Abstract
Background. Mosapride significantly improves intestinal motility in liver cirrhosis, ultimately leading to the reduction in plasma endotoxin levels and bacterial translocation.
Objectives. To investigate the effects of mosapride on intestinal microecology in cirrhotic rats and its potential mechanisms.
Materials and methods. Forty-five healthy male Sprague–Dawley rats that were pathogen-free (weight 200–220 g) were randomly divided into a control group (n = 15), model group (n = 15) and mosapride group (n = 15). Then, the pathological changes in the liver and intestine were determined through tissue staining and using transmission electron microscope (TEM). Bacterial translocation was examined. High throughput 16S rRNA sequencing was performed to determine the changes of gut microbiota in each group.
Results. Compared with the model group, mosapride treatment induced no attenuation in hepatic morphology and pathology changes. The TEM indicated no differences in intestinal structure in both groups. There was a significant decline in the rate of gut microbiota translocation in the mosapride group compared with the model group. There were intestinal microbiota changes in the mosapride group compared with that of the model group, including Bacteroidetes, Prevotellaceae, Alloprevotella, Ruminiclostridium, Negativicutes, Selenomonadales, Veillonellaceae, Anaerovibrio, Campylobacterales, Epsilonbacteraeota, Helicobacter, Oscillibacter, Verrucomicrobiales, Akkermansia, Intestinimonas, Eubacterium, Clostridiaceae, Clostridium, Bacteroides, Tyzzerella, Actinobacteria, and Bifidobacteriales. Among these bacteria, Alloprevotella showed a strong correlation with the other bacteria.
Conclusions. Taken together, we concluded that mosapride may reduce intestinal bacterial translocation through regulating the gut microbiota in rats with hepatic cirrhosis.
Key words: liver cirrhosis, mosapride, bacterial translocation, gut microbiota, 16S rRNA
Background
Liver cirrhosis, the end-stage of the development of various chronic liver diseases, is characterized by the progressive replacement of functional hepatic architecture with nonfunctional fibrotic tissues.1 Patients with cirrhosis are prone to endotoxemia, spontaneous bacterial peritonitis and other bacterial infections.2 Severe complications, including hepatic encephalopathy, hepatopulmonary syndrome and liver failure, may be triggered or aggravated by infection.3, 4, 5 Bacterial translocation (BT) from the intestinal lumen to extraintestinal sites is the initial and key step in the pathogenesis of infection in cirrhosis.6
Gut microbiota disorder and increased intestinal permeability are 2 important causes of BT.7, 8 Intestinal bacterial overgrowth (IBO) has been reported in both cirrhotic patients and experimental animal models of cirrhosis.9, 10 Cirrhosis with BT displayed increased intestinal permeability, which may be caused by intestinal mucosal peroxidation and the impairment of the intestinal epithelial barrier.7, 8
Mosapride, serving as an agonist of 5-hydroxytryptamine4 (5-HT4) receptors, has been commonly utilized as a prokinetic element for treating gastrointestinal dysfunction.11 It has been reported to be effective in enhancing gastric emptying and increasing intestinal frequency, together with improving gastric mucosal injury.12 Our previous study showed that mosapride significantly increased intestinal motility, effectively preventing the translocation of bacteria and reducing plasma endotoxin level in carbon tetrachloride (CCl4)-induced cirrhotic rats.13 However, we could not identify which genus of bacteria was involved in the improvement of endotoxin-mediated mosapride.
Objectives
In this study, we aimed to investigate gut microbiota changes after the administration of mosapride in rats with liver cirrhosis.
Materials and methods
Animals
This study was approved by the Committee of Animal Research and Ethics of Zhejiang Integrated Traditional Chinese and Western Medicine Hospital, Hangzhou, China (approval No. IACUC-20190128-04). All experimental procedures on animals were in accordance with the Guide for the Care and Use of Laboratory Animals.14 Forty-five healthy male Sprague–Dawley rats, weighing 200–220 g, were used for the following experiments. Rats were acclimatized to laboratory conditions for 1 week prior to the experiment. All rats were housed under specific pathogen-free conditions where temperature (25 ±1°C) and humidity (55 ±5%) were controlled. Rats were fed ad libitum with free access to water and food under a 12-hour light-dark cycle.
Induction of liver cirrhosis and mosapride treatment
Cirrhosis was induced with subcutaneous injection of CCl4. Animals were randomly divided into the control group (n = 15), model group (n = 15) and mosapride group (n = 15). The CCl4 (Sigma-Aldrich, St. Louis, USA) was dissolved in the olive oil at a dilution rate of 2:3 (v/v). The animals were administered subcutaneously with CCl4 at an initial dose of 5 mL/kg, and then 3 mL/kg twice every week for 12 weeks. During this period, animals in the mosapride group were additionally treated by gavage with mosapride (Dainippon Sumitomo Pharmaceutical Co., Ltd., Osaka, Japan), at a daily dose of 3 mg/kg, using an orogastric feeding tube. In the control group, animals received the same volume of olive oil subcutaneously and normal saline intragastrically.
Liver and intestine histological analysis
Liver and terminal ileum specimens were fixed in 4% paraformaldehyde, followed by dehydration and paraffin embedding. The tissues were cut into sections of 4 μm in thickness and stained with hematoxylin and eosin (H&E). The Image-Pro plus v. 5.0 software (Media Cybernetics, Inc., Rockville, USA) was utilized to measure the height and the width of the villi, and the thickness of the mucosa. Picrosirius red (Polysciences, Warrington, USA) staining was performed to evaluate hepatic collagen deposition. Five randomly selected fields were obtained for each rat liver. The area of fibrosis was analyzed according to the following formula: Picrosirius red-positive area/(total area − vascular lumen area) × 100%.15
Ultrastructure analysis of the intestine
Intestinal tissues were obtained at a position that was about 15–20 cm from the ileocecal junction. Tissues were fixed using glutaraldehyde and osmic acid. Tissues were dehydrated using a series of increasing concentrations of acetone, cleared in toluene and embedded with resin. Tissues were cut with an ultramicrotome (Leica, Wetzlar, Germany) and 70 nm-thick ultrathin sections were obtained. The sections were stained with lead citrate and uranyl acetate. Microvillus and organelles were observed under the transmission electron microscope (TEM) (FEI, Eindhoven, the Netherlands), the images were captured with SIS Mega View III CCD camera and visualized using AnalySIS software (SIS, Münster, Germany).
Determination of plasma endotoxin level
Plasma endotoxin was detected using Pyrochrome® chromogenic endotoxin testing kit (Associates of Cape Cod, Falmouth, USA), according to the manufacturer’s instructions. A standard curve was plotted with endotoxin standard (Escherichia coli O113:H10). Pyrochrome lyophilized powder was gently dissolved in 3.2 mL of the reconstitution buffer. Appropriately, 50 μL of plasma sample was mixed with 50 μL of reconstitution buffer in 96-well plates, and incubated at 37°C for 12 min. The reaction was terminated by adding 25 μL of acetic acid (1:1). The samples were examined at the optical density of 405 nm.
Bacterial translocation assessment
All the procedures were performed under strict sterile conditions to avoid contamination. Animals were anesthetized, shaved and subjected to laparotomy. Mesenteric lymph nodes (MLNs) obtained from the ileocecal junction and mesenteric roots, liver and spleen tissues were homogenized in phosphate-buffered saline (PBS; 0.1 mL per 0.1 g). Then, 100 μL of homogenates were transferred to a blood plate medium and incubated for 48 h at a constant temperature of 37°C. The BT was defined as the positive culture of enteric bacteria in the MLNs, liver or spleen, regardless of the species.
High throughput 16S rRNA sequencing
Fresh fecal samples from the ileum were collected before the sacrifice, and immediately transferred into liquid nitrogen container for temporary storage. Fecal genomic DNA was extracted using QIAamp DNA Stool Mini Kit (Qiagen, Hilden, Germany). The polymerase chain reaction (PCR) amplification was conducted to amplify V3 and V4 regions of 16S rRNA based on specific primers of 338F-806R (5’-ACTCCTACGGGAGGCAGCAG-3’; 5’-GGACTACHVGGGTWTCTAAT-3’). Purified PCR products for V3 and V4 regions in 16S rRNA were subject to the high-throughput sequencing, using the Illumina MiSeq device (Illumina, San Diego, USA). Quantitative Insights Into Microbial Ecology (QIIME 2; http://qiime.org) platform was used to study the effects of mosapride treatment on the microbiome in cirrhotic rats.16 The DADA2 was used to remove noise signals.17
Statistical analyses
GraphPad Prism v. 6.0 software (GraphPad Software, San Diego, USA) was used for the data analysis. Measurement data were presented as the median and quartiles (1st and 3rd quartile (Q1, Q3)). Statistical significance was ascertained using nonparametric Mann–Whitney test. The value of p < 0.05 was considered statistically significant.
Community composition and clustering analysis were presented using stacked bar charts and a heat map. Alpha diversity was evaluated by calculating Chao1, Good’s coverage, operational taxonomic units (OTUs), and Shannon’s and Simpson’s indices with the script from QIIME 2.16 The Kruskal–Wallis test was used to test the differences in alpha diversities among the groups. The Chao1 index for abundance data was used to estimate species richness, which is suitable for the estimation of the number of shared species in multiple assemblages.18, 19 The Good’s coverage assessed sampling depth coverage.20 The OTUs represent the number of different features. The Shannon’s diversity index was used to measure the diversity of species in a community.21 The Simpson’s index shows the representativeness of a species.22 The beta diversity was used to measure the similarity in the bacterial composition between the different groups. Unweighted UniFrac distances were used for evaluating beta diversity with principal coordinates analysis (PCoA).23 The analysis of similarities (ANOSIM) was used to calculate the p-values for beta diversity.24 Linear discriminant analysis effect size (LEfSe) was used for high-dimensional class comparisons.25 The Bugbase algorithm was used to predict bacterial phenotype associated with cirrhosis.26
Results
Mosapride did not alleviate liver injury in cirrhotic rats
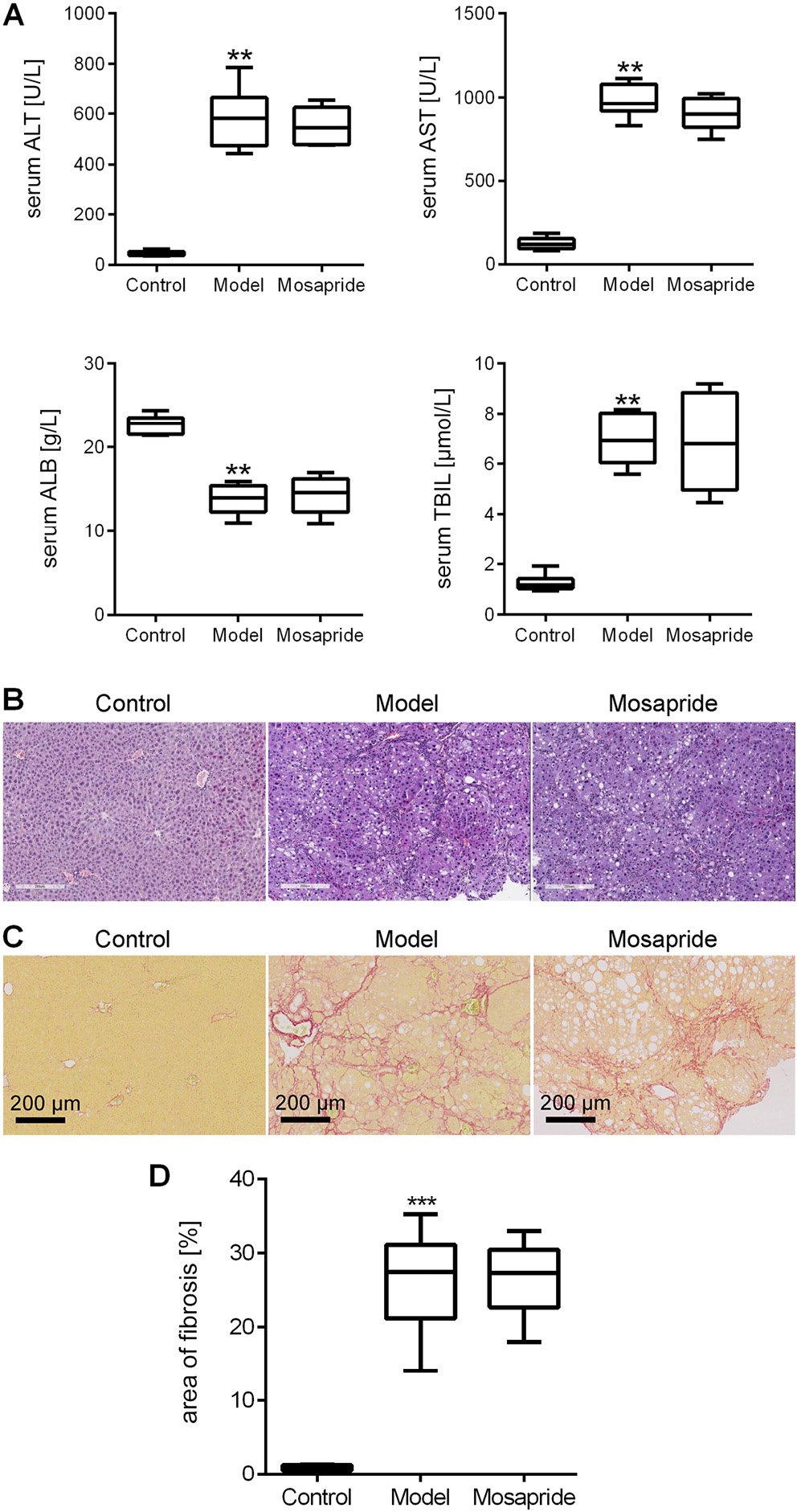
Three rats in the model group and 2 rats in the mosapride group died at the end of the 12-week experiment. The CCl4 significantly increased the level of alanine aminotransferase (ALT) (p = 0.0022), aspartate aminotransferase (AST) (p = 0.0022) and total bilirubin (TBIL) (p = 0.0022) (all p-values were calculated using Mann–Whitney test). Serum albumin (ALB) was significantly decreased by CCl4 (p = 0.0022; Mann–Whitney test) (Figure 1A). The H&E and Picrosirius red staining indicated that the normal liver structure was severely destroyed with significant fibrous septa and cirrhotic nodules in the model group (Figure 1B,C). The quantification of Picrosirius red staining showed that the fibrotic area was 0.70 (0.52, 1.15) in the control group, 27.42 (22.33, 30.16) in the model group, and 27.32 (23.21, 29.87) in the mosapride group (Figure 1D). The CCl4 significantly induced fibrosis in the model group compared to the control group (p < 0.0001; Mann–Whitney test), while no significant difference was observed between model and mosapride groups (p = 0.9502; Mann–Whitney test).
Mosapride reduced plasma
endotoxin level and BT
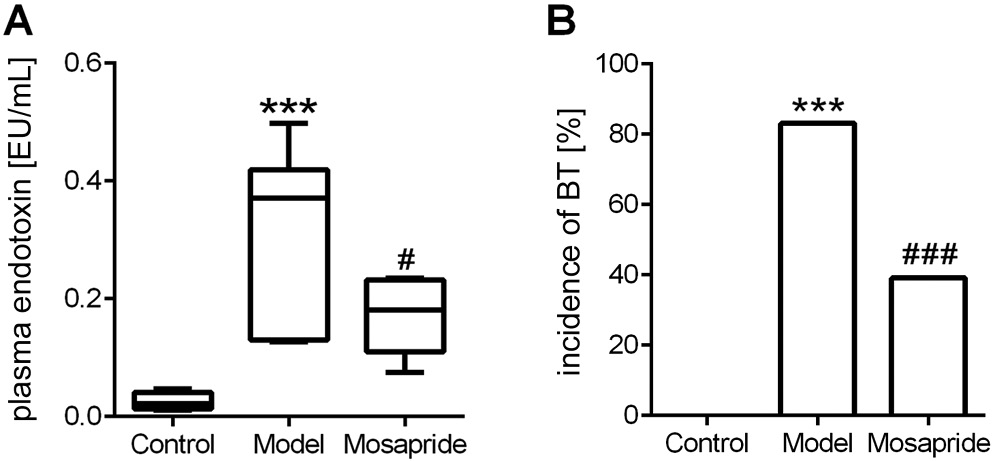
As shown in Figure 2A, the plasma endotoxin level in the model group was significantly increased compared to the control group: 0.02 (0.01, 0.04) EU/mL (control) compared to 0.37 (0.14, 0.41) EU/mL in the model group (p < 0.0001; Mann–Whitney test). The plasma endotoxin level in the mosapride group was 0.18 (0.11, 0.22) EU/mL, which was significantly decreased compared to the model group (p = 0.0223; Mann–Whitney test).
The BT was detected in 10 out of 12 rats (83.3%) in the model group, whereas its occurrence was nil in the control group (p < 0.0001; Fisher’s exact test). The incidence of BT was significantly lower in the mosapride group (5 out of 13, 38.5%) than in the model group (p < 0.0001; Fisher’s exact test) (Figure 2B). Table 1 shows the bacteria and the sites where bacteria were isolated. Escherichia coli from MLNs and liver tissues were counted.
Mosapride did not improve intestinal mucosa damage
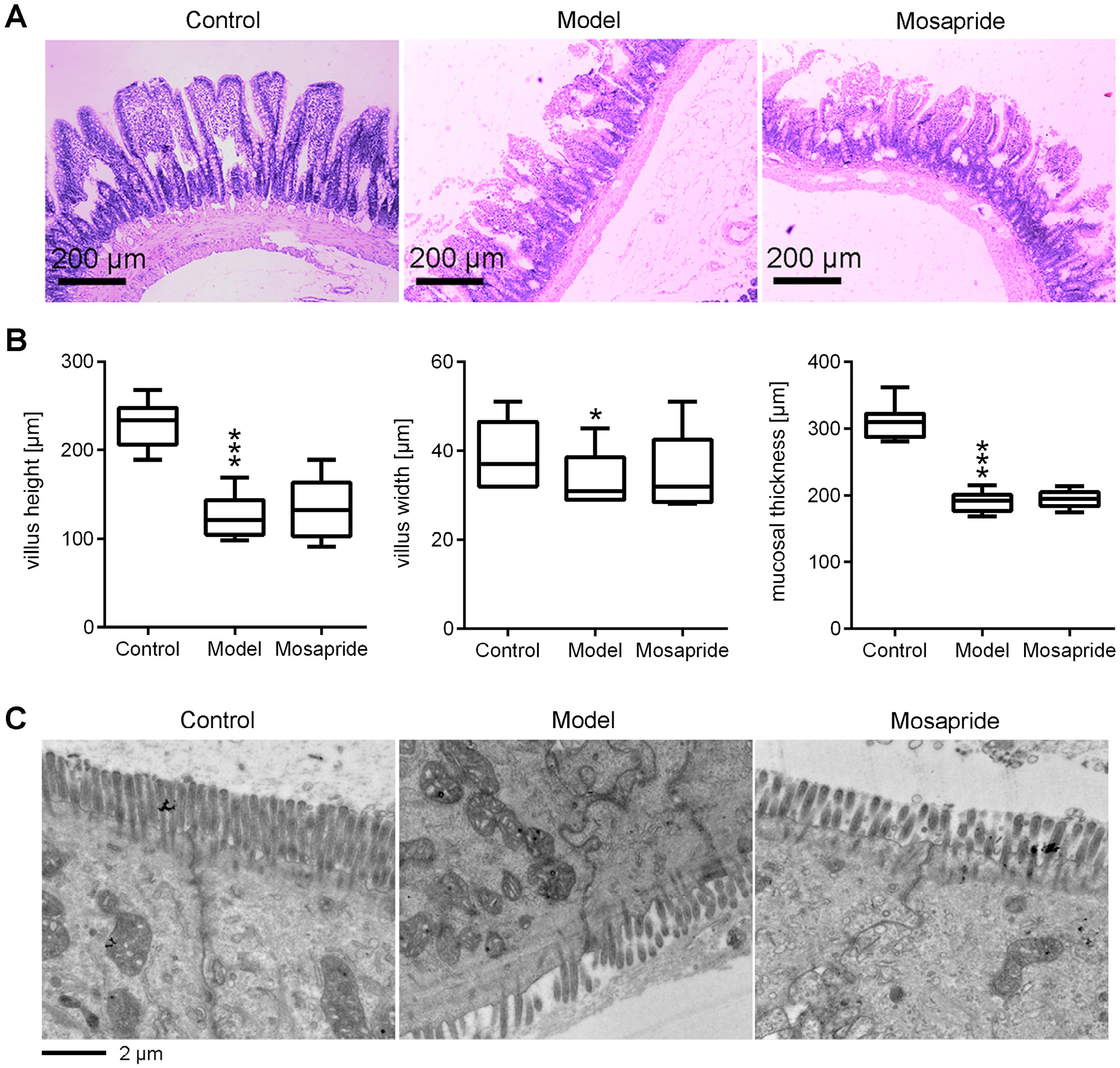
As illustrated in Figure 3A, the ileum tissues of the control rats displayed a normal intact architecture of intestinal mucosa without hyperemia, edema or inflammatory cell infiltration. The villi were well arranged, smooth and intact. In the model and mosapride groups, the mucosa was destructed with atrophic and shorter villi. As presented in Figure 3B, the villi height and width, as well as the thickness of mucosa in the model group were significantly decreased compared to the control group (p < 0.0001, p = 0.0417 and p < 0.0001, respectively; Mann–Whitney test). There were no differences in the villi height, villi width and thickness of mucosa between the model group and the mosapride group (p = 0.5601, p = 0.9155 and p = 0.7135, respectively; Mann–Whitney test). Representative images of intestinal ultrastructural changes were shown in Figure 3C. The epithelial cells in the control group were tightly arranged with neatly arranged surface microvilli. The paracellular spaces were narrow and the tight junctions of the epithelial cells in the control group were intact and clear. By contrast, the tight junctions were discontinuous and the paracellular spaces were widen in both, the model group and the mosapride group.
Effects of mosapride on the abundance of intestinal bacteria at the phylum and genus levels
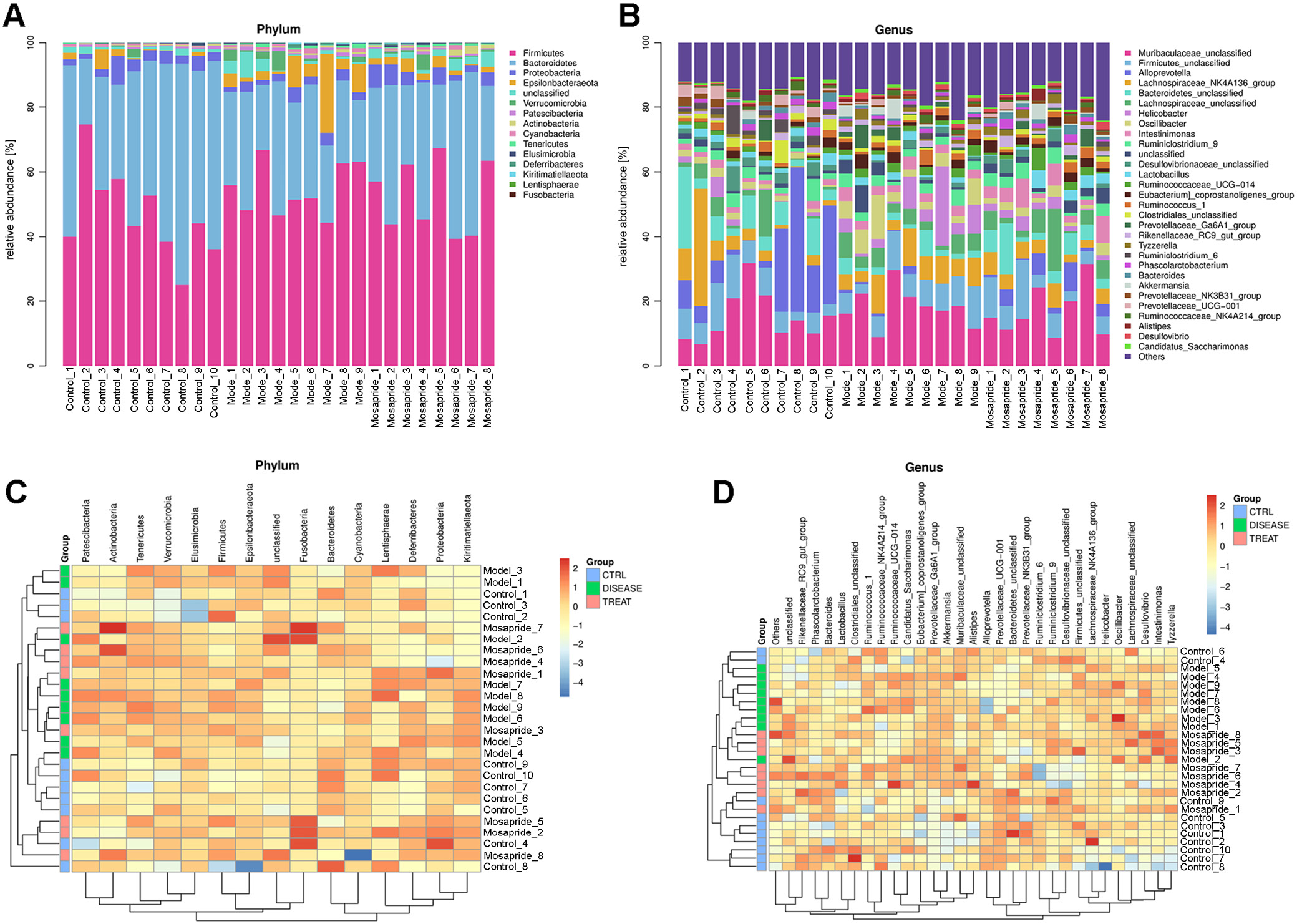
We analyzed the abundance of intestinal bacteria among 3 groups at the phylum and genus levels, and the results are shown in Figure 4A,B. In the control group, Bacteroidetes were the most abundant, followed by Firmicutes and Proteobacteria. In the model group, Firmicutes were the most abundant, followed by Bacteroidetes and Epsilonbacteraeota. In the mosapride group, the top 3 abundant bacteria were Firmicutes, Bacteroidetes and Proteobacteria. Figure 4C,D showed the clustering analysis of intestinal bacteria in the heat map. In the model group, there was a decrease of Prevotellaceae, Bacteroidetes, Alloprevotella, Desulfovibrionaceae, Phascolarctobacterium, Eubacterium, Clostridiales, and Ruminiclostridium when compared to the control group, which showed an increase after the administration of mosapride. In contrast, Firmicutes, Prevotella, Oscillibacter, Helicobacteria, Candidatus, Muribaculaceae, and Akkermansia were increased in the model group, and they decreased after the administration of mosapride. The levels of Lachnospiraceae, Intestinimonas and Tyzzerella showed an increase in the model group compared to the control group, and a further increase after mosapride administration.
Effect of mosapride on bacterial diversity in cirrhotic rats
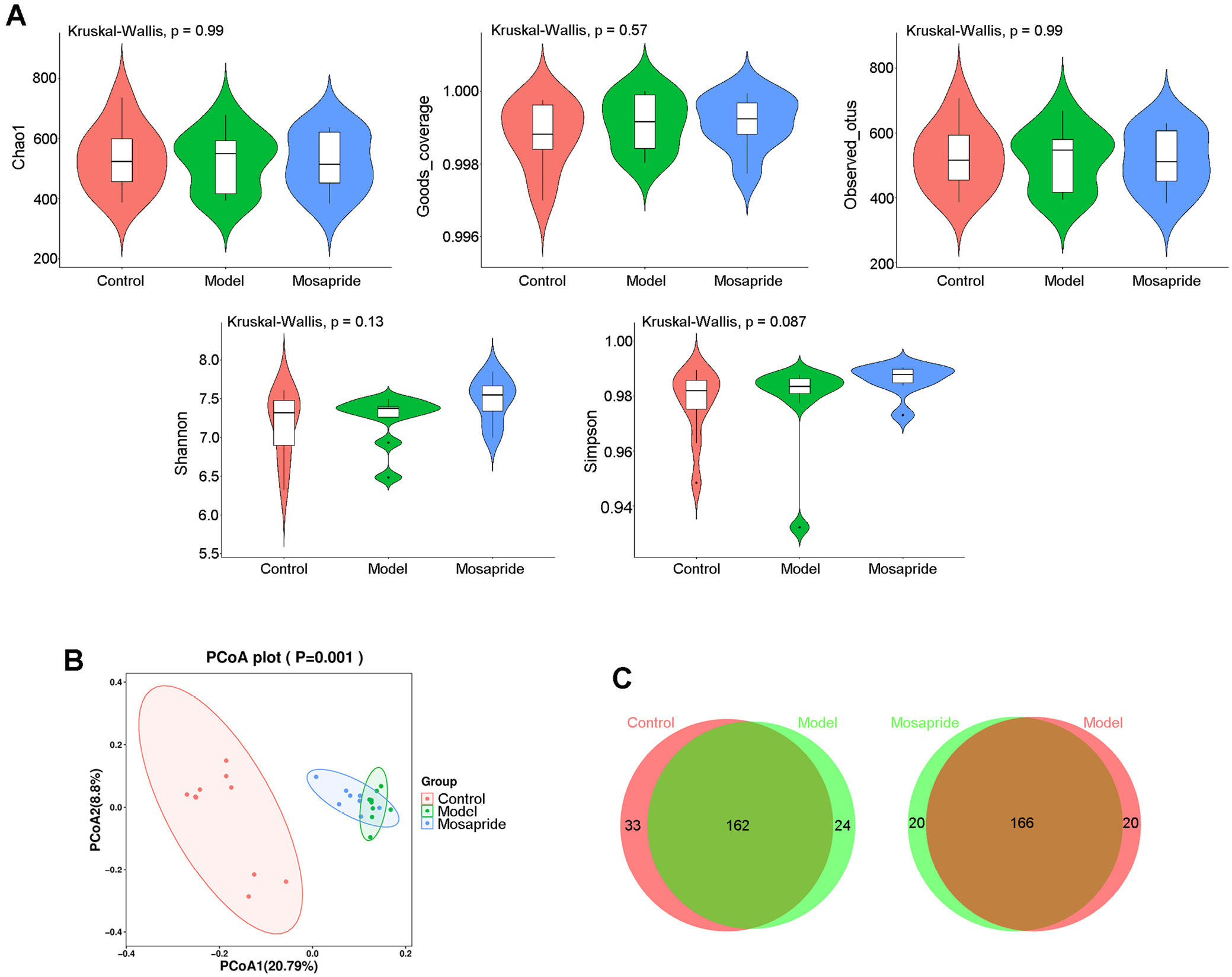
The alpha diversity analysis was carried out to determine the bacterial diversity at the genus level among the 3 groups. The violin plot showed no statistical differences in alpha diversity among the control, model and mosapride groups based on Chao1 (p = 0.99), Good’s coverage (p = 0.57), OTUs (p = 0.99), Shannon’s index (p = 0.13), and Simpson’s index (p = 0.087) (all p-values were calculated using Kruskal–Wallis test) (Figure 5A and Supplementary Information, File 2). This indicated that there was no statistical difference in alpha diversity of gut microbiota among the control group, the model group or the mosapride group. Unweighted UniFrac-based PCoA was conducted to evaluate beta diversity of the gut microbiota among the 3 groups. The PCoA plot showed that the constitution of the gut microbiota in the treatment group was significantly different compared with that of the control group and the model group, respectively (p = 0.001; ANOSIM) (Figure 5B). The unweighted UniFrac distance matrix was shown in Supplementary Information (File 3). Figure 5C showed the number of common and unique species between control and model groups or between model and mosapride groups. The specific genus was listed in Supplementary Information (File 4).
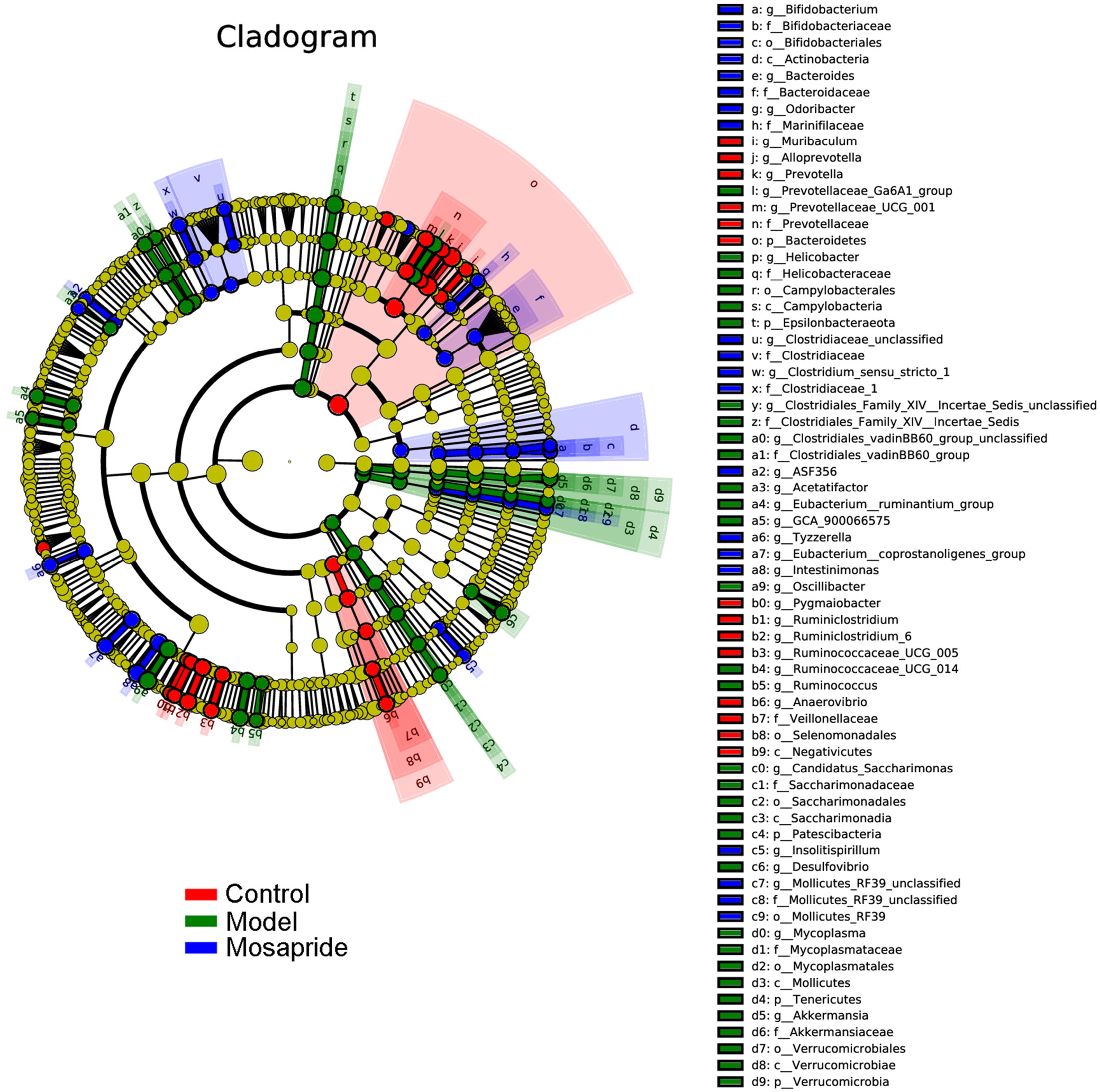
LEfSe analysis of gut microbiota in cirrhotic rats after mosapride treatment
The LEfSe analysis was conducted to identify bacteria that may be involved in the pathological changes of the lesions. The LEfSe cladogram showed the bacteria that played the most important roles in each group. As shown in Figure 6, 14 dominant bacteria were found in the control group, 32 dominant communities in the model group and 20 dominant communities in the mosapride group. In the control group, the crucial genera included Bacteroidetes, Prevotellaceae, Alloprevotella, Ruminiclostridium, Negativicutes, Selenomonadales, Veillonellaceae, and Anaerovibrio. In the model group, the closely related bacteria included Campylobacterales, Epsilonbacteraeota, Helicobacter, Oscillibacter, Prevotellaceae_Ga6A1_group, Verrucomicrobiales, and Akkermansia. In the mosapride group, the bacteria included Intestinimonas, Eubacterium, Clostridiaceae, Clostridium, Bacteroidaceae, Bacteroides, Tyzzerella, Actinobacteria, and Bifidobacteriales.
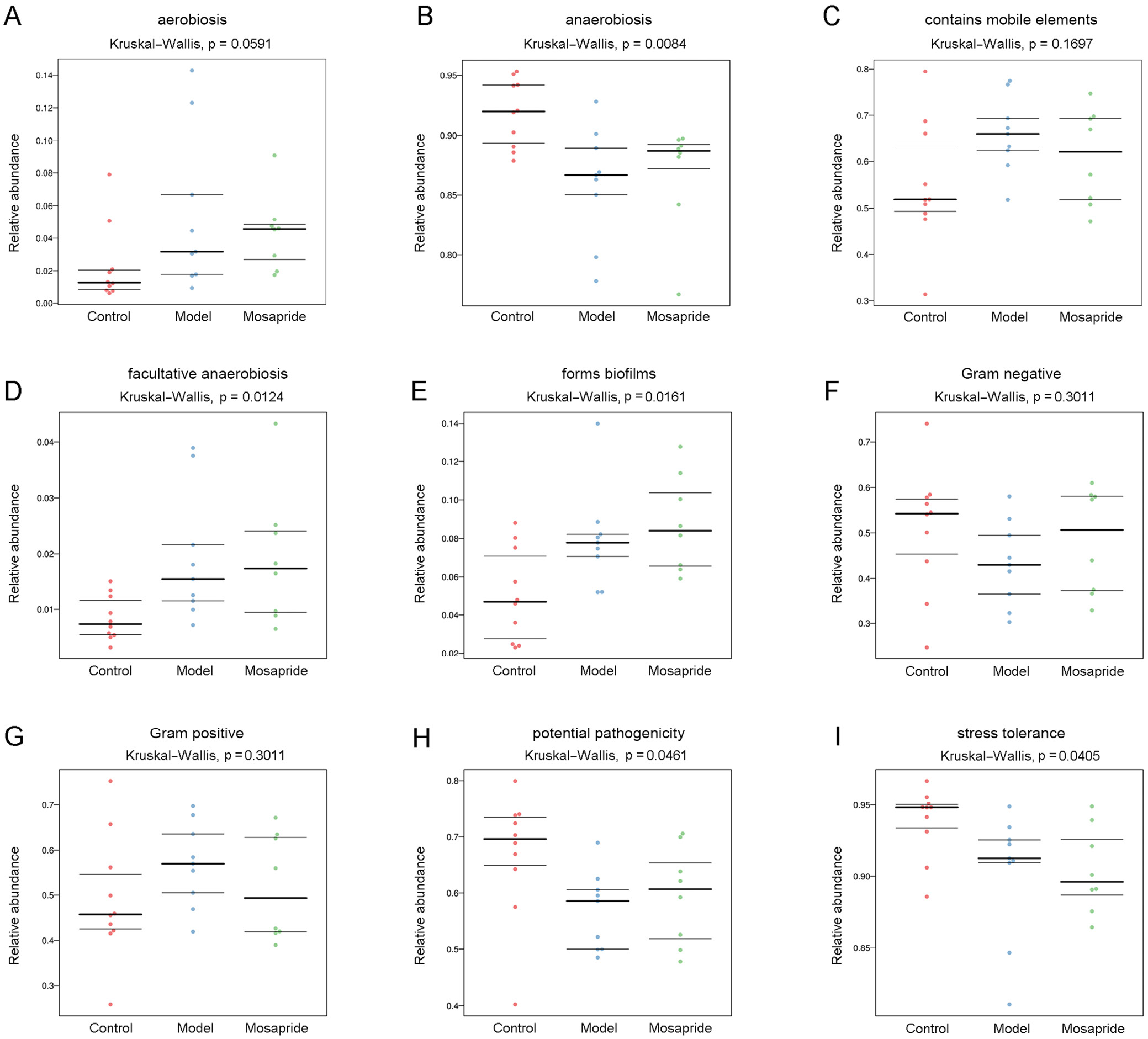
Prediction of bacterial phenotype associated with cirrhosis
Biologically interpretable phenotypes were predicted to investigate the association of cirrhosis and microbiota. Figure 7A–I showed that the differential microbiota may not be associated with aerobiosis (p = 0.0591; Kruskal–Wallis), and that it contains mobile elements (p = 0.1697; Kruskal–Wallis), Gram-negative phenotype (p = 0.3011; Kruskal–Wallis) and Gram-positive phenotype (p = 0.3011; Kruskal–Wallis). Cirrhosis-related phenotypes of bacteria included anaerobiosis (p = 0.0084; Kruskal–Wallis), facultative anaerobiosis (p = 0.0124; Kruskal–Wallis), formed biofilms (p = 0.0161; Kruskal–Wallis), potential pathogenicity (p = 0.0461; Kruskal–Wallis), and stress tolerance (p = 0.0405; Kruskal–Wallis). This finding may have implications for understanding the relationship between different microbial phenotypes and cirrhosis.
Discussion
In this study, we investigated the efficiency of mosapride in hepatic cirrhosis by illustrating the pathological changes and the intestinal microbiota changes after the administration of mosapride. In addition, we tried to discover the potential cause of the improvement of liver condition after the administration of mosapride. In our previous study, mosapride mediated bacterial translocation and endotoxin levels in rats. Besides, the attempts have been made to screen for potential bacteria involved in the translocation that may lead to increased permeability of the intestinal wall in vivo. Our data showed that mosapride could improve gut microbiota translocation, which then resulted in gut microbiota changes that may be associated with the pathogenesis of hepatic cirrhosis in rats.
Mosapride has been commonly utilized for treating liver diseases as it induces no significant adverse events.13 For instance, in nonalcoholic steatohepatitis mice, mosapride citrate could improve hepatic histological damage and colonic inflammation.27 Our previous studies have shown that mosapride could significantly reduce plasma endotoxin concentrations and reduce the incidence of bacterial translocation in cirrhosis.13 Additionally, mosapride did not improve hepatic and intestinal injuries. In this study, we tried to validate that mosapride did not change the hepatic morphology of rats. However, to some extent, it could attenuate intestinal injuries. This was not similar with our previous findings. Compared with the model group, there was a significant attenuation in intestinal mucosa injury after the mosapride treatment. Additionally, after the mosapride treatment, villus showed shortening together with partial rupture and distortion. There were no differences in the width of the villus on intestinal mucosa. Compared with the control group, there was a significant decline in villus width, mucosal thickness and intestinal wall thickness in the model group. In view of this, we investigated the potential mechanism that intestinal mucosa damage may associate with the change of intestinal microbiota.
In LEfSe analysis, we screened several bacteria that were associated with the pathological changes of intestinal injury, including Bacteroidetes, Prevotellaceae, Alloprevotella, Ruminiclostridium, Negativicutes, Selenomonadales, Veillonellaceae, Anaerovibrio, Campylobacterales, Epsilonbacteraeota, Helicobacter, Oscillibacter, Verrucomicrobiales, Akkermansia, Intestinimonas, Eubacterium, Tyzzebella, Clostridiaceae, Clostridium, Vacteroidaceae, Bacteroides, Tyzzerella, Actinobacteria, and Bifidobacteriales. The Oscillibacter genus has been reported to be associated with atrial fibrillation,28 stroke and transient ischemic attack,29 and human infection.30 However, no previous studies reported the roles of Oscillibacter genus in hepatic diseases. For Alloprevotella, genus has been reported to be less abundant in rheumatoid arthritis patients.31 In patients with chronic kidney disease, Alloprevotella were enriched in the fecal samples.32 A study focusing on screening of pathogens for oral cavity squamous cell cancer (OC-SCC) reported the enrichment of Alloprevotella.33 Similarly, Alloprevotella were abundant in patients with oral cancer.34 For Bacteroides, a beneficial gut Bacteroides-folate-liver pathway was demonstrated to regulate lipid metabolism.35 Ruminiclostridium has been closely involved in the obesity and metabolic dysfunction.36 Anaerovibrio are the key microbiota signature after quercetin treatment.37 Besides, they were considered to play a certain role in the evolution of gut microbiota following acute human immunodeficiency virus type 1 (HIV-1) infection.38 Prevotellaceae alternation was proposed to be involved in the protection of the central nervous system (CNS) autoimmunity and oral cancer.34, 39 The human gut bacteria Akkermansia are newly investigated to be associated with many healthy problems40 and their regional distribution is associated with metabolic syndromes.41 However, their roles in the pathogenesis of hepatic cirrhosis are not well defined. In this study, our data showed that the mosapride treatment would lead to significant changes in the gut microbiota in hepatic cirrhosis rats. In the future, further studies are required to investigate the exact roles of these microbiota.
After screening gut microbiota, the prediction of bacterial phenotypes was conducted. Mosapride may change bacterial phenotypes related with cirrhosis, and these phenotypes included anaerobiosis, facultative anaerobiosis, biofilm formation, potential pathogenicity, and stress tolerance. However, little is known about the exact roles of microbiota in the pathogenesis of liver diseases. In the future, more studies are required to investigate the correlation between bacterial phenotypes and the pathogenesis of hepatic cirrhosis.
Limitations
There are some limitations to our study. In this study, we could not find out which bacteria are the direct causes for the attenuation of intestinal lesions mediated by mosapride. As there are so many signaling pathways involved in the intestinal lesion and repairment, we could not illustrate the exact pathway affected by the intestinal microbiota after mosapride treatment. Besides, the sample size is another limitation in this study. We will focus on these limitations in our subsequent studies.
Conclusions
Our data showed that mosapride could modulate the gut microbiota changes and trigger the decrease of gut microbiota translation and attenuation of endotoxin concentrations. This may help to illustrate the pathogenesis of hepatic cirrhosis in rats.
Data availability statement
The original contributions presented in the study are included in the Supplementary Information containing:
File 1: Bacterial abundance and clustering analysis in control, model and mosapride rats.
File 2: Alpha diversity at the genus level was evaluated using Chao1, Good’s coverage, OTUs, Shannon’s, and Simpson’s indices.
File 3: The unweighted UniFrac distance matrix was used for evaluating beta diversity with principal coordinates analysis.
File 4: Abundance of the detected bacteria between control and model rats or between mosapride and model rats.
Further inquiries can be directed to the corresponding author. The Supplementary Information is available online at https://doi.org/10.5281/zenodo.6021342.