Abstract
Background. The expression of miR-127 has been reported to be decreased in the breast tissue of patients with breast cancer (BRC). However, the mechanism of miR-127 involvement in the pathogenesis of BRC is still unclear and requires urgent clarification.
Objectives. To explore the role of miR-127 in the pathogenesis of BRC.
Materials and methods. In this study, we measured the expression of miR-127 in blood samples of 60 BRC patients and 60 controls, investigated the influence of miR-127 on the viability and apoptosis of MCF-7 and MDA-231 cells, identified a miR-127 target gene, and determined the expression level of the target gene in the blood samples of BRC patients and controls.
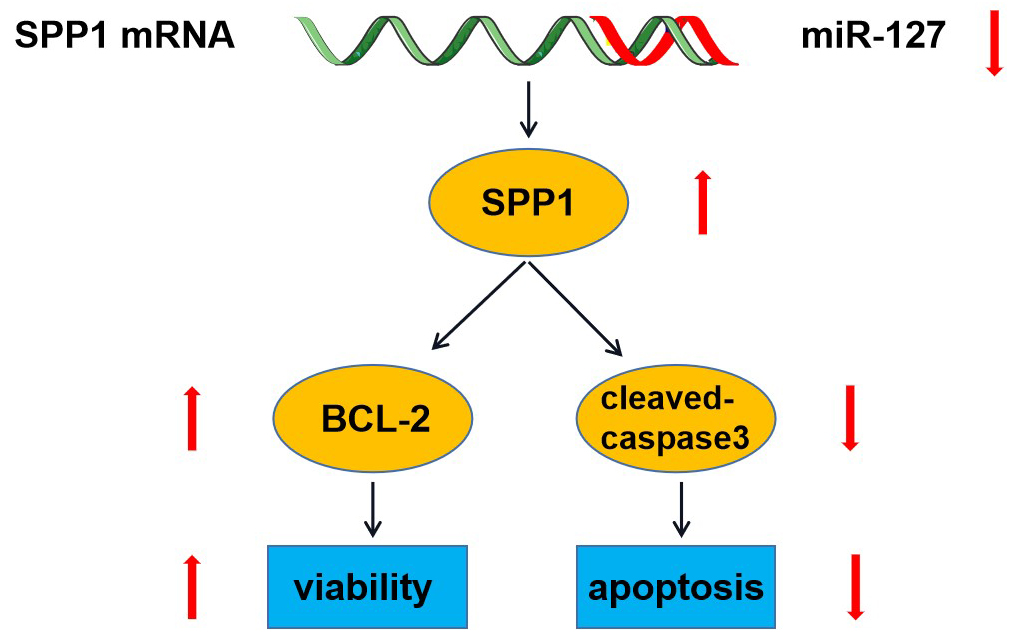
Results. We found that miR-127 expression was significantly decreased in the plasma of BRC patients compared to controls. Additionally, the upregulation of miR-127 in MCF-7 and MDA-231 cells inhibited their proliferation and promoted their apoptosis. Conversely, the downregulation of miR-127 promoted cell proliferation and inhibited their apoptosis. The SPP1 was successively predicted and validated as a target gene of miR-127. Finally, the expression level of SPP1 was significantly increased in the plasma of BRC patients compared to controls.
Conclusions. Our study demonstrated that decreased miR-127 may promote BRC cell proliferation, inhibit apoptosis and promote the occurrence of BRC through increasing the SPP1 expression level.
Key words: pathogenesis, breast cancer, SPP1, miR-127
Background
Cancer is a cause of a great economic burden and a severe public health crisis.1 Among various kinds and types of cancers, breast cancer (BRC) is the most prevalent among females worldwide.2 Although the mortality rate of BRC dropped by 40% from 1989 to 2017 in the USA, the incidence rate has still been rising by 0.3% per year.3 In China, BRC accounted for approx. 16.5% of new female cancer incidence and 7.82% of female cancer mortality in 2014.4 Traditionally, high body mass index (BMI), advanced age, as well as high plasma glucose and alcohol consumption were considered to be related to the etiology of BRC.5, 6 In addition, similar to other cancers, the etiology of BRC was closely related to the aberrant expression of multiple genes, both coding7, 8 and non-coding.9, 10 Recently, the role of non-coding gene expression, in particular microRNAs (miRs), in the occurrence of BRC has gradually become a focus of research into the etiology of BRC.11, 12 The miRs often consist of 18–25 nucleotides. They mainly regulate the translation of coding RNAs via biding to their 3’ untranslated regions, and affect the functions of coding RNAs in various pathophysiological processes.13, 14, 15
The miR-127 is a miR located at the q32 region of chromosome 1. The miR-127 gene encodes 2 kinds of mature miRs, namely miR-127-3p16, 17 and miR-127-5p. In this study, miR-127-5p was the object of our research. According to previous studies, miR-127 was involved in the pathogenesis of several diseases such as osteoarthritis,18, 19 severe pneumonia20 and premature ovarian insufficiency,21 among others. Regarding cancers, the expression of miR-127 was found to be decreased in blood samples of patients suffering from endometrial adenocarcinoma22 and colon cancer,23 according to the data from next-generation sequencing. Moreover, miR-127 was reported to participate in the occurrence and progression of squamous cell carcinoma,24, 25 gastric cancer,26 cervical cancer,27 and hepatocellular carcinoma,28, 29, 30 through the regulation of different target genes such as ADCY7, CDC6, SORT1, and FOXD1. Recently, a decrease in miR-127 expression was also reported in the BRC tissues when compared to para-carcinoma tissues, which may be caused by the hypermethylation of the miR-127 gene.31 Additionally, an RNA-protein nanoplex including miR-127 was reported to be effective in the inhibition of tumor growth in a mouse model of BRC.32 Self-assembling nanoplexes were formed by mixing the CXCR4-scFv-protamine fusion protein with miR-127. Although the results of these studies appear promising for the development of diagnostic and therapeutic applications in BRC, the expression of miR-127 in the breast tissues or blood samples of BRC patients has yet to be verified, and the mechanism of miR-127 in the pathogenesis of BRC requires urgent clarification.
Herein, we measured the expression of miR-127 in blood samples of BRC patients from Guangxi region in China, investigated the influence of miR-127 on the viability and apoptosis of MCF-7 and MDA-231 cells, and identified the target gene of miR-127. Finally, we verified the expression level of the target gene in the blood samples of BRC patients and revealed a preliminary mechanism of miR-127 involvement in the pathogenesis of BRC.
Objectives
This study aimed to explore the role of miR-127 in the pathogenesis of BRC and to identify a potential diagnostic or therapeutic biomarker for BRC.
Materials and methods
Study population
We performed a case-control study, with the approval from the Ethics Committee of Youjiang Medical University for Nationalities (approval No. 2020036688), and received written informed consent from all BRC patients and controls examined. The BRC patients (n = 60) were recruited from the Department of Breast and Thyroid Surgery (Affiliated Hospital of Youjiang Medical University for Nationalities, Baise, China) from January 2020 to June 2020. In the same period, data of age-matched controls (n = 60) were collected from the Department of Physical Examination of the same hospital. Clinical parameters, including age, progesterone receptor status, estrogen receptor status, and clinical stage were collected. Cases from families with a history of BRC, recurrent BRC or other types of cancers were removed from the study population. Controls whose family members suffered from cancer, breast lesions or mastitis, or other major human diseases such as stroke, were also removed from the study population.
qRT-PCR
A total RNA extraction kit (Qiagen, Hilden, Germany) was employed for extracting total RNA from the plasma of clinical samples or BRC cell lines. The extraction product was immediately stored at −80°C. Following complementary DNA strand synthesis from total RNA, real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR) was performed on a PCR machine (Bio-Rad, Hercules, USA) in triplicate, according to the manufacturer’s instructions (Qiagen). The primers for qRT-PCR were supplied by RiboBio (Guangzhou, China). The cel-miR-39 was used as the internal control, and the relative miR-127 level was calculated using the 2−ΔΔCT method.
Cell transfection
Two typical BRC cell lines MCF-7 and MDA-231, were purchased from Guangdao Corporation (Shanghai, China). The MCF-7 BRC cell line was isolated from the pleural effusion of a 69-year-old Caucasian female patient with BRC and retained multiple characteristics of differentiated mammary epithelium. The MDA-231 BRC cell line was isolated from the pleural effusion of a 51 year-old Caucasian female patient with BRC and barely expressed epidermal growth factor receptor and transforming growth factor-α receptor. It could form poorly differentiated adenocarcinoma in nude mice and BALB/c mice treated with amyotrophic lateral sclerosis (ALS). Cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium (Gibco, Carlsbad, USA), which was supplemented with 10% fetal bovine serum (HyClone Laboratories, Logan, USA), 80 U/mL penicillin (Gibco) and 100 μg/mL streptomycin (Gibco) under sterile conditions at 37°C and 5% CO2. The miR-127 mimics (5’-UAGUCUCGGGAGACUCGAAGUC-3’), inhibitors (5’-GACUUCGAGUCUCCCGAGACUA-3’) and negative control (5’-UAGUCUCGGGAGACUCACUACC-3’) were supplied by RiboBio. Before the transfection, MCF-7 and MDA-231 cells were transferred to a 6-well plate with 3×105 cells per well and cultured until 30% confluence. Subsequently, miR-127 mimics, inhibitors and negative control were transfected into MCF-7 and MDA-231 cells with Lipofectamine™ 3000, according to the manufacturer’s instructions. Finally, qRT-PCR was used to examine the effect of cell transfection.
Cell viability measurement
The viability of MCF-7 and MDA-231 cells was measured using Cell Counting Kit-8 (CCK-8) assay (TransGen Biotech, Beijing, China). Cells transfected with miR-127 mimics, inhibitors and negative control were seeded into a 96-well plate with 4×103 cells per well, and were cultured under sterile conditions at 37°C and 5% CO2. At 0, 24, 48, and 72 h, 100 μL of CCK-8 solution was added into partial wells of the plate and then incubated at 37°C for 2 h. Finally, the absorbance was measured at 450 nm using an enzyme-linked immunosorbent assay (ELISA) reader (Bio-Rad). Each experiment was performed in triplicate.
Cell apoptosis measurement
The apoptosis rates of MCF-7 and MDA-231 cells were measured with flow cytometry using Annexin-V-FITC/PI kit provided by Solarbio (Beijing, China). Cells transfected with miR-127 mimics, inhibitors and negative control were inoculated into a 96-well plate as above for 48 h, then collected and washed twice with cold 1× phosphate-buffered saline (PBS) and adjusted to 1×106 cells/mL using 1× binding buffer. Subsequently, 5 μL of Annexin-V-FITC and 5 μL of propidium iodide (PI) were successively added into 100 μL of cell suspension. Then, the mixture was incubated at 25°C for 15 min in the dark. Finally, flow cytometry was used to analyze the apoptosis rate of cells. Each experiment was performed in triplicate.
Western blotting
A western blotting kit (Solarbio) was used to examine the expression of protein. The harvested cells were washed twice with cold 1× PBS. Then, cold 1× lysate solution containing phenylmethylsulfonyl fluoride was added, and samples were kept on ice for 30 min. The samples were centrifuged (10,000 × g for 10 min at 4°C) and the concentration of extracted proteins was determined using a bicinchoninic acid assay (BCA). Then, proteins were separated with a 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) assay and transferred to polyvinylidene fluoride (PVDF) membranes. The membranes were completely soaked in 1× PBS with 5% bovine serum albumin (BSA) and 0.1% Tween, and shaken gently at 4°C overnight. Afterward, the membranes were incubated in the primary antibodies, namely anti-Bcl-2, anti-BAX, anti-Cytochrome c (anti-Cyt-c), anti-cleaved caspase-3, anti-β-actin, and anti-SPP1 antibodies, which were diluted in 1× PBS with 5% BSA and 0.1% Tween at 4°C overnight. After washing with 1× PBS and 0.1% Tween 5 times, the membranes were incubated in the secondary horseradish peroxidase (HRP)-linked antibodies (1:5000 diluted) at 25°C for 60 min. After washing 5 times with 1× PBS and 0.1% Tween, the bands on the membranes were detected with enhanced chemiluminescence and analyzed using ImageJ software (National Institutes of Health, Bethesda, USA). The β-actin was used as the internal control, and each experiment was performed in triplicate.
Dual-luciferase reporter assay
The SPP1 was predicted as the target gene of miR-127 according to online bioinformatics software (TargetScan; https://www.targetscan.org) analysis and a search of previous literature.33 The DNA fragments including wild-type and mutant binding sites of the 3’UTR of SPP1 were synthesized by Sangon Biotech (Shanghai, China) and then inserted into the psiCHECK-2 vector. Human embryonic kidney 293 cells were inoculated in a 24-well plate and cultured under sterile conditions at 37°C and 5% CO2. After the cells reached 70% confluence, 500 ng of recombinant psiCHECK-2 vector and 5 pmol miR-127 mimics or negative control were transfected into human embryonic kidney 293 cells using Lipofectamine™ 3000 transfection system (Invitrogen). After 48 h, firefly luciferase activity and Renilla luciferase activity were measured using the Dual-Luciferase Reporter Assay System (Promega, Madison, USA). Each experiment was performed in triplicate.
Enzyme-linked immunosorbent assay
An ELISA kit (Shunran Biotechnology, Xinzheng, China) was utilized to measure SPP1 levels in the plasma of BRC patients and controls. Plasma was diluted 1:3 in a dilution buffer and successively added into a 96-well plate by a micropipette. The mixture was incubated at 37°C for 30 min and then discarded. The wells were washed 5 times with 1× washing buffer, then the HRP-linked secondary antibody was added and incubated at 37°C for 30 min. After washing 5 times with 1× washing buffer, 50 μL of tetramethylbenzidine solution was added to the wells and incubated at 37°C for 15 min in the dark. Finally, 50 μL of terminating solution was added to the wells, and the absorbance was read at 450 nm with an ELISA reader (Bio-Rad).
Statistical analyses
The IBM SPSS v. 21 software (IBM Corp., Armonk, USA) was used for statistical analysis. Data were presented as the mean ± standard deviation (M ±SD). The normality distribution and the homogeneity of variance were evaluated using the Kolmogorov–Smirnov test and Levene’s test, respectively. Discrete variables including age and relative miR-127 level as well as SPP1 level of cases and controls were compared using the Student’s t-test, whereas continuous variables including the optical density (OD) 450 nm values, the apoptosis rates, the relative luciferase activity, and the expression levels of Bcl-2, BAX, Cyt-c, cleaved caspase-3, and SPP1 were compared using one-way analysis of variance (ANOVA) followed by the Tukey’s post hoc analysis. A value of p < 0.05 was considered statistically significant. Clinical parameters of BRC patients and controls are presented in Table 1. The results of the normality distribution and the homogeneity of variance analyses are shown in Supplementary Table 1. The results of the ANOVA followed by the post hoc analysis are shown in Supplementary Table 2.
Results
Clinical parameters and relative
miR-127 level of BRC patients and controls
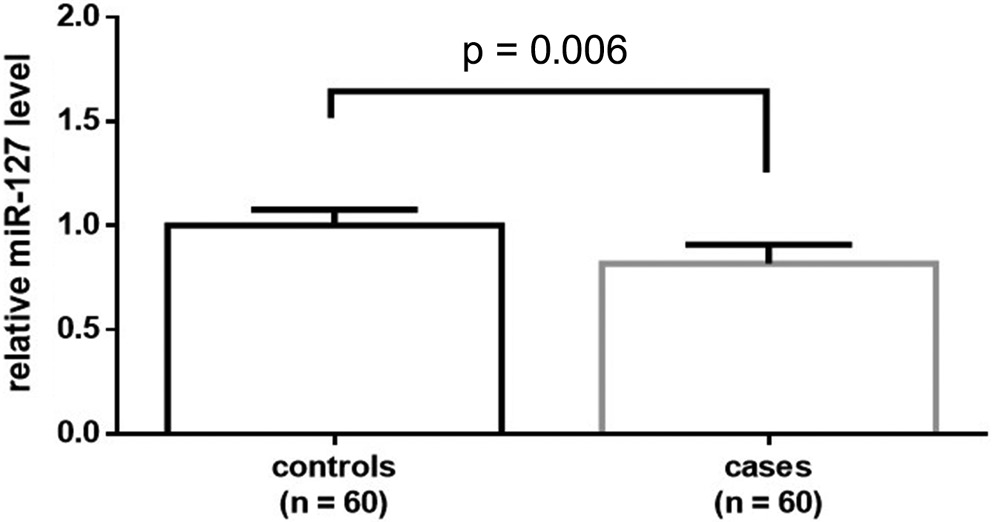
The age of BRC patients was not significantly different from that of controls (Student’s t-test, t = 0.252, degrees of freedom (df) = 118, p = 0.824) (Table 1). The percentages of BRC patients with positive estrogen receptor and progesterone receptor were 46.67% and 40.00%, respectively. The BRC patients at clinical stage IV accounted for 21.66% of cases. The relative miR-127 level was significantly decreased in the plasma of BRC patients compared to controls (Student’s t-test, t = 3.061, df = 118, p = 0.006) (Figure 1).
miR-127 inhibited viability and promoted apoptosis in MCF-7 and MDA-231 cells
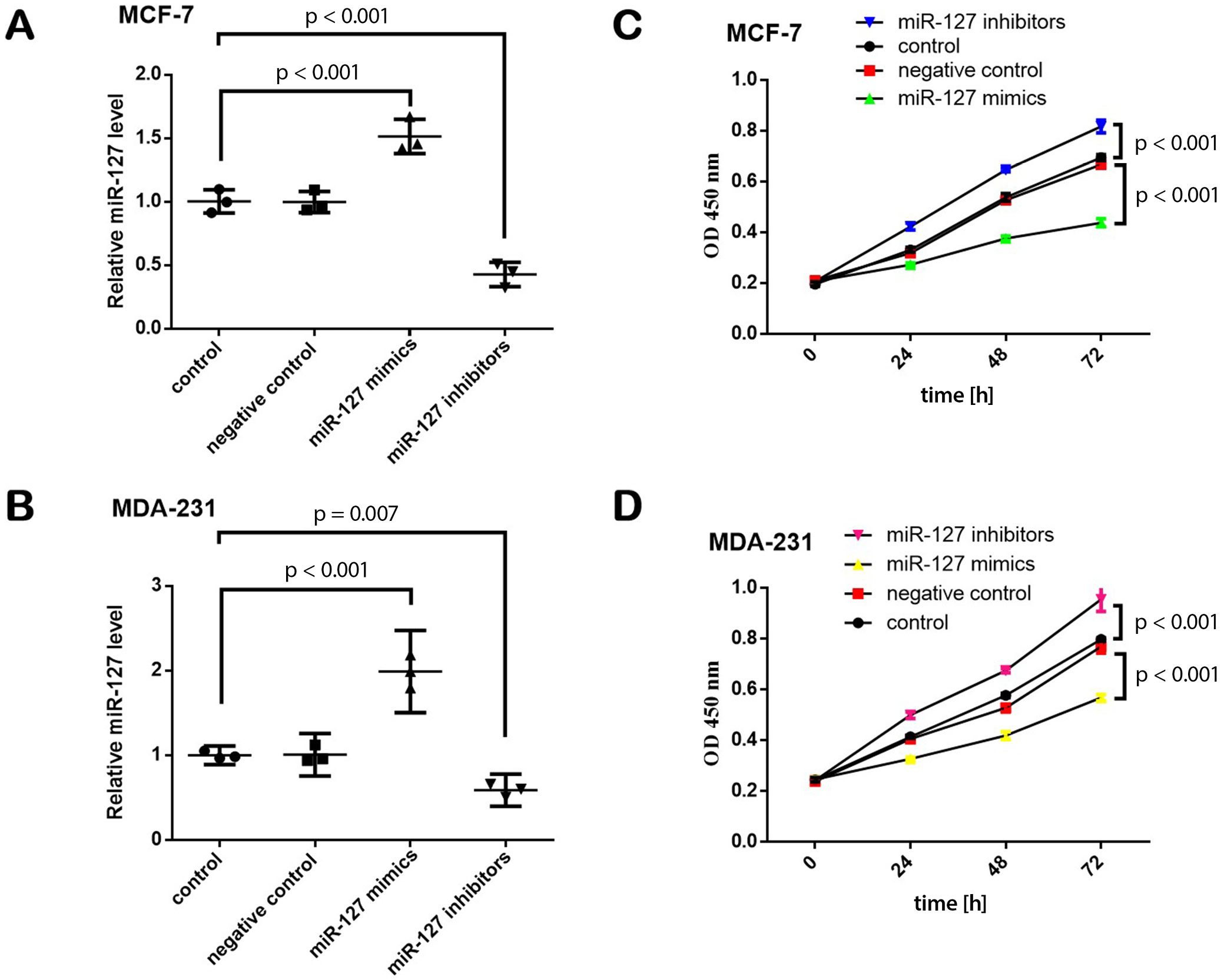
As demonstrated using the ANOVA test, there was a statistically significant effect of the treatments on the relative miR-127 level in both MCF-7 (F (3,8) = 55.436, p < 0.001) and MDA-231 cells (F (3,8) = 75.642, p < 0.001) (Figure 2A,B). When compared with controls, the relative miR-127 level in MCF-7 and MDA-231 cells was significantly upregulated after the cells were transfected with miR-127 mimics. However, the relative miR-127 level in MCF-7 and MDA-231 cells was downregulated after the cells were transfected with miR-127 inhibitors.
The treatment had a statistically significant effect on both MCF-7 viability (24h-F (3,8) = 102.191, p < 0.001; 48h-F (3,8) = 307.466, p < 0.001; 72h-F (3,8) = 256.069, p < 0.001) and MDA-231 (24h-F (3,8) = 208.510, p < 0.001; 48h-F (3,8) = 222.030, p < 0.001; 72h-F (3,8) = 96.235, p < 0.001) viability (Figure 2C,D). When compared with controls, the viability of MCF-7 and MDA-231 cells was inhibited after the cells were transfected with miR-127 mimics. Conversely, the viability of MCF-7 and MDA-231 cells was promoted after the cells were transfected with miR-127 inhibitors.
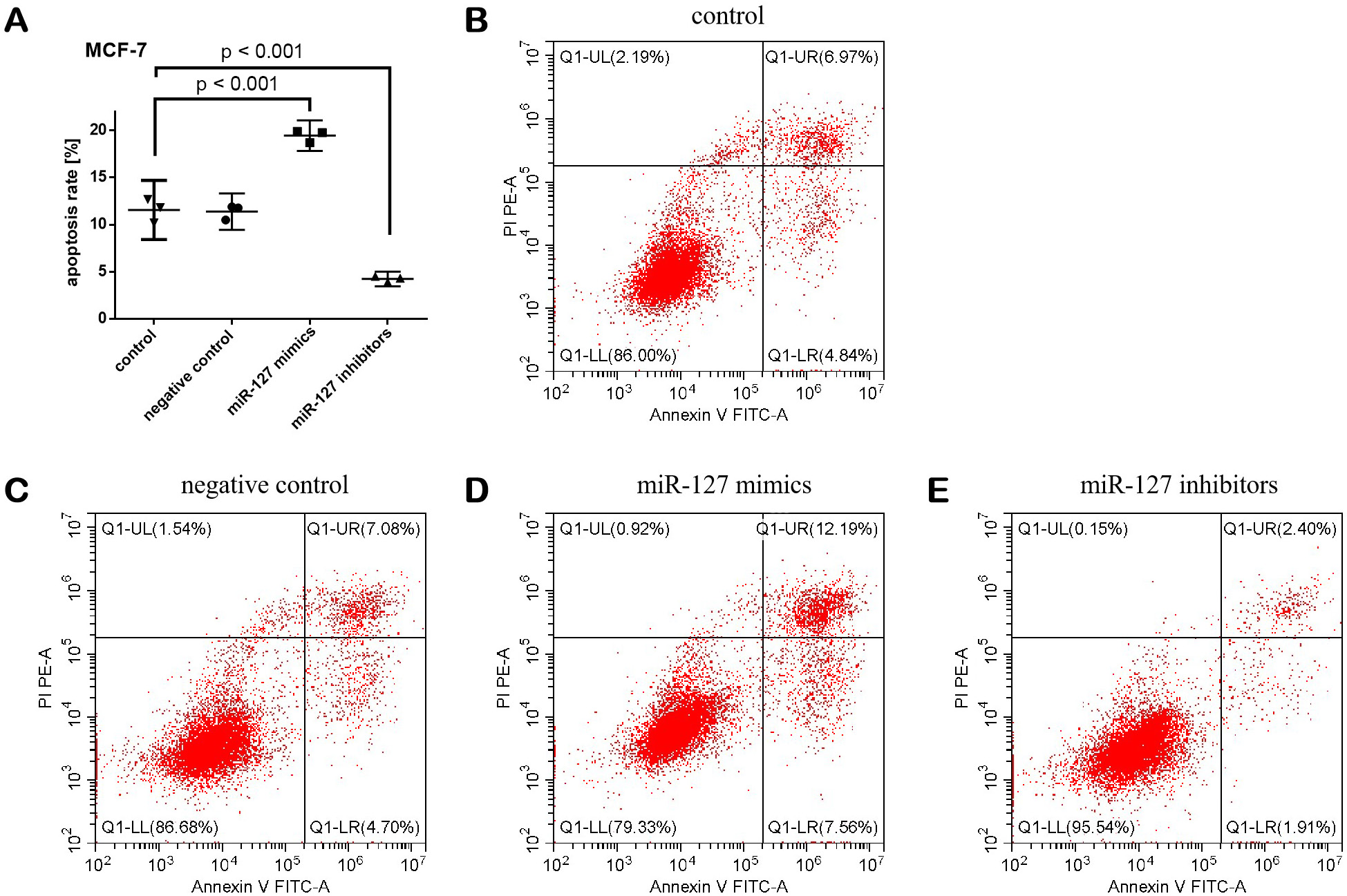
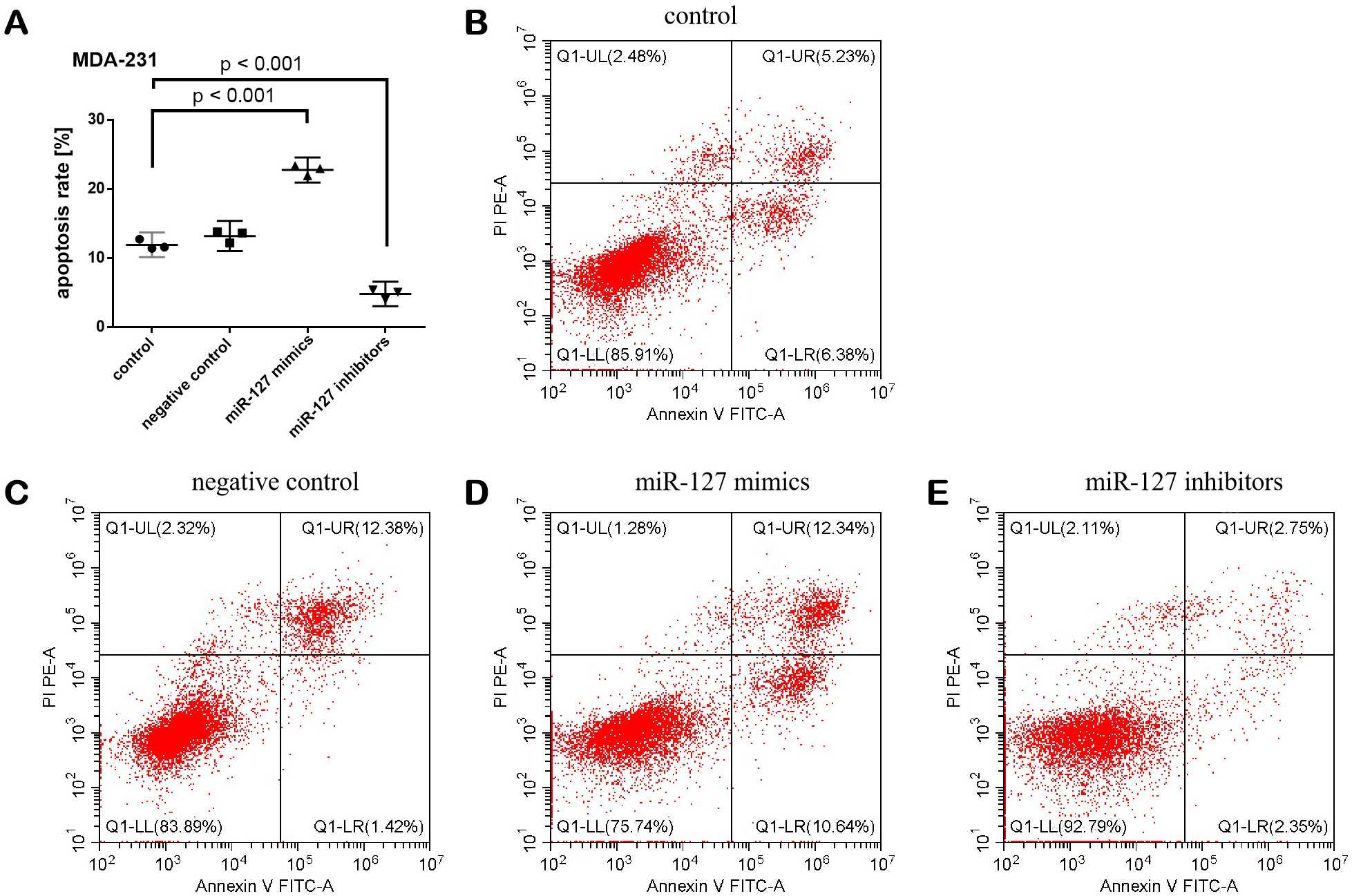
Moreover, the treatments had a statistically significant effect on both the MCF-7 apoptosis rate (F (3,8) = 169.603, p < 0.001) and the MDA-231 apoptosis rate (F (3,8) = 279.029, p < 0.001) (Figure 3, Figure 4). When compared with controls, the apoptosis rate of MCF-7 and MDA-231 cells was significantly increased after the cells were transfected with miR-127 mimics. In contrast, the apoptosis rate of MCF-7 and MDA-231 cells was inhibited after the cells were transfected with inhibitors.
MiR-127 decreased the expression level of Bcl-2 and increased the expression levels of BAX, Cyt-c and cleaved caspase-3 in MCF-7 and MDA-231 cells
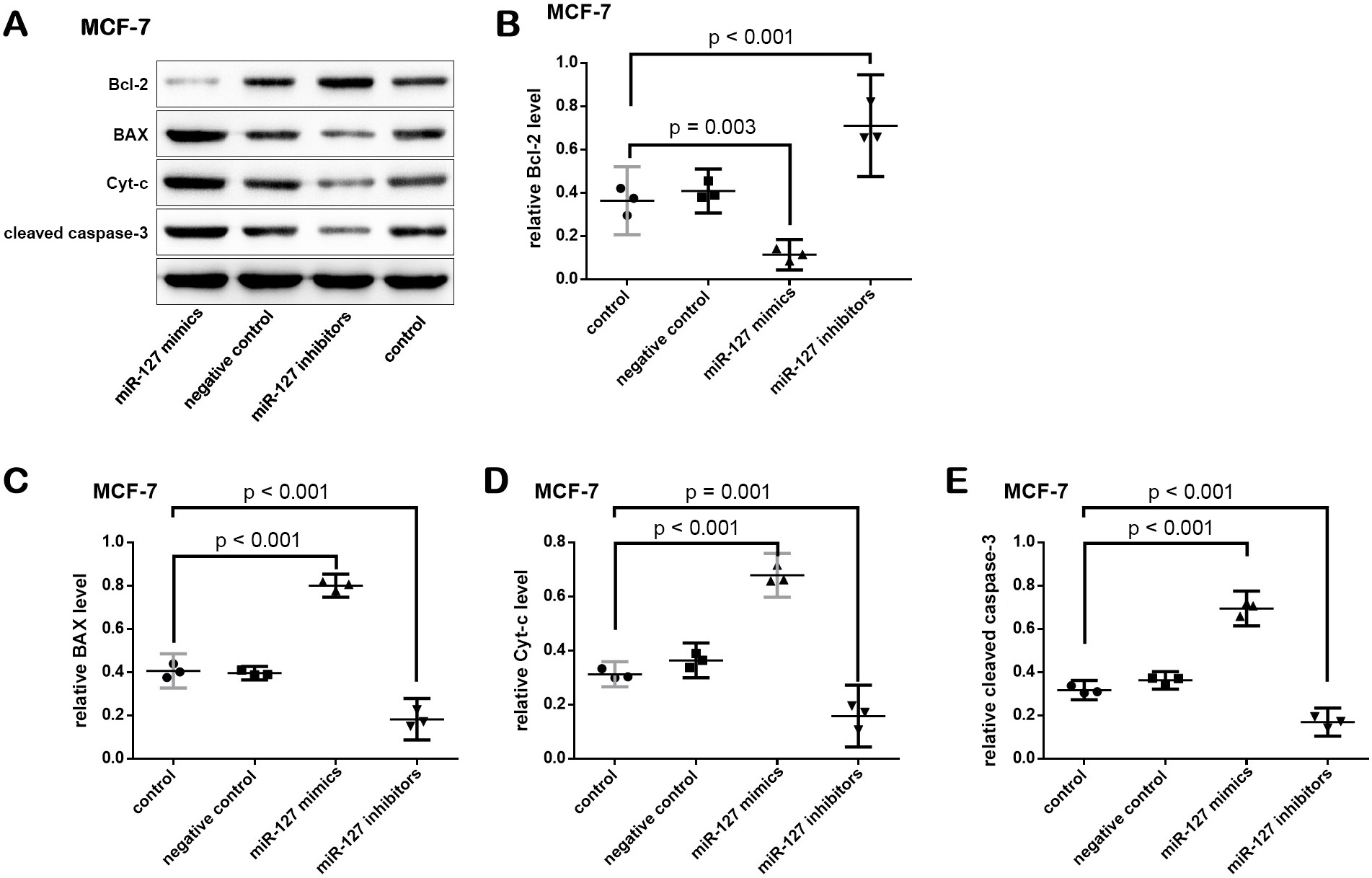
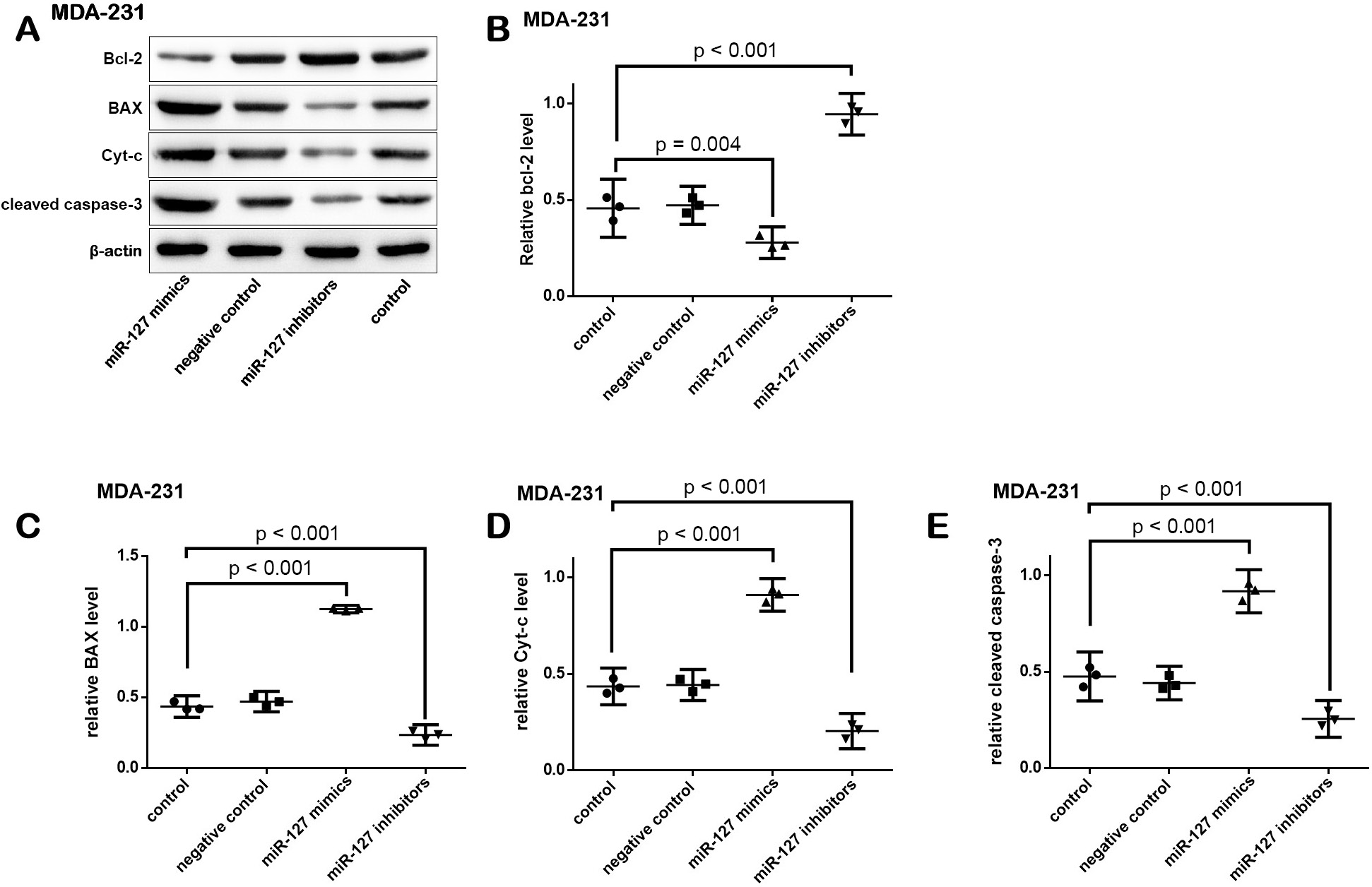
The effect of the treatments on the expression levels of Bcl-2 (F (3,8) = 46.271, p < 0.001), BAX (F (3,8) = 255.628, p < 0.001), Cyt-c (F (3,8) = 137.014, p = 0.001), and cleaved caspase-3 (F (3,8) = 255.751, p < 0.001) in MCF-7 cells was statistically significant (Figure 5). A similar effect was found in the expression levels of Bcl-2 (F (3,8) = 118.263, p < 0.001), BAX (F (3,8) = 650.679, p < 0.001), Cyt-c (F (3,8) = 209.512, p < 0.001), and cleaved caspase-3 (F (3,8) = 128.638, p < 0.001) in MDA-231 cells (Figure 6).
When compared with controls, the expression of Bcl-2 was significantly decreased, but the expression levels of BAX, Cyt-c and cleaved caspase-3 were significantly increased after MCF-7 and MDA-231 cells were transfected with miR-127 mimics. Conversely, the expression level of Bcl-2 was increased, but the expression levels of BAX, Cyt-c and cleaved caspase-3 were decreased after the cells were transfected with miR-127 inhibitors.
miR-127 inhibited SPP1 expression level through targeting 3’UTR of SPP1
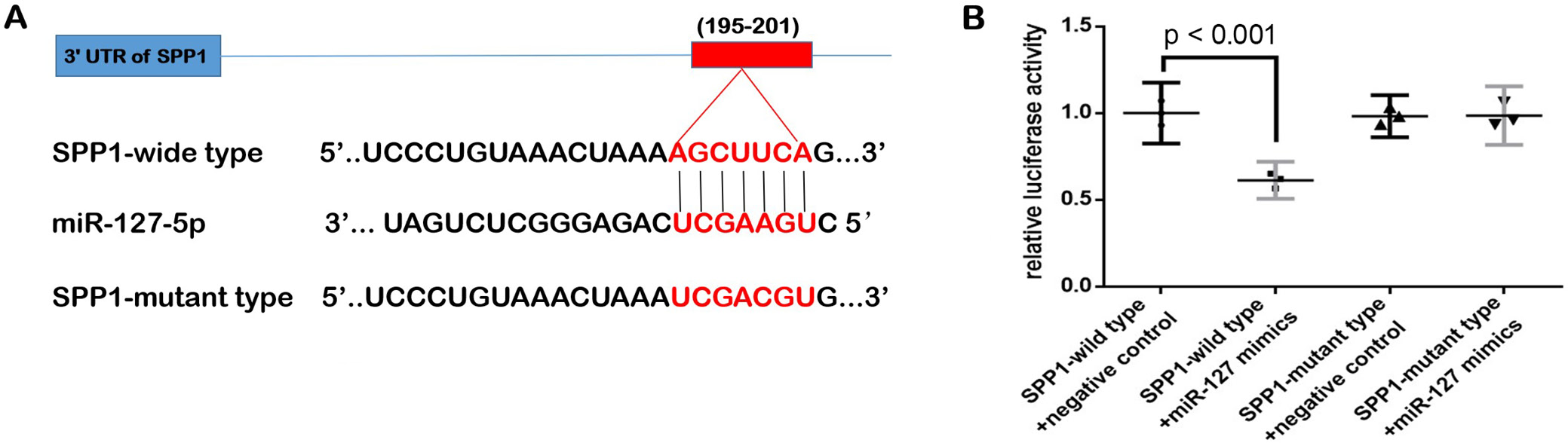
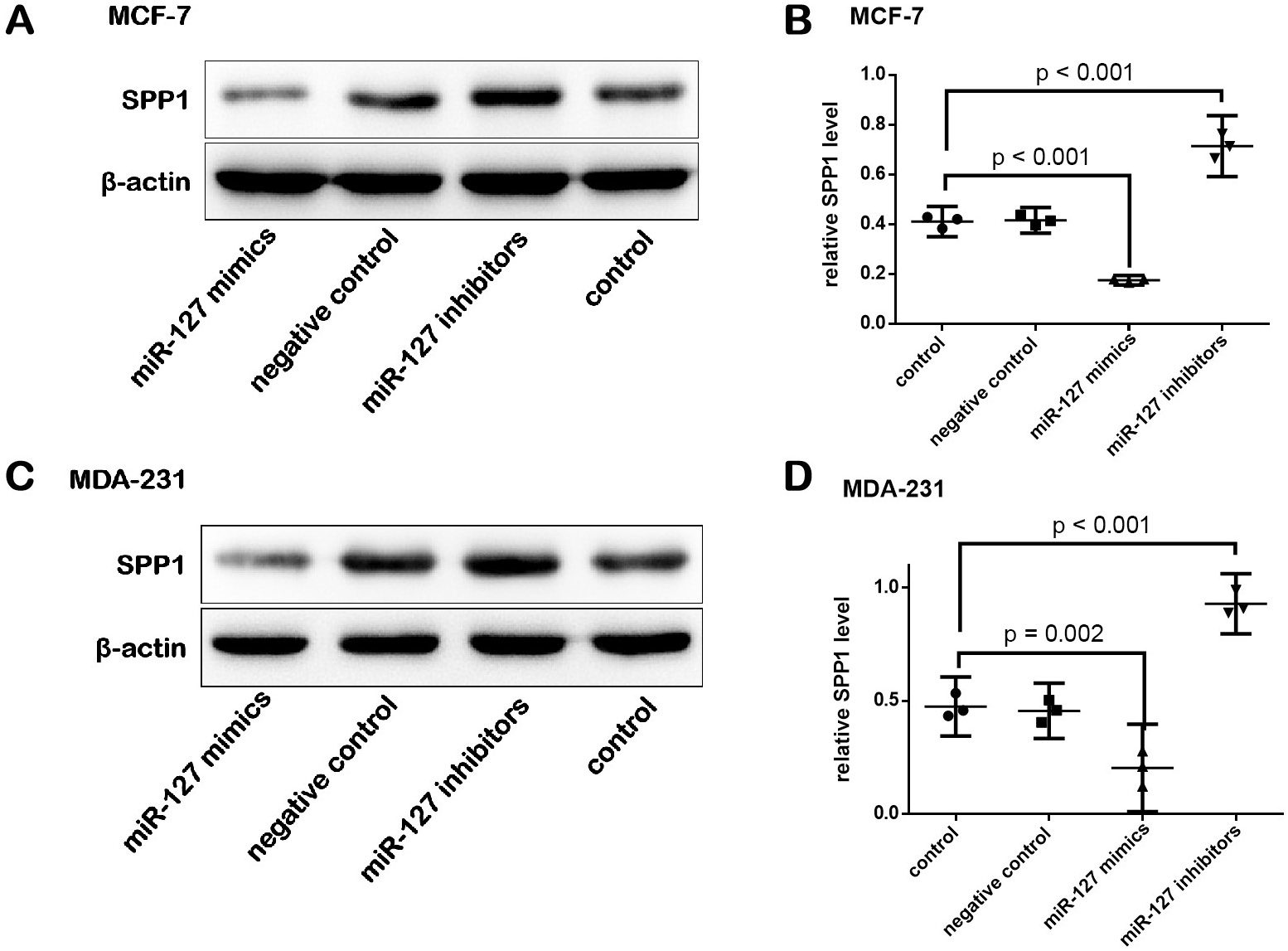
With the help of online bioinformatic software (TargetScan) analysis and previous literature,31 SPP1 was predicted as the target gene of miR-127. The predicted binding site between miR-127 and the 3’UTR of MTHFR is shown in Figure 7. Subsequently, the relationship between miR-127 and SPP1 was validated using dual-luciferase reporter assay. The results showed that miR-127 mimics significantly decreased the relative luciferase activity of SPP1-wild type instead of SPP1-mutant type compared to miR-127 inhibitors (ANOVA, F (3,8) = 30.884, p = 0.001). Furthermore, miR-127 could inhibit SPP1 expression level by targeting the 3’ untranslated region of SPP1, making SPP1 a target gene of miR-127.
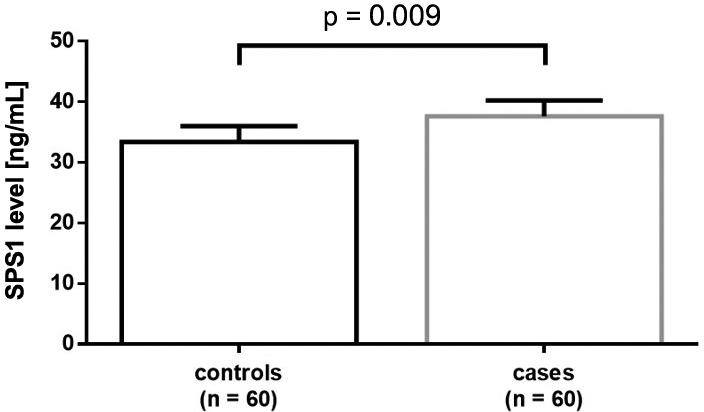
miR-127 decreased the relative SPP1 level in MCF-7 and MDA-231 cells and SPP1 was increased in the plasma of BRC patients and controls
A statistically significant effect of the treatments on both the relative SPP1 level in MCF-7 cells (F (3,8) = 165.636, p < 0.001) and MDA-231 cells (F (3,8) = 73.849, p < 0.001) was found (Figure 8). When compared with controls, the relative SPP1 level in MCF-7 and MDA-231 cells was significantly decreased after transfection with miR-127 mimics. Conversely, the relative SPP1 level in MCF-7 and MDA-231 cells was increased after cells were transfected with miR-127 inhibitors. The SPP1 level was significantly increased in the plasma of BRC patients compared to controls (Student’s t-test, t = 2.292, df = 118, p = 0.009) (Figure 9).
Discussion
Breast cancer is the most prevalent cancer type among female adults in the whole world. According to previous studies, the aberrant expression of non-coding genes like microRNAs was closely related to the etiology of BRC. The miR-127 was a kind of microRNA firstly reported to arise from the region q32 of chromosome 1 and its distinctive expression patterns might be associated with the karyotype of acute myeloid leukemia.34, 35 Based on the evidence from previous studies, the aberrant expression of miR-127 was correlated with various diseases, such as osteoarthritis. Recently, the expression level of miR-127 was also reported to be decreased in the BRC tissues compared to para-carcinoma tissues. In our study, the expression level of miR-127 in the blood samples of BRC patients from Guangxi, China was firstly measured. Similar to the study above, the relative miR-127 level was found to be significantly decreased in the plasma of BRC patients compared to controls, and the dysregulation of miR-127 in the pathogenesis of BRC was further verified. However, the effect of the dysregulation of miR-127 in the pathogenesis of BRC was not clear.
According to research on esophageal squamous cell carcinoma and hepatocellular carcinoma, the upregulation of miR-127 could inhibit the proliferation of cancer cells. As for BRC, a designed RNA-protein nanoplex including miR-127 could effectively inhibit tumor growth in a BRC mouse model. In the current study, 2 typical BRC cell lines, MCF-7 and MDA-231, were chosen to investigate the influence of miR-127 on the viability and apoptosis of BRC cells. The miR-127 successfully inhibited cell viability and promoted apoptosis in MCF-7 and MDA-231 cells. We further explored the influence of miR-127 on the expression levels of pro-proliferation factors such as Bcl-2, and pro-apoptosis factors such as BAX, Cyt-c and cleaved caspase-3 in MCF-7 and MDA-231 cells. We found that miR-127 decreased the expression of Bcl-2 and increased the levels of BAX, Cyt-c and cleaved caspase-3 in MCF-7 and MDA-231 cells. These data highlight that the upregulation of miR-127 in MCF-7 and MDA-231 cells inhibited their proliferation and promoted their apoptosis, but the downregulation of miR-127 promoted their proliferation and inhibited their apoptosis. However, the mechanism of miR-127 in the regulation of viability and apoptosis needs further clarification.
Previously, some studies reported that miR-127 might participate in the occurrence and progression of several kinds of cancer like squamous cell carcinoma via regulating some target genes like ADCY7. However, the target gene of miR-127 and its role in the pathogenesis of BRC have not been reported. With the help of online bioinformatic software analysis, previous literature33 and dual-luciferase reporter assay, SPP1 was successfully predicted and validated as a target gene of miR-127. To further explore the influence of miR-127 on SPP1, we measured the expression of SPP1 in MCF-7 and MDA-231 cells after miR-127 was up- or downregulated. The miR-127 decreased the expression level of SPP1 in MCF-7 and MDA-231 cells. The SPP1 is a gene located in the q22 region of chromosome 4 and has been investigated by numerous groups regarding the pathogenesis of various cancers, including BRC. Qin et al. reported that SPP1 was upregulated in the blood samples of patients with nasopharyngeal carcinoma, and when upregulated in CEN-2Z cancer cells, it promoted their proliferation and suppressed their apoptosis.36 Similarly, Liu et al. mentioned that SPP1 downregulation in human renal cancer cells inhibited their proliferation and increased their apoptosis.37 In addition, miR-181a was found to promote the apoptosis of cervical cancer cells by inhibiting the expression level of SPP1, and SPP1 could facilitate chemotherapy resistance of BRC cells by preventing the activation of caspase-3 into cleaved caspase-3.38 In our study, the expression level of SPP1 was significantly increased in the plasma of BRC patients compared to controls. Based on the evidence above, we could make preliminary conclusions that the downregulation of miR-127 increased the expression level of SPP1, promoted BRC cell proliferation, inhibited their apoptosis, and, finally, promoted the occurrence of BRC.
Limitations
All the patients and controls in our study were enrolled in the Affiliated Hospital of Youjiang Medical University for Nationalities. However, the data collected from other research centers will be examined in the future by our team in cooperation with researchers in other countries. Second, due to limited funding and research resources, the role of miR-127 in the pathogenesis of BRC was explored in clinical samples and cellular models, but not in animal models, and the sample size was limited in our in vitro experiments. The data from animal models will be collected, and a larger sample size in the cellular experiments will be further explored in future studies.
Conclusions
In summary, our study measured the expression of miR-127 in blood samples of BRC patients from Guangxi, China, investigated the influence of miR-127 on the viability and apoptosis of BRC cell lines, and revealed a target gene of miR-127, potentially highlighting the mechanism involving miR-127 in the pathogenesis of BRC. These outcomes may provide critical information for a new explanation of BRC pathogenesis and a potential diagnostic or therapeutic biomarker.
Supplementary data
The Supplementary Tables are available at https://doi.org
/10.5281/zenodo.7624083. The package contains the following files:
Supplementary Table 1. Normality distribution and the homogeneity of variance.
Supplementary Table 2. Results of ANOVA followed by the post hoc analysis.