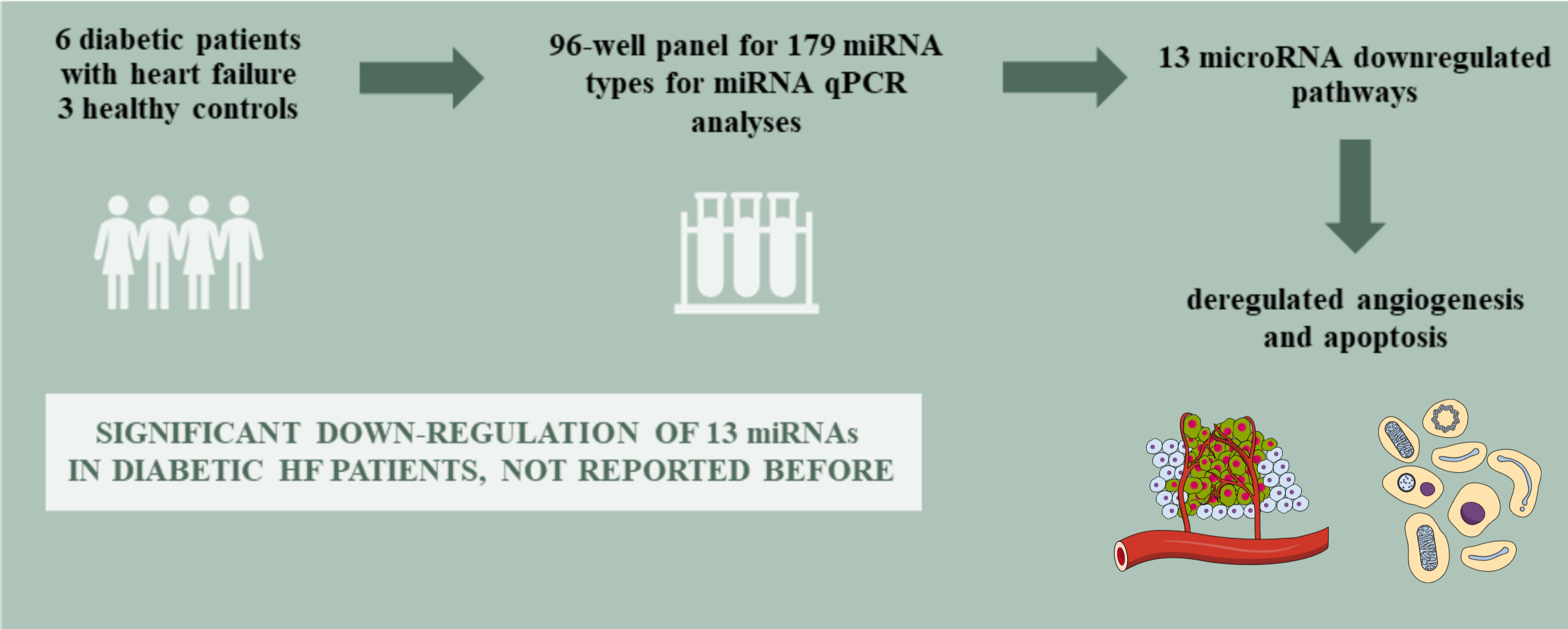
Abstract
Background. Diabetic patients present an increased risk for heart failure (HF) independently of the presence of coronary artery disease (CAD) and hypertension. However, little is known about circulatory microRNA (miRNA), an important regulatory RNA in this population.
Objectives. To evaluate serum miRNA profile of patients with diabetes mellitus (DM) and HF and analyze its relationship with pathophysiological pathways involved.
Materials and methods. The accumulation of 179 miRNAs was measured in serum of diabetic patients with HF and compared to the same measurements in healthy control subjects. The miRNAs were assayed using quantitative polymerase chain reaction (qPCR) on the Serum/Plasma Focus microRNA PCR panel (Qiagen) with LightCycler® 96 Real-Time PCR System (Roche). A pairwise comparison of mean relative miRNA accumulation levels was performed to establish those miRNAs that are differently expressed in patients with: 1) HF; 2) HF and chronic coronary syndrome (HF-CAD); and 3) HF without chronic coronary syndrome (HF-nonCAD) compared to healthy controls. To gain insight into these functions of miRNAs, we applied Gene Ontology (GO) enrichment analysis of Biological Processes and Molecular Functions of their predicted targets.
Results. The pairwise comparison revealed that 12 miRNAs were significantly downregulated in HF-CAD patients compared to controls, whereas 4 miRNAs were considerably deregulated in HF-nonCAD patients, with miRNA-15b-5p being downregulated in both groups. The GO analysis revealed that differentially accumulated targets of miRNAs include genes involved in potassium channel function, MAPK kinase activity and DNA transcription regulation, with similar alterations observed in the whole HF group and HF-CAD subgroup as well as a response to stress and apoptosis (in HF group), and genes involved in the development (in HF-CAD group). No oriented specialization of deregulated miRNA targets was observed in the HF-nonCAD subgroup.
Conclusions. We observed a significant downregulation of 13 miRNAs in diabetic HF patients, which was not reported previously either in HF or diabetic patients. Downregulated miRNAs regulate angiogenesis and apoptosis.
Key words: angiogenesis, cytokine, glucocorticoids, lipid, apoptosis
Introduction
Diabetes mellitus (DM) is an independent risk factor for heart failure (HF). Diabetic patients manifest 4 times greater risk for HF and earlier HF onset compared to the general population.1, 2 Myocardial dysfunction in DM has a heterogeneous phenotype and multifactorial origin, as it frequently occurs due to concomitant coronary artery disease (CAD) and hypertension. It may also stem from DM itself, more specifically, a condition termed diabetic cardiomyopathy.3, 4 In vitro and in vivo studies demonstrated that microRNAs (miRNAs) play important roles in the pathogenesis of cardiovascular complications of DM by exerting a direct effect on cardiomyocytes and influencing vascular smooth muscle cells, platelets, as well as processes such as lipid metabolism and inflammation.5, 6 However, little is known about circulating miRNAs in diabetic patients.7
Objectives
In this study, we aim to evaluate profiles of circulating miRNAs of diabetic patients with HF diagnosis and the correlation between deregulated miRNAs and pathophysiology of HF in DM.
Materials and methods
Study group
We included 6 adult diabetic patients, in whom HF had been diagnosed according to the European Society of Cardiology Guidelines at least 3 months before the onset of the study.8 Diabetes was diagnosed according to current guidelines.9 The control group consisted of 3 healthy age- and sex-matched subjects with no history of cardiovascular disease. The study was approved by the Medical Ethics Committee of the Medical University of Warsaw and was conducted conforming to the tenets of Declaration of Helsinki. All participants provided written informed consent. The exact inclusion and exclusion criteria are described in Supplementary Text S1.
miRNA isolation
Blood samples from 6 patients and 3 healthy donors were centrifuged at 1000 × g for 20 min at 4°C. The serum was stored at −80°C until analyzed. The miRNA was extracted from serum using a commercial column-based system (Micro RNA Concentrator, cat. No. 035-25C; A&A Biotechnology, Gdańsk, Poland), following the manufacturer’s instructions. The obtained concentration and quality of miRNA were measured using a NanoDrop Lite spectrophotometer (cat. No. LD-LITE-PR; Thermo Fisher Scientific, Waltham, USA) at 260 nm and a purity analysis based on a 260/280 ratio.
cDNA synthesis and real-time polymerase chain reaction
Briefly, 8 μL of RNA eluate was reverse transcribed in 20-μL reactions using the miRCURY LNA™ Universal RT cDNA Synthesis Kit (cat. No. 339340; Exiqon, Copenhagen, Denmark). The cDNA was diluted 30 times and assayed using real-time polymerase chain reaction (PCR), according to the protocol for the miRCURY LNA™ Universal RT microRNA PCR System (cat. No. 3393306; Exiqon); each microRNA was assayed once using quantitative PCR (qPCR) on the Serum/Plasma Focus microRNA PCR panel (cat. No. 3393325; Qiagen, Hilden, Germany). A no-template control (NTC) of water was purified with the samples and profiled like the samples. The amplification was performed in a LightCycler® 96 Real-Time PCR System (cat. No. 05815916001; Roche, Basel, Switzerland) in 96-well plates. The amplification curves were analyzed using the Roche LC software (Roche), both for the determination of the quantification cycle (Ct; using the second derivative method) and for the melting curve analysis. The amplification efficiency was calculated using algorithms similar to the LinRegPCR software (https://medischebiologie.nl/files/). All assays were inspected for distinct melting curves, and the melting temperature was checked to be within known specifications for the assay.
Quantification of miRNA accumulation
The Ct values for all miRNAs and all samples were extracted from the reverse transcription results reports and uploaded into R/Bioconductor environment (https://www.bioconductor.org/), where the analysis was performed. In particular, for data quality control and normalization, the “HTqPCR” package was employed. Two forms of normalization were applied. To compensate for the bias of obtaining Ct values in different runs, normalization to Interplate Calibrator (IPC) UniSp3 (a component of the miRCURY LNA™ Universal RT microRNA PCR System (cat. No. 3393306; Exiqon)) was performed. The second form was normalization to endogenous normalizer. As in serum samples, widely used controls such as snoRNAs or snRNAs are not accumulated on the appropriate level; therefore, we decided to perform normalization to invariant miRNA, and the intra-normalization was performed to hsa-miR-21. As a result, we obtained ΔCt values that are equal to log2-transformed miRNA accumulation levels. Differential miRNA accumulation profiles were performed using fold change calculations based on ΔCt values.
Statistical analyses
First, a heatmap was used to visualize the hierarchical clustering of 179 miRNAs. Second, principal component analysis (PCA) was applied to verify the differences in miRNA profiles between the patients. Subsequently, a pairwise comparison of mean relative miRNA accumulation levels was performed to establish those miRNAs that are differently expressed in patients with: 1) HF; 2) HF and chronic coronary syndrome (HF-CAD); and 3) HF without chronic coronary syndrome (HF-nonCAD) compared to healthy controls. The cutoff values for identifying significantly differentially accumulated miRNAs were adjusted p-value < 0.05 and fold change (FC) >2 or FC < 0.5 (same as |log2FC| > 1). To gain insight into these functions of miRNAs, we applied Gene Ontology (GO) enrichment analysis of Biological Processes and Molecular Functions of their predicted targets, selected with TargetScan (https://www.targetscan.org) and miRDB (https://mirdb.org). Data were analyzed with the R Bioconductor environment. A more detailed description is given in Supplementary Text S2.
Results
In total, 6 male patients with DM and HF were included in the preliminary analysis (Table 1). The median age of the patients was 69 years (range: 61–70 years). All patients were in the New York Heart Association (NYHA) class I or II, with median left ventricular ejection fraction (LVEF) equal to 41% (range: 35–50%) and median glycated hemoglobin (HbA1c) equal to 6.9% (6.6–7.5%). The control group consisted of 3 healthy males (age: 66 ±8 years, body mass index (BMI): 28 ±6 kg/m2).
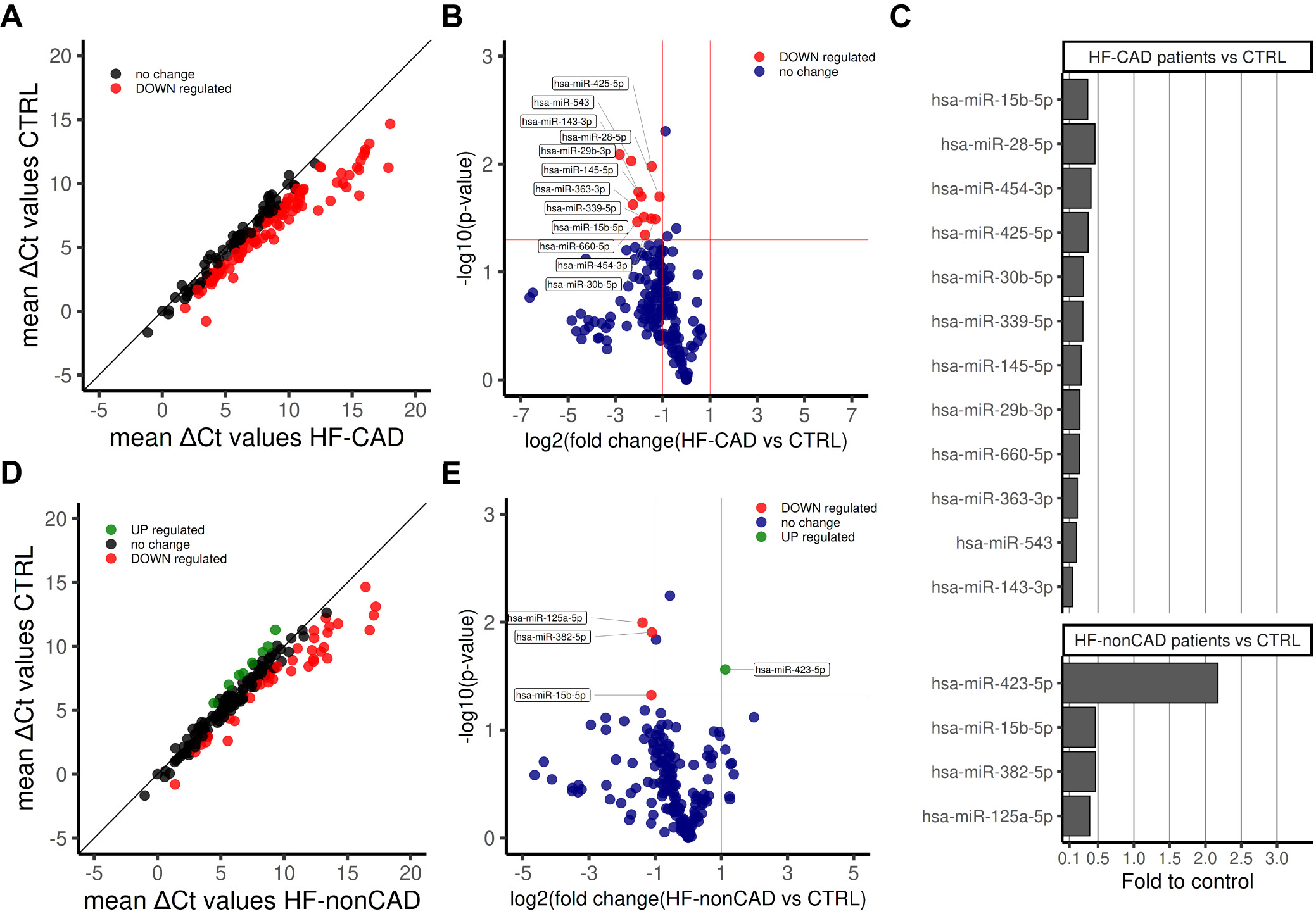
A comprehensive analysis of circulating miRNA profiles revealed that analyzed individuals fall into 4 separate clusters, with all 3 healthy individuals clustered together, as uncovered by hierarchical clustering (Supplementary Figure 1A,B) and PCA (Supplementary Figure 1C). We observed a clear shift in HF patients’ miRNA levels, with 65 miRNAs (36.3%) being downregulated (Supplementary Figure 2A). In the case of 13 miRNAs, the downregulation was significant compared to healthy controls (Supplementary Figure 2B,C). The pairwise comparison revealed that 12 miRNAs are significantly downregulated in HF-CAD patients compared to controls (Figure 1), whereas 4 miRNAs are considerably deregulated in HF-nonCAD patients (Figure 1), with miRNA-15b-5p being downregulated in both groups.
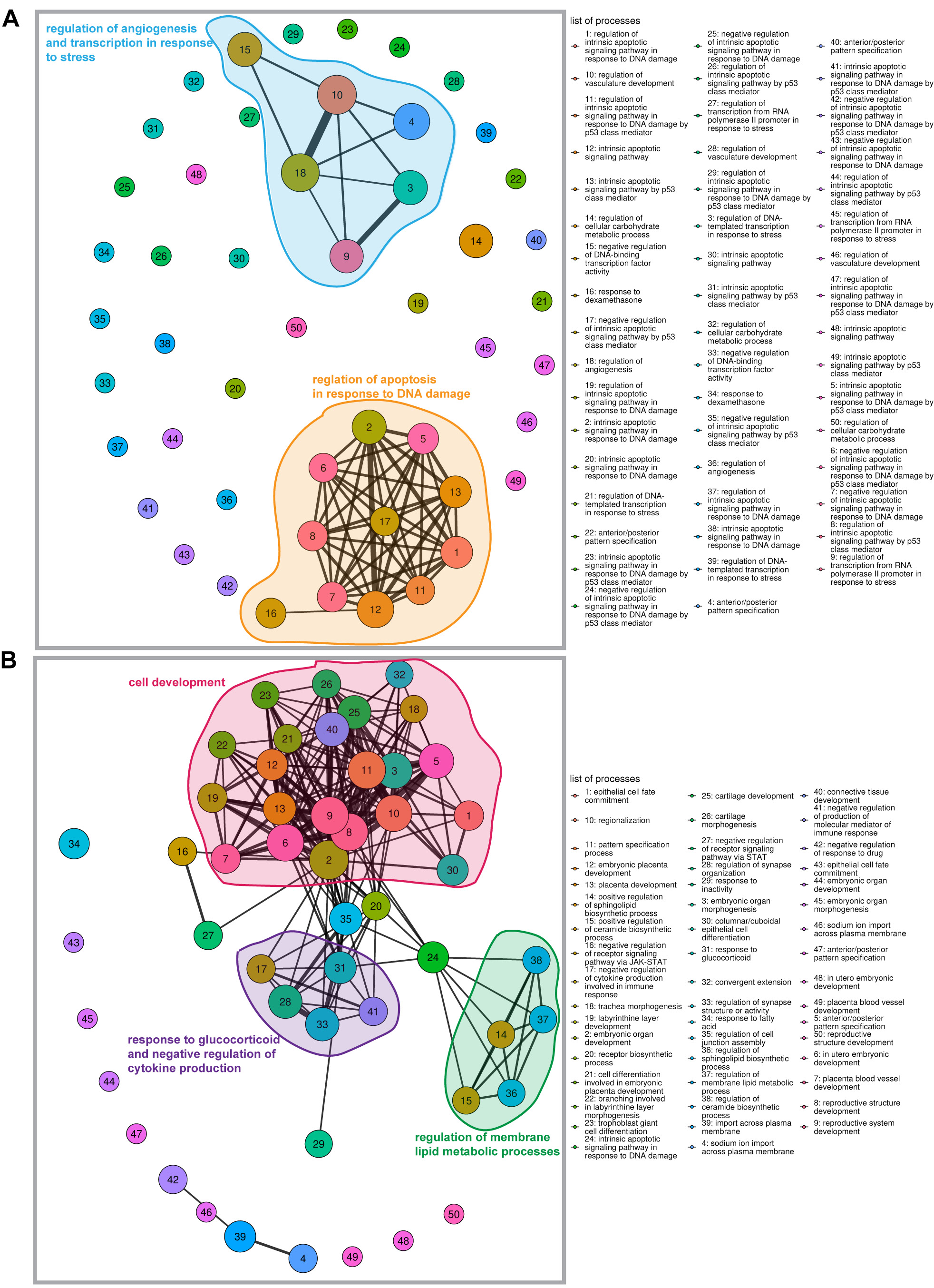
The GO analysis revealed that differentially accumulated targets of miRNAs include genes involved in potassium channel function, MAPK kinase activity and DNA transcription regulation, with similar alterations observed in the whole group and HF-CAD subgroup (Supplementary Fig. 3A) as well as genes involved in response to stress and apoptosis (HF), and genes involved in the development (HF-CAD) (Supplementary Fig. 3B). No oriented specializations of deregulated miRNA targets were observed for the HF-nonCAD subgroup. Functional networks of pathways regulated by mRNA targets of significantly altered miRNAs are presented in Figure 2.
Discussion
This is the first study that investigated serum miRNA profile in HF patients with DM. We identified significant differences in serum miRNA accumulation compared to healthy controls. The pathway analysis revealed that deregulated miRNAs might influence angiogenesis and apoptosis in response to stress in HF patients, as well as cell development, response to glucocorticoids, negative regulation of cytokine production, and membrane lipid metabolism in HF-CAD patients.
MicroRNAs downregulated in HF-CAD patients have been investigated in the preclinical setting. The miR-15 group is implicated in the negative regulation of cardiomyocyte proliferation.10 The miR-30-b has been shown to repress cyclophilin D-induced necrosis and promote hypoxia-induced apoptosis, similarly to miR-145-5p.11, 12, 13 The miR-143/145 cluster is also involved in inhibiting the phenotype switch of the contractile, mature and differentiated vascular smooth muscle cell type to a dedifferentiated, synthetic and proliferative one.14 The mir-29b-3p was identified as anti-apoptotic and anti-fibrotic to the myocardium.15, 16 Substantial evidence supports the use of anti-fibrotic miR-29b as a potential therapeutic target against myocardial remodeling, but this application still requires verification in the clinical setting.17
In the clinical context, none of the 13 downregulated miRNAs have been reported to be a diagnostic or prognostic biomarker in HF. Still, the higher circulating miR-145-5p expression is associated with a positive response to cardiac resynchronization therapy.18 According to the recent systematic review, none of the 13 circulating miRNAs, except for the miR-30 family, are deregulated in CAD patients.19 Similarly, none is deregulated in the case of DM alone.20 Therefore, we hypothesize that the identified miRNA set is distinctive for HF-CAD patients.
Limitations
In this article, we focused on male patients with HF and CAD. Thus, the results should not be extrapolated to patients with HF without an ischemic component, neither pure diabetic cardiomyopathy nor women.
Conclusions
We observed a significant downregulation of 13 miRNAs in diabetic HF patients, which was not reported previously either in HF or diabetic patients. Downregulated miRNAs regulate angiogenesis and apoptosis. Heart failure in DM is a heterogeneous disease, and these preliminary data should be viewed with caution. Further patient recruitment is ongoing to confirm these results.
Supplementary data
The Supplementary Files are available at https://doi.org/10.5281/zenodo.7295018. The package contains the following files:
Supplementary Text S1. Supplemental methods – study population.
Supplementary Text S2. Supplemental methods – statistical analysis.
Supplementary Fig. 1. Explorative analysis of serum miRNA accumulation profile in heart failure patients.
Supplementary Fig. 2. Profiling of serum miRNA differential accumulation between patients and healthy control individuals.
Supplementary Fig. 3. Gene Ontology (GO) analysis of miRNAs differentially accumulated in heart failure patients.