variation analysis
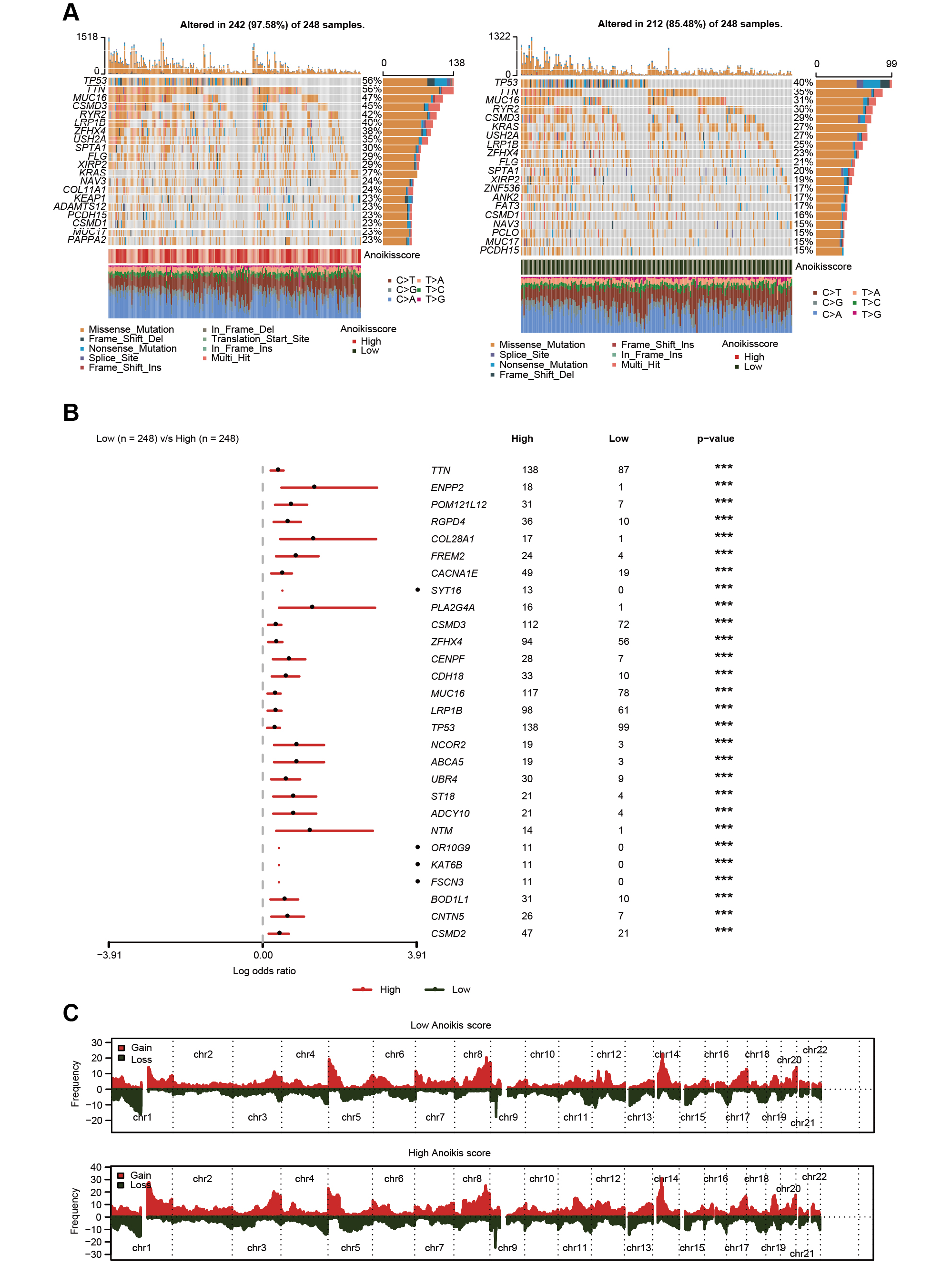
Data presented in Figure 6A visualizes the top 20 mutated genes in LUAD patients. Our analysis revealed that a mutation in one of these genes was present in 97.58% of patients in the HAS group compared to 85.48% in the LAS group. In addition, TP53 mutations were found in 56% of the patients in the HAS group and in 40% of the patients in the LAS group. The frequency of gene mutants was analyzed to further compare the number of patients with mutations between the 2 groups. Genes that showed a higher frequency of mutations in the HAS group compared to the LAS group (p < 0.001, χ2 test) included TTN (HAS: n = 138; LAS: n = 87), ENPP2 (HAS: n = 18; LAS: n = 1), MUC16 (HAS: n = 117; LAS: n = 78), LRP1B (HAS: n = 98; LAS: n = 61), TP53 (HAS: n = 138; LAS: n = 99), NCOR2 (HAS: n = 19; LAS: n = 3), CNTN5 (HAS: n = 26; LAS: n = 7), and CSMD2 (HAS: n = 47; LAS: n = 21) (Figure 6B and Supplementary Table 4). Moreover, the CNV frequency was found to be different between the 2 groups, as illustrated in Figure 6C. Our findings indicated a significant association between the anoikis score, gene mutation and CNV in LUAD patients.
Discussion
Anoikis plays a significant role in the metastatic characteristics of tumors.25 Our research has identified anoikis-associated prognostic features for LUAD. The constructed genetic feature, known as the anoikis score, has been shown to predict patient prognosis, and it outperformed multiple other prognostic models for LUAD.26, 27 The anoikis score encompasses the TLE1, GLI2 and PLK1 genes, among other factors, which play a crucial role in LUAD prognosis. Specifically, TLE1, a transcriptional repressor protein, is involved in cell growth and anti-apoptotic processes.28, 29 The TLE1 hinders Bit1-mediated anoikis by reducing the formation of the pro-apoptotic Bit1-Amino-terminal Enhancer of Split (AES) complex in the nucleus30, 31 and inhibits E-cadherin expression, promoting epithelial–mesenchymal transition.32 In our study, high levels of TLE1 were a prognostic risk factor for LUAD,33 and we demonstrated that patients in the high-TLE1 group had a lower survival rate. Similarly, the transcription factor GLI2 is positively correlated with the human epidermal growth factor receptor 2 (HER2)34, 35 and increases the resistance to anoikis in HER2-overexpressing breast cancer cells.35 The PLK1, as a mitotic master Ser/Thr kinase, is involved in cell cycle regulation and cell proliferation.36 It regulates the β-catenin protein level, which in turn protects cancer cells from anoikis by inhibiting ubiquitination and degradation of β-catenin protein.37 Therefore, high levels of GLI2 and PLK1 in LUAD could lead to poor prognosis as they enhance anti-apoptosis and migration ability, contributing to anoikis resistance. Overall, our findings illustrated the vital role of TLE1, GLI2 and PLK1 in the prognosis of LUAD and their relationship with anoikis resistance.
In both HAS and LAS groups, TTN- and TP53-mutated genes were altered, and a high anoikis score predicted a higher likelihood of TTN and TP53 mutations. These gene mutations have been shown to affect the chemotherapy response in patients with LUAD.38 The TP53 mutation is one of the most common mutations in early LUAD, observed in about 50% of patients.39 Additionally, the presence of both TP53 and PLK1 mutations in primary breast tumors has been linked to a worse prognosis.40 Tumors with TP53 deletion and high PLK1 expression may potentially benefit from PLK1 inhibitors, although the available data are controversial.36 Meanwhile, TP53 mutant cancers have been associated with higher levels of anti-tumor immunity markers, suggesting a possible relationship between TP53 mutation and the immune response.41
Following our analysis, it became evident that the increase in anoikis score led to a decrease in the levels of natural killer cells, CD4+ T cells, CD8+ T cells, B cells, and macrophages. The Hedgehog (Hh) protein, which is a signaling molecule present between cells, plays a crucial role in cancer tissue homeostasis and repair. The transcription factor GLI2, which responds to Hh, attenuates T-cell activation and proliferation by driving gene transcription.42 However, knocking out GLI2 can exclude immunosuppressive bone marrow cells and inhibit tumor growth by recruiting natural killer cells.43 The expression of GLI2 is negatively correlated with T cell activation and natural killer cells. Our analysis concluded that the anoikis-related gene GLI2 might participate in immune cell infiltration in LUAD. Meanwhile, the transcriptional activity of GLI2 can regulate cytokine communication in the tumor microenvironment, promote immunoglobulin secretion and increase interleukin (IL)-6 expression.44 By regulating GLI2, the expression of tumor growth factor (TGF)-β1 in human CD4+ T cells can be reversed.45 These findings are consistent with our functional analysis, which indicates that the anoikis score has a negative regulatory relationship with the immune response, natural killer cell activation and T cell activation in the context of LUAD.
Ultimately, our results suggested that TLE1, GLI2 and PLK1 genes may be associated with cancer cell progression, gene mutations and immune cell infiltration in LUAD. Further in vitro and in vivo experiments are necessary to elucidate the mechanisms underlying these genes and to identify potential therapeutic strategies.
Limitations
Due to limited funds, expression verification and functional analysis of anoikis-related genes were not further examined in this study. These topics will be our next research direction.
Conclusions
Our study identified a group of anoikis-related prognostic features in LUAD. High scores on these features were found to be highly predictive of poor patient outcomes. Moreover, we found a strong correlation between the anoikis score, TP53 mutation and immune cell infiltration. These findings suggest that targeting anoikis-related genes could be a promising approach for the treatment of LUAD.
Supplementary data
The supplementary materials are available at https://doi.org/10.5281/zenodo.8271767. The package contains the following files:
Supplementary Fig. 1. Patient survival analysis in the GSE31210 dataset.
Supplementary Table 1. Forty-seven anoikis-related genes were obtained by univariate analysis.
Supplementary Table 2. Correlation analysis between immune cells and anoikis score.
Supplementary Table 3. Correlation analysis between immune checkpoints and anoikis score.
Supplementary Table 4. Comparison of gene mutation frequencies.