Abstract
The advent of structural magnetic resonance imaging (sMRI) at the end of the 20th century opened the way toward a deeper understanding of the neurophysiology of psychiatric disorders, substantiating regional structural abnormalities underlying this group of clinical conditions. However, despite abundant and flourishing scientific research, sMRI methodologies are not currently integrated into daily diagnostic practice. One reason behind this failed translation may be the prevailing approach to logical reasoning in neuroimaging: The forward inference via frequentist-based statistics. This reasoning prevents clinicians from obtaining information about the selectivity of results, which are therefore of limited use regarding the definition of biomarkers and refinement of diagnostic processes. Recently, another type of inferential approach has started to emerge in the neuroimaging field: The reverse inference via Bayesian statistics. Here, we introduce the key concepts of this approach, with a particular emphasis on the clinical sMRI environment. We survey recent findings showing significant potential for clinical translation. Clinical opportunities and challenges for developing reverse inference-based neural markers for psychiatry are also discussed. We propose that a systematic sharing of imaging data across the human brain mapping community is an essential first step toward a paradigmatic clinical shift. We conclude that a defined synergy between forward-based and reverse-based sMRI research can illuminate current discussions on diagnostic brain markers, offering clarity on key issues and fostering new tailored diagnostic avenues.
Key words: magnetic resonance imaging, psychiatry, gray matter, biological markers, neurophysiology
Neuronal markers in psychiatry: (Still) an open issue
In the context of clinical research, the term “diagnostic biomarker” denotes a diverse spectrum of medically relevant signals or objective indices of clinical status with the potential to detect and/or confirm the presence of a condition of interest.1 The advent of structural magnetic resonance imaging (sMRI) nearly 3 decades ago generated fervent enthusiasm about its potential to revolutionize our comprehension of the neuroanatomical basis of psychiatric disorders and to identify clinically valuable brain-based markers. Although the development of several sMRI-based techniques and the associated exponential proliferation of studies have facilitated substantial progress in elucidating the former goal (graphical overview in Figure 1), it is crucial to emphasize that no discernible metrics or models stemming from these advanced techniques have been integrated into daily clinical diagnostic practice thus far.2, 3, 4, 5, 6, 7, 8, 9, 10, 11 As a result, contemporary psychiatric diagnoses rely on descriptive data collected through clinical observation,12 leaving the development of neuroimaging biomarkers for predicting diagnostic categories or disease progression an open challenge.
What underlies this translational gap in psychiatric neuroimaging? While some researchers advocate for the need to reform the current standards concerning group-level sample size,13, 14 others posit that the further development of single-subject methods is essential to capture inter-individual features of this group of conditions characterized by aberrations in mood, cognition and behavior.15, 16, 17 Choices pertaining population stratification, phenotypic profile selection, MRI data transformation, imaging modalities, multimodal measures, and algorithms are also a matter of ongoing debate.18, 19, 20, 21, 22, 23, 24 No less important, the clinical and biological heterogeneity of patients in terms of type of symptomatology, medication status, neurodevelopmental stage, and medical comorbidities makes it challenging to detect a sound neurophysiological signature for the clinical population of interest.25, 26, 27, 28, 29, 30
Here, we add further complexity to the issue by stating that clinical inferences drawn from canonical group-level sMRI techniques with case-control designs fall short of translational goals. The main claim of this editorial is to elucidate the inferential drawbacks evident in current sMRI psychiatric research and provide suggestions for overcoming these drawbacks within an existing knowledge framework. We propose that reverse inference reasoning via Bayesian statistics provides an ideal stepping stone to suit specific clinically relevant questions across the spectrum of translational neuroimaging.
Inferential reasoning in clinical neuroimaging
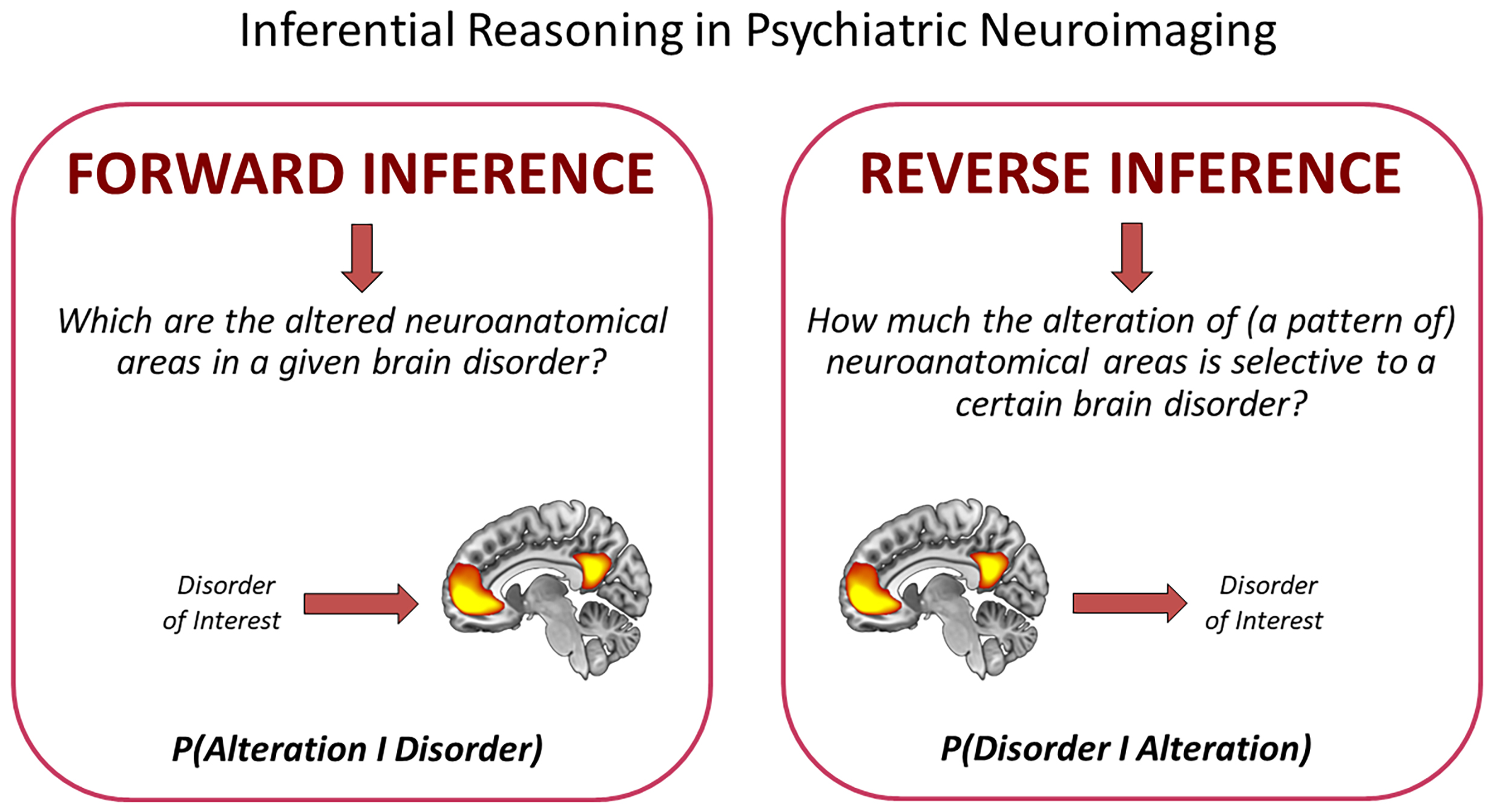
In the field of clinical neuroscience, sMRI techniques such as voxel-based morphometry (VBM), cortical thickness (CT), diffusion-weighted imaging (DWI), and diffusion tensor imaging (DTI) play a pivotal role. As well as their specific biological significance, these advanced computational techniques allow researchers to identify regional neuroanatomical variations associated with a clinical population of interest by means of a whole-brain data-driven voxel-wise intergroup comparison with a neurotypical control group.31, 32 Frequentist-based parametric statistics like the two-sample t-test are generally employed to this end, which reveals clusters of voxels where the null hypothesis (i.e., no difference in neuronal morphometry between the groups in question) is rejected using a certain user-selected p-value.33 In this scenario, a precise type of reasoning can be delineated: A “disorder-to-alteration estimation” or forward inference.34 In formal terms, the forward inference represents the probability p(alteration|disorder) of detecting alteration of some brain territories based on the hypothesis of the presence of a given disorder (Figure 1). For example, it answers the question “How probable is it to observe this pattern of neuroanatomical variation since I hypothesize that my patient is affected by this given disorder?”
Forward inference-based reasoning is used extensively in quantitative clinical sMRI research due to its role in uncovering the neural substrates co-occurring with a mental illness.35, 36 Nevertheless, it is important to note that such reasoning was not designed with translational goals in mind and has a number of fundamental limitations, most importantly its inability to establish differential diagnosis and estimate the selectivity of the effect under investigation.24, 37, 38 In other words, researchers can only infer the presence of a neuroanatomical pattern of variation in the disorder under study, but not “whether” and “to what extent” this pattern is pathognomonic for the disorder of interest or observable in other clear-cut clinical conditions. In recent years, this issue has been repeatedly accentuated in MRI-derived meta-analyses, which have demonstrated a shared neuroanatomical/functional abnormal substrate for major psychiatric disorders.39, 40, 41, 42, 43, 44, 45 For example, recent studies46, 47 taking into account forward-inference based VBM data from 82 different brain disorders and over 19,000 study participants found that a broad array of cortical and subcortical regions exhibit alterations in the context of numerous, if not all, of the considered disorders (Figure 2). Hence, this substantial convergence hinders the utility of sMRI methods for diagnostic purposes in pinpointing a specific clinical condition, given the noticeable absence of selectivity in the alteration patterns within these regions.
Building on the seminal contributions of Poldrack,48, 49 a different reasoning pattern in the human brain mapping community has recently been introduced: The reverse inference via Bayesian statistics. In formal terms, the reverse inference represents the probability p(disorder|alteration) of the presence of a given disorder based on the observed alteration (Figure 1). Moreover, in practice, it can answer the question “How probable is that my patient is indeed affected with the disorder I hypothesize since I observe this pattern of neuroanatomical variation?” Of note, this type of experimental evidence can be quantified using Bayes’ rule50 (for methodological details and explanations specific to the MRI environment, refer to Poldrack49 and Liloia et al.51). This perspective underscores that the degree of belief of a reverse inference hinges on the selectivity of neuroanatomical alterations in the disorder under scrutiny (i.e., the ratio of disorder-specific variation in the brain to the overall likelihood of variation across all other brain disorders), along with the prior belief of observing neuroanatomical variations associated with the disorder of interest. Consequently, we can obtain an estimate of how likely the disorder is given the observed pattern of neuroanatomical variation. It is crucial to acknowledge that this reasoning offers numerous advantages over canonical forward-inference methods.52 Primarily, it overcomes the binary decision of rejecting or not rejecting null hypotheses, instead providing a quantitative assessment of the evidence supporting the hypothesis under scrutiny.37
Neuronal markers and reverse inference: State of the field
While the reverse inference reasoning has provided important new information about the functional architecture of the human brain in normative populations using task-based fMRI data,48, 53, 54, 55 its application in the field of clinical sMRI has been systematically neglected. Recently, a new trend has started to emerge to address this issue. Specifically, in Cauda et al.,52 the authors delved into the 2 distinct selective volumetric alteration patterns observed in Alzheimer’s disease and schizophrenia. This was achieved by analyzing published data derived from the entire VBM BrainMap database56 utilizing the Bayes factor (BF),57 a statistical measure that quantifies the strength of evidence for one hypothesis over another. Taking the research a step further, the same group developed a meta-analytic reverse inference toolbox called Bayes fACtor mOdeliNg (BACON).58 This user-friendly and open access resource facilitates the generation of whole-brain maps, which display the selective alteration landscape at the voxel-level for a given disorder. In doing so, it compares the pattern of alteration that can be observed based on VBM results about a given disorder of interest (e.g., schizophrenia) against the pattern of alteration that can be observed based on VBM results about all the possible disorders excluding that of interest (e.g., everything but schizophrenia). Then, the BF is computed to quantify the strength of evidence for selective alterations associated with the disorder of interest (in our example, the pattern of alteration that can be observed in schizophrenia is much more likely than in any other disorder) (Figure 2).
Recently, BACON was used to explore the presence of selective brain abnormalities in autism spectrum disorder across an extensive dataset of 849 VBM experiments, accounting for over 22,000 clinical study participants diagnosed with 132 different brain disorders.51 Intriguingly, findings highlighted the existence of significant abnormalities in both cortical and cerebellar regions, with a selectivity value ≥90% (i.e., p0.9) (Figure 2). Significantly, this study not only introduced a new perspective on understanding the autistic brain, but also presented a reverse inference framework with wide-reaching potential. In fact, the BACON approach is applicable to any other clinical condition that exhibits regional variations in brain structure, as detected through VBM, CT, DWI, DTI, or other group-level sMRI techniques with case-control designs. Furthermore, this toolbox is also potentially applicable to the study of disorder-selective functional brain aberrations as revealed using several whole-brain voxel-wise nuclear medicine and fMRI techniques, such as positron emission tomography (PET), arterial spin labeling (ASL), regional homogeneity (ReHo), or amplitude of low-frequency fluctuations (ALFF), broadening its utility in the field of clinical neuroimaging. Nonetheless, it is crucial to note the current lack of studies in this specific area of research, which underscores the necessity for future multimodal imaging endeavors.
Methodological considerations and future (clinical) directions
The studies surveyed above offer hope for understanding the neuropathological basis of psychiatric disorders and yielding valuable diagnostic applications. Nevertheless, there are essential initial ways in which modeling efforts may undergo to start a paradigm shift.
Current reverse inference literature is based on meta-analytic data that, by definition, are characterized by more spatial uncertainty than native statistical parametric mapping data, thereby diminishing the ability to identify smaller but potentially highly selective brain areas.59, 60 As a matter of fact, so far, the use of meta-analytic and coordinate-based repositories (i.e., BrainMap56 or NeuroSynth53) is the prime approach to create posterior probability maps capable of giving an overarching picture of the disorder-specific brain variations utilizing VBM and fMRI data exclusively.48, 51, 61 While the trend toward sharing is gaining traction,62, 63, 64, 65 the establishment of publicly accessible, automated repositories for storing, sharing and querying voxel-wise whole-brain multimodal data from published neuroimaging studies is still in its infancy. In this context, we expect that the systematic and widespread use of fine-grained maps derived from a plethora of advanced imaging techniques will enhance the relevance of reverse inference research. The culture of sharing is also crucial to improve the generalizability of brain reverse inference models. For translational utility, these models must generalize to new individual cases, and their key signatures should be globally distributed across laboratories. This can enable testing in diverse settings with reduced complexity and error potential.
Can these methodological enhancements initiate the development of imaging biomarkers for psychiatry, or even modify current diagnostic criteria, in the absence of a definitive biological gold standard? We anticipate an iterative process in the near future, where prior neuroimaging-based prediction findings inform new hypotheses and serve as foundational knowledge for future diagnostic-oriented studies. In this context, we claim that the Bayesian reverse inference approach offers a significant advantage over traditional frequentist methods, as it allows for continuous updating of data supporting the hypotheses of interest (i.e., Bayesian updating).58, 66 Furthermore, if this process proves capable of identifying highly selective regions of interest among studies, future clinical efforts will be able to adopt tailored measurements on these specific areas, e.g., utilizing targeted strategies based on biological approaches at the micro- and meso-levels.
Another concrete goal of clinical neuroimaging based on reverse inference can be seen in its potential ability to integrate information from daily clinical practice. We expect that selective brain maps can assist in convincing clinicians of the robustness of their behavioral-based predictions, especially when the highlighted areas have been independently linked to the outcome by other reports, or to resolve uncertainties regarding complex scenarios.
In summary, this editorial discusses the latest developments in the role of reverse inference reasoning in structural MRI in psychiatry. The essence of this work is programmatic, aiming to delineate the interplay and potential synergies between forward-based and reverse-based research approaches. We posit that this distinction can illuminate current discussions on diagnostic brain markers, offering clarity on key issues, and fostering new avenues for methodological and clinical consideration in this domain.

















.jpg)
.jpg)