Abstract
Background. microRNAs are involved in a variety of physiological and pathophysiological processes, but their role in the pathogenesis of hypertrophic scars (HS) is not fully understood. Transforming growth factor β1 (TGF-β1) plays an important role in the genesis and development of HS.
Objectives. In this study, we hypothesized that a post-translational miRNA mechanism regulates the expression of TGF-β1 in HS fibroblasts (HSFBs) and participates in the development of HS.
Materials and methods. Predictions from EBCORI, PicTar and miRBase databases showed that miR-124-3p can target and regulate the expression of TGF-β1. We collected HS tissue and corresponding normal tissue from 25 patients with HS who had been operated on for the first time.
Results. The expression level of miR-124-3p in HS tissue was significantly lower than in normal tissue, while the expression level of TGF-β1 mRNA was significantly higher than in normal tissue (p < 0.05), showing a negative correlation between them. Results from a luciferase reporter assay showed that miR-124-3p targets the 3’-UTR of TGF-β1 and inhibits its expression. After miR-124-3p mimics were transfected into HSFBs, the expression of TGF-β1, α-smooth muscle actin (α-SMA), collagen I, survivin, and Bcl-2 were reduced and the expression of Bax was increased, with significant decreases in DNA synthesis, proliferation and survival. However, after a miR-124-3p inhibitor was transfected into HSFBs, these effects were reversed as the expression of TGF-β1, α-SMA, collagen I, survivin, and Bcl-2 increased, expression of Bax decreased, and DNA synthesis, proliferation and survival cells increased significantly.
Conclusions. miR-124-3p can inhibit the proliferation of HSFBs by targeting TGF-β1, and miR-124-3p may thus be a potential therapeutic target in HS.
Key words: TGF-β1, proliferation, fibroblasts, hypertrophic scarring, miR-124-3p
Background
Hypertrophic scar (HS) is a type of pathological scar, which is the result of local over-healing after dermal injury of human skin. The main pathological characteristics are excessive proliferation and differentiation of fibroblasts and enhanced deposition of collagen-based extracellular matrix (ECM) components.1 Histologically, HS contains myofibroblasts, which participate in the contraction of scars.2 These cells cause contraction of the surrounding ECM, resulting in an increase in the density of scar tissue, which not only affects the appearance and function of patients’ skin, but may also cause psychological trauma. Hypertrophic scar is one of the most difficult problems to solve in burn, plastic surgery and in the general field of wound repair. Because of the lack of effective treatment methods, HS treatment is very difficult and often requires repair surgery, which leads to the formation of HS again.3 At present, the mechanism of HS formation is not fully understood. Elucidating the molecular mechanism of HS will help contribute to the discovery of new therapeutic targets.
Hypertrophic scar is characterized by excessive proliferation of fibroblasts and aberrant ECM formation. It consists of collagen, fibronectin (FN) and aminoglycan. In normal fibroblasts, the synthesis and decomposition of ECM are in a dynamic equilibrium, which maintains the relative stability of ECM. In HS, this balance is impaired due to increased ECM synthesis, and deficiencies in its degradation and turnover. It has been shown that the amount of FN synthesized by HS fibroblasts in vitro is 4 times that of normal fibroblasts, and the amount of collagen synthesized is 3 times that of normal fibroblasts.4 The expression level and activity of collagenase in HS fibroblasts were significantly lower than that in normal fibroblasts.5
Scar formation can be divided into 3 different stages: inflammation, hyperplasia and remodeling.6 In the inflammatory stage, a large number of chemokines and inflammatory factors are produced, which can induce angiogenesis, re-epithelialization, fibroblast recruitment, proliferation, and ECM deposition. The balance between the recruitment and proliferation of fibroblasts and the production and degradation of ECM is mainly regulated by fibroblast growth factors, including insulin-like growth factor, platelet-derived growth factor and transforming growth factor β (TGF-β). Among these growth factors, TGF-β1 is widely reported to be related to the formation of HS, and mediates fibroblast proliferation, collagen production, ECM deposition, and myofibroblast differentiation during wound healing. In HS fibroblasts, TGF-β1 has been shown to increase the phosphorylation of Smad2/3 and Smad4 and decrease Smad7 phosphorylation, resulting in the over-accumulation of Collagen-1 (Col-1).11 In addition, TGF-β1 can induce collagen production and contraction of human skin fibroblasts (HSFBs) derived from HS.12 This suggests that TGF-β1 may be a key target for the development of new strategies for HS treatment. The TGF-β 1 can induce macrophage M2 polarization in the process of damage repair, making M2 macrophages a “double-edged sword.” On the one hand, they are essential for tissue repair, but on the other hand, they are also potential mediators of fibrosis and scar formation.13 Studies show that inhibition of macrophage M2 polarization can inhibit fibroblast activation and the formation of scars.14, 15
MicroRNAs (miRNA) are endogenous non-coding RNAs with regulatory function found in eukaryotes. They are involved in the regulation of cellular processes such as cell proliferation, differentiation and apoptosis. They can also aggravate or inhibit the onset and development of various diseases, including inflammation, cancer and fibrosis.18 Through bioinformatic approach, we predicted that miR-124-3p may target the 3’-UTR of TGF-β1 to inhibit its expression. Recent reports show that miR-124-3p is considered to be a tumor suppressor in some cancers, including hepatocellular carcinoma,19 cervical cancer20 and gastric cancer.21 Therefore, it is speculated that miRNAs may posttranscriptionally regulate the expression of TGF-β1 in HS.
Objectives
In this study, we evaluated the expression levels of miR-124-3p in HS and normal skin tissues and found that miR-124-3p was downregulated in HS and was negatively correlated with TGF-β1 expression. We found that modulating expression of miR-124-3p in HSFBs could affect cell proliferation.
Materials and methods
Samples
From January 2018 to January 2020, tissue samples from HS patients and corresponding normal tissues (NC) were collected from the plastic surgery department. Fresh tissue was stored directly in liquid nitrogen. Written informed consent was obtained from all patients. This study was approved by the ethics committee of Zhejiang Tongde Hospital (Hangzhou, China).
Cell culture and transfection
Human skin fibroblasts were prepared according to a previously reported method.22 The human skin fibroblast cell line HSF2 was purchased from National Biomedical Experimental Cell Resource Bank (Beijing, China). They were cultured with high sugar Dulbecco’s modified Eagle’s medium DMEM (Gibco, Carlsbad, USA) containing 10% fetal bovine serum (FBS; Gibco), 100 U/mL penicillin G, and 100 U/mL streptomycin sulfate at 37°C and 5% CO2.
The TGF-β1 expression vector pEX-3-TGF-β1, pGPU6-TGF-β1, miR-124-3p mimic, and miR-124-3p inhibitor were purchased from Gene Pharma (Shanghai, China). They were transfected into HSFBs and HSF2 cells. All transfections were performed with Lipofectamine 3000 Reagent (Life Technologies, Carlsbad, USA) according to the manufacturer’s protocol. pEX3 and pGPU6 empty vectors were used as negative controls.
EdU incorporation assay
EdU (5-ethynyl-2’-deoxyuridine) is a thymidine analogue, which can get incorporated into synthetic DNA molecules instead of thymine (T) during DNA replication. The ethynyl on EdU can covalently react with fluorescently labeled small molecule azide probes (such as Azide Alexa Fluor 488, Azide Alexa Fluor 555, Azide Alexa Fluor 594, and Azide Alexa Fluor 647) through the catalysis of a univalent copper ion, forming a stable triazole ring so that the newly synthesized DNA can be labeled by the corresponding fluorescent probe, allowing for the detection of proliferating cells using appropriate fluorescence detection equipment.
In order to detect DNA synthesis in different groups of cells, 2 × 104 cells were added into each well of a 24-well plate to which EdU was added at a final concentration of 20 μm; the cells were cultured for 24 h. The cells were washed twice with pre-chilled PBS and fixed for 20 min with pre-chilled 4% paraformaldehyde. They were treated with 0.5% Triton X-100 and stained with Cell-LightTM EdU In Vitro Imaging kit (Shanghai Beyotime Biotechnology Co., Ltd., Shanghai, China) according to the manufacturer’s protocol. Nuclei of cells were stained with DAPI (Shanghai Beyotime Biotechnology), and EdU-positive cells were observed with an inverted immunofluorescence microscope.
Cell viability assay
Cells were harvested (from tissue), suspended in serum-free medium and plated into 96-well plates (1 × 104/well, 5 wells in each group). The cells were cultured in an incubator at 37°C and 5% CO2 for 12 h. The MTT (10 μL, 5 mg/mL; Sigma-Aldrich, St. Louis, USA) was added into each well at 0 h, 24 h, 48 h, and 72 h respectively, and cultured for 4 h. After incubation, the culture media was discarded and dimethyl sulfoxide (DMSO; 200 μL/well) was added to the wells. The plates were placed on a shaker for 5–10 min at room temperature in the dark. Absorbance values at 570 nm were measured with a Microplate Reader (Multiskan MK3; Thermo Fisher Scientific, Waltham, USA). The experiment was repeated 3 times.
Cell apoptosis detection by flow cytometry
Apoptosis analysis was performed using the Annexin V-FITC Analysis Kit (Shanghai Beyotime Biotechnology) according to the manufacturer’s protocol. Cells were collected, digested with trypsin without EDTA and washed 3 times with ice-cold phosphate-buffered saline (PBS) and centrifugation. Cells (1 × 106) were collected and resuspended in 300 μL of 1X binding buffer (Thermo Fisher Scientific). Annexin V-FITC (195 μL) and PI-PE (5 μL) were added to the cells according to the manufacturer’s protocol. After 10 min of incubation at 4°C in the dark, pre-chilled 1X binding buffer (200 μL) was added. Apoptosis was detected using flow cytometry. The results were analyzed using CELLQUEST software (BD Biosciences, Franklin Lakes, USA). The experiment was repeated 3 times.
Double luciferase reporter gene analysis
In order to examine the potential interaction between miR-124-3p and the 3’-UTR of TGF-β1, we constructed wild-type (wt-pGL3-TGF-β1) and mutant (mut-pGL3-TGF-β1) luciferase reporter genes based on the predicted binding site. The constructs were co-transfected with miR-124-3p and Renilla luciferase into HEK293 cells. The cells were lysed using the Dual-Luciferase Reporter Assay System (Promega, Madison, USA) according to the instructions after culture for 48 h. The results were detected using Panomics Luminometer (Affymetrix, Santa Clara, USA) after the luminescence was added. The sea renin fluorescence was used as an internal reference.
RNA extraction and qRT-PCR
Total RNA was extracted using Trizol reagent (Invitrogen, Carlsbad, USA) according to the manufacturer’s protocol. RNA concentration and purity were determined using a NanoDrop™ 1000 spectrophotometer (Thermo Fisher Scientific). Total RNA (1 μg) was subjected to reverse transcription using miScript II RT Kit (Qiagen, Hilden, Germany). Real-time polymerase chain reaction (RT-PCR) was performed using Taqman Universal Mix II No UNG (Thermo Fisher Scientific), PCR primers, miScript SYBR® Green PCR Kit (Qiagen), and the ABI StepOne Plus system (Applied Biosystems, Waltham, USA). At the end of each reaction, a melting curve analysis was performed to confirm the absence of primer dimers. The glyceraldehyde 3-phosphate dehydrogenase (GAPDH) gene was used as an internal control for normalization of RNA quantity and quality differences in the samples. Quantification of target gene expression was performed using the2−ΔΔCt method. Primer sequences are listed in Table 1.
Western blotting detection
Cells were harvested and lysed with Cell Lysis Solution (Sigma-Aldrich). The supernatant was collected after centrifugation at 4°C (10,000 rpm) for 5 min. Total proteins were extracted and protein concentration was determined using the bicinchoninic acid assay (BCA) assay. Proteins (50 μg per lane) were separated using 12% SDS-PAGE. Proteins were then electrotransferred to a polyvinylidene fluoride (PVDF) membrane (Amersham Biosciences, Piscataway, USA). The PVDF membrane was rinsed with TBS for 10–15 min and placed in TBS/T blocking buffer containing 5% (w/v) skimmed milk powder. Then, it was incubated at 4°C overnight following the addition of an appropriate dilution of primary antibodies (1:2000 α-smooth muscle actin (α-SMA); 1:1000 collagen I; 1:2000 survivin; 1:2000 Bcl-2; 1:2000 Bax; 1:5000 GAPDH; all from Abcam, Cambridge, UK). The membrane was then rinsed with TBS-Tween 20 (TBST) 3 times and incubated with a horseradish peroxidase (HRP)-labeled goat anti-mouse IgG secondary antibody (1:50,000; Abcam) at room temperature for 1 h. After incubation, the membrane was rinsed 3 times with TBST. Protein bands were detected using an enhanced chemiluminescence kit (Perkin-Elmer, Waltham, USA) and quantified as a ratio to GAPDH. Quantification was performed using Imagequant LAS4000 (GE Healthcare, Tokyo, Japan).
Statistical analysis
Statistical analysis was performed using SPSS v. 17.0 software (SPSS Inc., Chicago, USA). The data are expressed as mean ± standard deviation (SD). The 2 groups of variables, which were comparable with F-test, were compared with unpaired Student’s t-test. The mean values of multiple groups were compared using one-way analysis of variance (ANOVA). The correlation of the 2 genes was examined with Spearman correlation test. A p-value <0.05 was considered to be statistically significant.
Results
miR-124-3p expression was downregulated and negatively correlated with TGF-β1 mRNA expression in HS
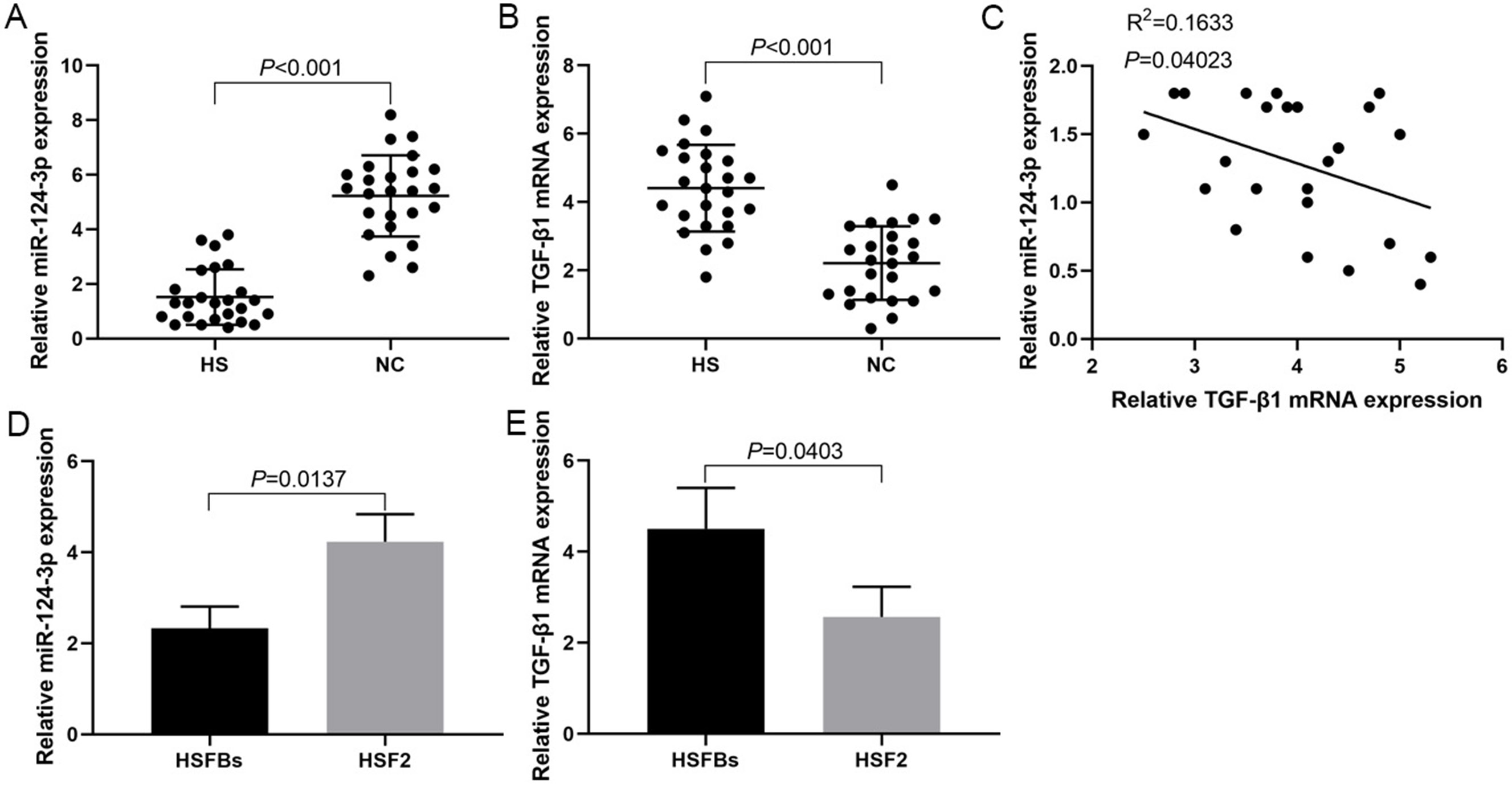
The RT-PCR results showed that the expression levels of miR-124-3p in HS tissues were significantly lower than that in NC tissue (Figure 1A), while the expression levels of TGF-β1 mRNA in HS tissues were significantly higher than in NC tissue (Figure 1B), and there was a negative correlation between them (Figure 1C). The miR-124-3p expression in HSFBs was significantly lower than in HSF2 cells (Figure 1D), while TGF-β1 mRNA expression in HSFBs was significantly higher than in HSF2 (Figure 1E).
miR-124-3p inhibits HSFB cell proliferation
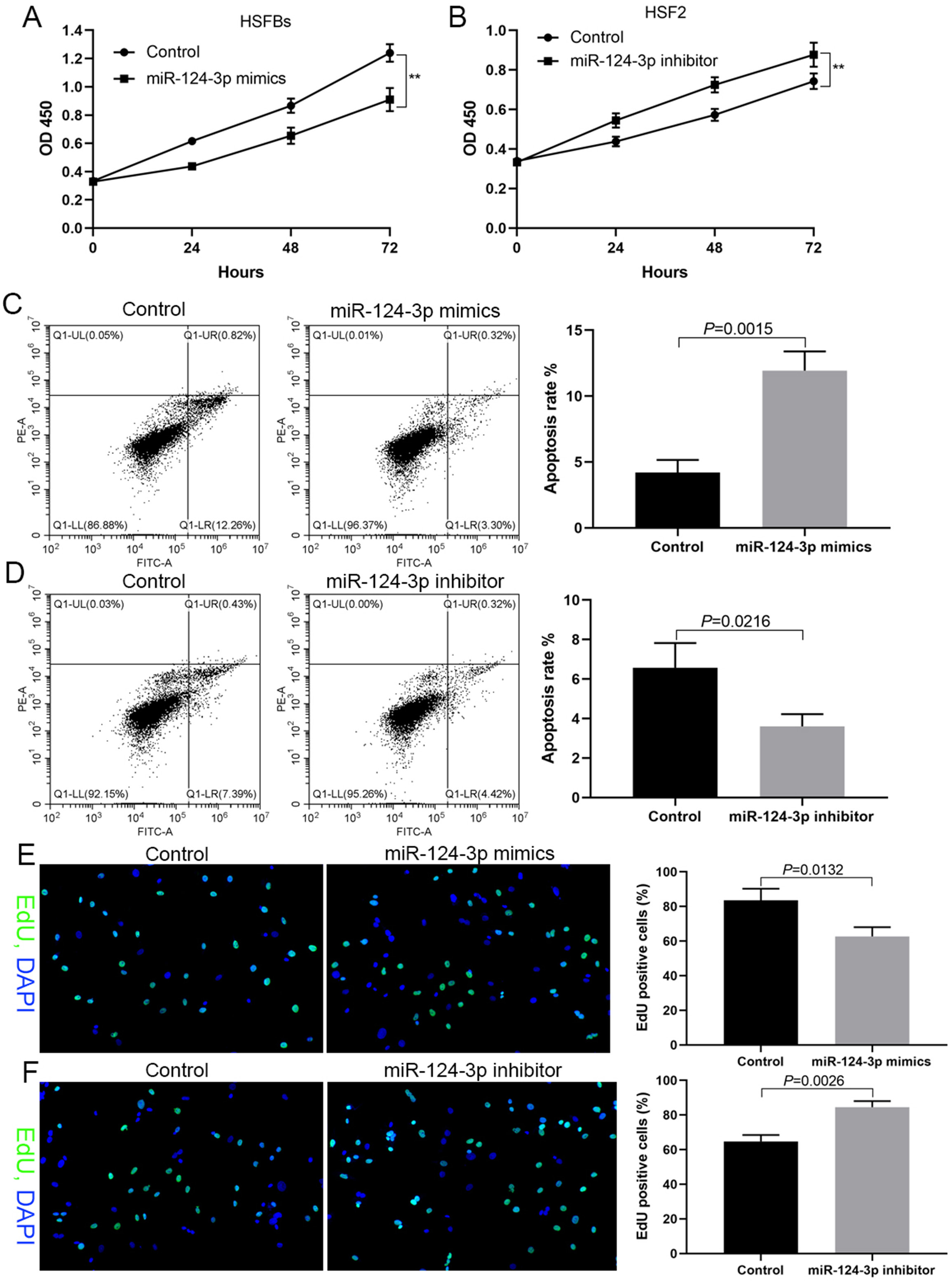
In HSFBs with low expression of miR-124-3p, DNA synthesis, cell proliferation and cell survival significantly decreased after being transfected with miR-124-3p mimics (Figure 2A,C,E). However, in HSF2 cells with high expression of miR-124-3p, DNA synthesis, cell proliferation and cell survival significantly increased after transfection transfected with a miR-124-3p inhibitor (Figure 2B,D,F).
miR-124-3p can target the 3’-UTR of TGF-β1 and inhibit TGF-β1 expression
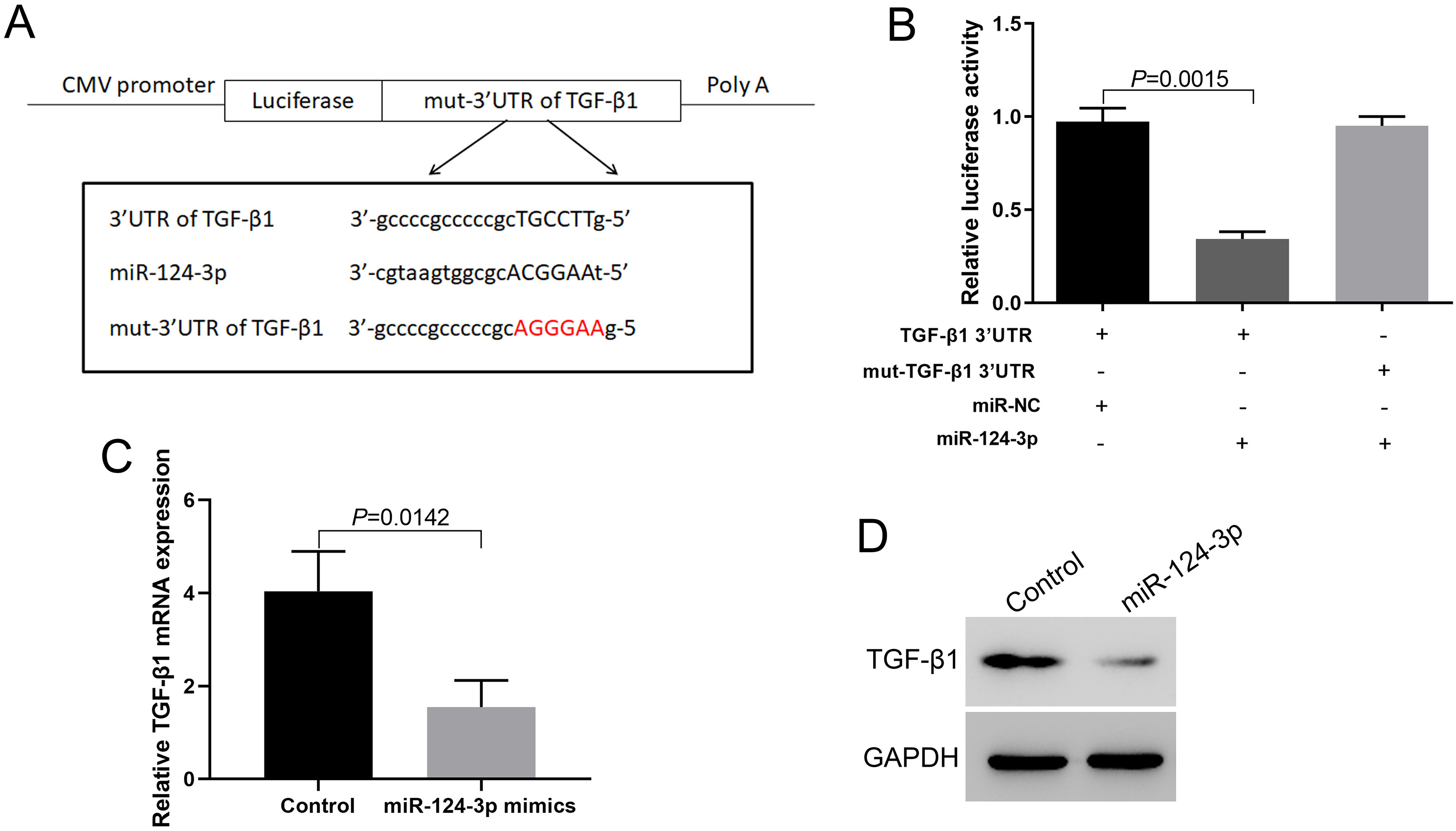
Bioinformatics prediction showed that miR-124-3p can target the 3’-UTR of TGF-β1 (Figure 3A). The double-luciferase reporter gene assay showed that miR-124-3p could target the 3’-UTR of TGF-β1 to inhibit the expression of luciferase. When the 3’-UTR of TGF-β1 was mutated, the inhibition of miR-124-3p was eliminated (Figure 3B). RT-PCR and western blotting results showed that miR-124-3p can directly downregulate TGF-β1 expression (Figure 3C,D).
miR-124-3p affected the proliferation of HSFBs by regulating TGF-β1 expression
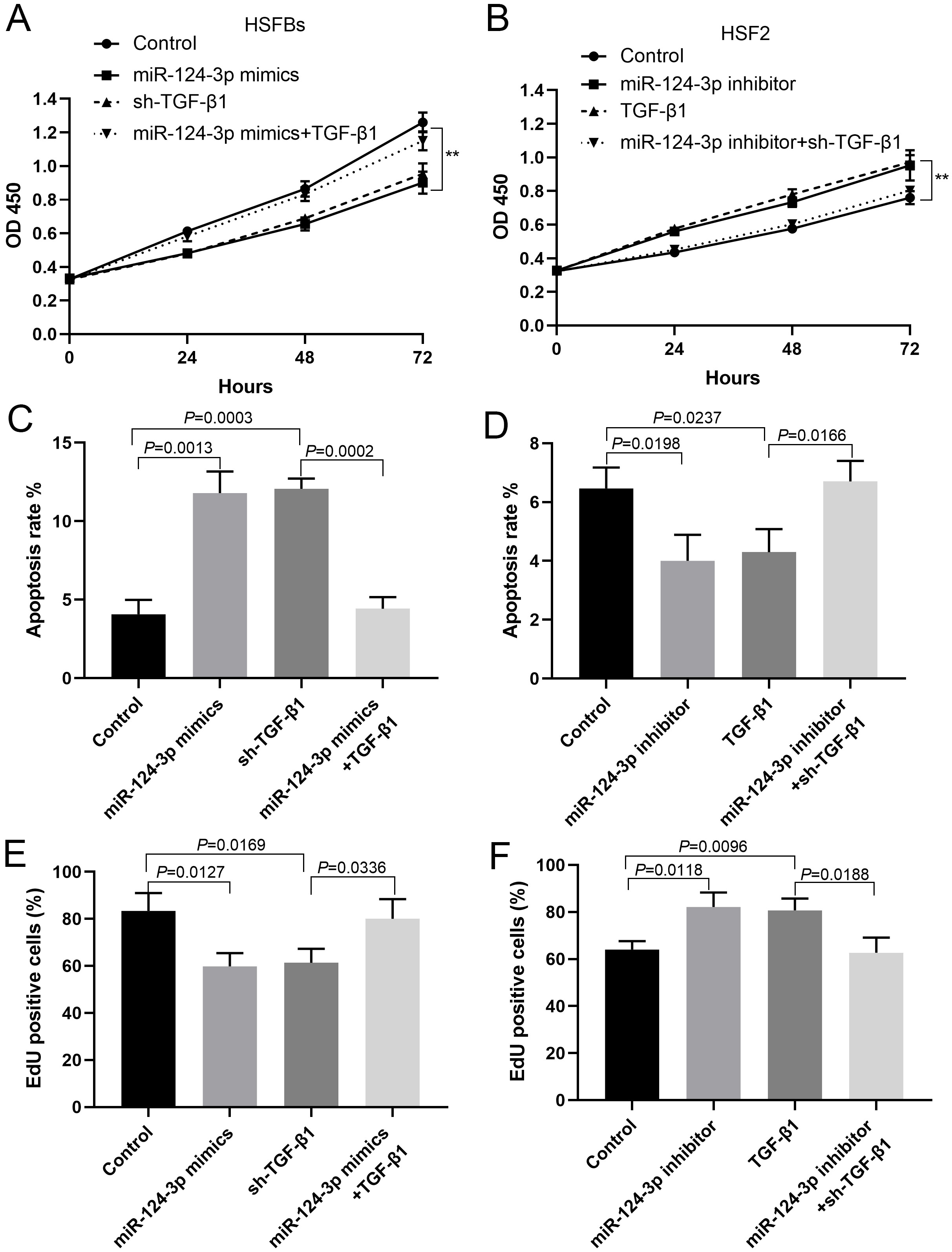
In HSFBs transfected with miR-124-3p mimics or shRNA against TGF-β1, cell proliferation was inhibited, while in HSFBs co-transfected with miR-124-3p mimics and TGF-β1, cell proliferation was restored (Figure 4A,C,E). In HSF2 transfected with miR-124-3p inhibitor or TGF-β1, cell proliferation was promoted, while in HSF2 co-transfected with miR-124-3p inhibitor and TGF-β1 shRNA, cell proliferation was inhibited (Figure 4B,D,F).
miR-124-3p affects the expression of downstream effectors related
to HS formation
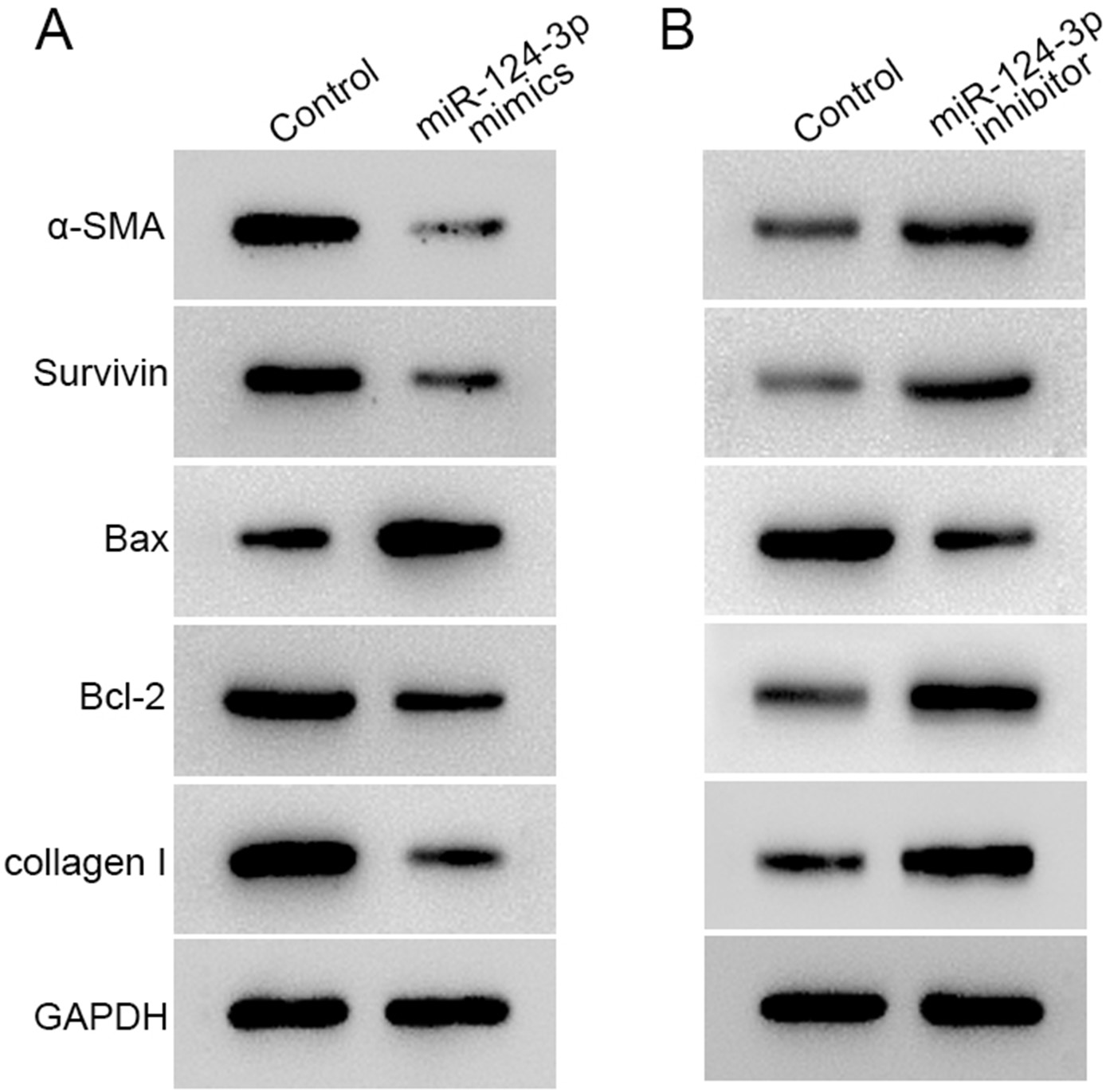
The expression of α-SMA, collagen I, survivin, and Bcl-2 was inhibited, and Bax expression increased after transfection of miR-124-3p mimics into HSFBs (Figure 5A). However, the expression of α-SMA, collagen I, survivin, and Bcl-2 was promoted, and Bax expression was inhibited with a miR-124-3p inhibitor (Figure 5B).
Discussion
miRNA is a type of endogenous, non-coding, small molecule RNA that has generally 19–22 nucleotides in length and is involved in the regulation of gene expression at the post-transcriptional level. It has been shown that miRNA can regulate the development of HS in many ways, including through the TGF-β/Smad signaling pathway,23 ECM synthesis and degradation,24 proliferation and differentiation of HSFBs, and epithelial–mesenchymal transition (EMT).25 miR-140-5p, miR-23b, miR-let-7b, and miR-153 can all regulate the TGF-β/Smad signaling pathway including TGF-β type I and type II receptors (RI and RII, respectively): miR-140-5p can negatively regulate the expression of TGF-βRII, while miR-23b and miR-let-7b can upregulate TGF-βRII expression. miR-17-5p and miR-20 are also closely related to HS through their regulation of TGF-βRII expression.26 miR-21 and miR-503 can also act on Smad7, a negative regulator of TGF-β/Smad signaling.26 miR-29b can also act on the 3’-UTR of collagen I and inhibit its expression directly.27 miR-10a and miR-181 can target PAI-1 and UPA to regulate the expression of collagen I in HS.28 In HSFBs, the expression of miR-196a is significantly decreased, resulting in increased expression of collagen I and III and promotion of scar formation.29 miR-146a inhibits the differentiation of HSFBs by acting on Smad4.30 The miR-200 family can affect EMT in HSFBs by regulating the expression of ZEB1 and ZEB2.31
The miR-124 family of miRNAs is highly expressed in differentiated and mature neurons. They were first found in large numbers in the mouse brain, accounting for 5–48% of total miRNAs in the organ.32 It was found that miR-124-3p belonged to the miR-124 family, members of which not only participate in brain development and nerve function, but also as tumor suppressors in some cancers. For example, miR-124-3p can target BCL2L11 to inhibit cell proliferation in angiomyolipoma cells,33 and in gastric cancer, miR-124-3p can target Rac1 and SP1 to inhibit cell proliferation and could be used as an independent indicator of survival and treatment strategies.20 In hepatocellular carcinoma, the long non-coding RNA (lncRNA) lncRNA MALAT1 can act as a molecular sponge to absorb miR-124-3p to inhibit its effects and upregulate expression of the EMT marker Slug to promote tumor metastasis.34 Aberrant cell proliferation similar to tumors is observed in HS. Whether miR-124-3p is involved in regulating HSFB cell proliferation, has not yet been reported.
In this study, we found that expression levels of miR-124-3p in HS tissue was significantly lower than that in corresponding normal skin tissue, and miR-124-3p in HSFBs was also significantly lower than that in HSF2 cells. miR-124-3p targeted the 3’-UTR of TGF-β1 to inhibit its expression. The expression of TGF-β1 in HS tissue was significantly higher than that in corresponding normal skin tissue, and the expression levels of miR-124-3p was negatively correlated with TGF-β1 expression.
The TGF-β1 has a variety of biological functions. After skin injury, high amounts of TGF-β1 are released to increase collagen and ECM deposition.35 The TGF-β1 can induce collagen production and contraction of fibroblasts in HS. It can also upregulate the expression of α-SMA and promote the differentiation of normal fibroblasts into myofibroblasts.36 In a study by Shah et al., it was found that neutralization of TGF-β1 could reduce scar formation in rats, which provided direct evidence of a role for TGF-β1 in the formation of scars.37 Loiselle et al. found that regulation of the TGF-β signaling pathway prevents scar formation during flexor tendon repair.38 Fibroblasts are the main participants in wound repair, being involved in the formation of granulation tissue, collagen synthesis and interaction with the ECM to promote scar hyperplasia. In the process of HS formation, the proliferation of fibroblasts promotes the secretion of collagen, resulting in a large amount of collagen deposition. At the same time, fibroblasts also release a lot of growth factors to promote scar hyperplasia. Therefore, the uncontrolled proliferation of fibroblasts is the basis of scar formation.
Limitations
The main limitation of this study is the small sample size. More clinical samples need to be analyzed to clarify the clinical significance of miR-124-3p in hypertrophic scarring.
Conclusions
In this study, we found that transfection of miR-124-3p mimics in HSFBs inhibit the expression of TGF-β1, α-SMA, collagen I, survivin, and Bcl-2, while promoting Bax expression. However, inhibition of miR-124-3p in normal human skin fibroblast cells (HSF2) promotes the expression of TGF-β1, α-SMA, collagen I, survivin, and Bcl-2, and inhibits Bax expression. These results suggested that in HS, miR-124-3p may target TGF-β1 to inhibit the expression of anti-apoptotic proteins such as survivin and Bcl-2, and promote the expression of the pro-apoptotic protein Bax. The transformation of fibroblasts into myofibroblasts is one of the important mechanisms of scar formation. There are a large number of myofibroblasts in HS tissue; α-SMA is a myofibroblast marker.39 Myofibroblasts increase the deposition of ECM. The number and function of myofibroblasts determine the speed and degree of scar formation. Inhibition of fibroblast to myofibroblast transformation is an important strategy in preventing scar tissue development. This study showed that miR-124-3p inhibits the expression of α-SMA and collagen I, suggesting that miR-124-3p plays an important role in TGF-β1-induced differentiation of HS fibroblasts. In summary, we found that miR-124-3p inhibited the proliferation of HSFBs by inhibiting TGF-β1 expression, which suggests that miR-124-3p may be a potential therapeutic target in HS.