Abstract
Background. A growing number of studies have shown that long-chain non-coding RNA (lncRNA) plays an important role in the progression of non-small cell lung cancer (NSCLC).
Objectives. To explore the role and potential molecular mechanism of lncRNA PSMA3-AS1 in promoting the proliferation, migration and invasion of NSCLC.
Materials and methods. The expression of PSMA3-AS1, miR-17-5p and PD-L1 in a human bronchial epithelial cell line, BEAS-2B, and NSCLC cell lines, H226 and A549, were detected with quantitative real-time polymerase chain reaction (qRT-PCR) or western blot. The PSMA3-AS1 shRNA transfection was used to reduce the expression of PSMA3-AS1. Double fluorescent enzyme reporting was used to detect the relationship between PSMA3-AS1, miR-17-5p and PD-L1. Cell Counting Kit-8 (CCK-8), wound-healing and transwell assays, as well as western blot, were used to detect the expression of proliferation, migration, invasion, and epithelial–mesenchymal transition (EMT)-related proteins in lung cancer cells.
Results. The expression of PSMA3-AS1 in NSCLC cells was significantly higher than in human bronchial epithelial cells. The PSMA3-AS1 knockdown significantly reduced the proliferation, migration and invasion of lung cancer cells. In addition, double fluorescent enzyme results showed that PSMA3-AS1 could competitively bind miR-17-5p to PD-L1. The expression of miR-17-5p is low in lung cancer cells, while the expression of PD-L1 in them is high. Overexpression of PD-L1 reversed the inhibitory effect of PSMA3-AS1 knockdown on the proliferation, migration and invasion of lung cancer cells.
Conclusions. Generally speaking, PSMA3-AS1 is highly expressed in NSCLC. The PSMA3-AS1 can promote the proliferation, migration and invasion of NSCLC cells by regulating miR-17-5p/PD-L1.
Key words: non-small cell lung cancer, proliferation, miR-17-5p, invasion, lncRNA PSMA3-AS1
Background
Non-small cell lung cancer (NSCLC) is a common pathological type of lung cancer, accounting for about 80% of all cases.1 At present, the cause of NSCLC is often not completely clear and, due to concealment of NSCLC diagnoses by patients and atypical early clinical features, the early diagnosis rate of NSCLC is low and most patients have metastasis with poor overall prognosis.2 Therefore, further in-depth study of lung cancer is of great importance for treating lung cancer and improving the prognosis of patients.
Long chain non-coding RNA (lncRNA) is a non-coding RNA with a length of more than 200 bp that does not encode a protein. It can regulate gene expression through complex molecular mechanisms at many cellular levels, but the specifics of the mechanisms are not clear.3 Recent studies have shown that lncRNA, as a regulatory factor, participates in almost all cellular processes and plays a role in promoting or inhibiting cancer.4, 5 For example, high levels of expression of lncRNA PART1 was found to significantly accelerate the occurrence of NSCLC cancer.6 The lncRNA BRCAT54 overexpression promoted proliferation and migration, and activated apoptosis in NSCLC cells by binding RPS9.7 The lncRNA PSMA3-AS1 is located on chromosome 14 at position 14q23.1 and is overexpressed in esophageal cancer and glioma.8, 9 However, the study of PSMA3-AS1 in NSCLC is currently less pressing.
MicroRNAs (miRNAs) are a type of endogenous non-coding RNA with 19–25 nucleotides. They affect cell proliferation, differentiation, migration, and apoptosis by inhibiting the mRNA level of their target gene.10 MicroRNA is widely reported to be abnormally expressed in NSCLC.11 Bioinformatic analysis showed that PSMA3-AS1 could target and bind miR-17-5p, and that the expression level of miR-17-5p decreased significantly in lung cancer.12 Therefore, we speculate that PSMA3-AS1 can affect the progression of NSCLC by targeting miR-17-5p.
Objectives
This study aimed to primarily analyze whether PSMA3-AS1 targets miR-17-5p and affects biological processes in NSCLC cells, which could provide a new theoretical basis for clinical targeted therapy in NSCLC.
Materials and methods
Cell culture
BEAS-2B cells and NSCLC cell lines (H226 and A549, respectively; Shanghai Cell Bank of Chinese Academy of Sciences, Shanghai, China) were cultured in Dulbecco’s modified Eagle’s medium (DMEM; Thermo Fisher Scientific, Waltham, USA) containing 10% fetal bovine serum (FBS; Gibco, Waltham, USA), and subcultured in 5% CO2 and 37°C incubator. Follow-up experiments were carried out on the cells in the logarithmic phase.
Cell transfection
The shRNA targeting PSMA3-AS1 (sh-PSMA3-AS1), shRNA negative control (sh-NC), the PSMA3-AS1 overexpression plasmid (pcDNA-PSMA3-AS1), the negative control (pcDNA-NC), miR-17-5p mimic, mimic negative control (miR-NC), the PD-L1 overexpression plasmid (OE-PD-L1), and the negative control (OE-NC) were provided from the GenePharma Co., Ltd (Shanghai, China). The Lipofectamine 2000 transfection reagent (Invitrogen, Waltham, USA) was used to transfect the above plasmids into A549 cells.
qRT-PCR analysis
Total RNA was extracted from cells and synthesized into the cDNA according to a first strand cDNA synthesis kit (Roche Diagnostics, Basel, Switzerland). Quantitative real-time polymerase chain reaction (qRT-PCR) was performed according to the SYBR Green Fast qPCR Mix protocol (Beyotime, Shanghai, China). The 2−ΔΔCt method was used to analyze the expression of PSMA3-AS1 and miR-17-5p.
CCK-8 assay
Cells were digested with trypsin and inoculated in 96-well plates at a density of 5 × 105 cells/mL. Routine culture proceeded for 12 h. At the end of culture, Cell Counting Kit-8 (CCK-8) solution (10 μL; Beyotime) was added to each hole of the plates at 24 h, 48 h and 72 h. After 4 h of culture, the absorbance of each hole at 450 nm was detected using an automatic enzyme labeling instrument (Thermo Fisher Scientific).
Wound-healing assay
Cells were inoculated in 12-well plates at a density of 5 × 104 cells/mL. When the degree of cell fusion was more than 95%, a sterile 200-μL liquid transfer gun was used to cross the surface of the culture plate vertically. After 24 h of routine culture, the scratches were photographed using an optical microscope (Eclipse 80i; Nikon Corp., Tokyo, Japan).
Transwell assay
Cells were inoculated in the upper chamber (Corning, Inc., Corning, USA) and smeared with Matrigel glue (Becton Dickinson Biosciences, Franklin Lakes, USA) at a density of 5 × 105 cells/mL; the lower chamber contained 500 μL of DMEM complete medium. After 24 h of routine culture, the cells were washed with phosphate-buffered saline (PBS) twice and fixed with paraformaldehyde for 20 min. Then, the uninvaded cells in the upper chamber were gently wiped with a cotton swab. After crystal violet staining for 5 min, the number of invaded cells in each group was observed and counted using a light microscope (Eclipse 80i; Nikon).
Western blot analysis
Total cellular proteins were extracted from RIPA cell lysate (Beyotime), and the concentration was determined using BCA kit (Bio-Rad, Hercules, USA). The sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) gel electrophoresis was used to separate the proteins which were transferred to polyvinylidene difluoride (PVDF) membrane. Membranes were sealed with 5% skim milk at room temperature, incubated with 1:1000 primary antibodies E-cadherin (ab15148; Abcam, Cambridge, UK), N-cadherin (ab12221; Abcam), vimentin (ab45939; Abcam), matrix metalloproteinase (MMP)-2 (ab92536; Abcam), MMP-9 (ab137867; Abcam), and β-actin (ab8226; Abcam) at 4°C for overnight, and then incubated with 1:4000 second antibody anti-rabbit horseradish peroxidase (HRP)-conjugated IgG for another 2 h. The immune bands were visualized using enhanced electrochemiluminescence (ECL) photoluminescence solution (Wanleibio Co., Ltd., Shenyang, China).
Luciferase reporter assay
Wild-type (WT) and mutant (MUT) reporter plasmids were constructed based on the sequences of PSMA3-AS1 3’-UTR and the PD-L1 3’-UTR. PSMA3-AS1-WT/Mut or PD-L1-WT/Mut were co-transfected into HEK293T cells with miR-NC and miR-17-5p mimic, respectively. The luciferase activity in the transfected cells was tested using the dual luciferase assay kit (KenReal, Shanghai, China) after 48 h.
Statistical analyses
Experimental data are presented as mean ± standard deviation (SD), and the statistical analyses were conducted using GraphPad Prism v. 8 software (GraphPad Software, San Diego, USA). The comparison of differences between 2 groups was carried out with Student’s t-test and between multiple groups with one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test. The differences were statistically significant at p < 0.05.
Results
Knockdown of PSMA3-AS1 significantly inhibited the proliferation, migration, invasion, and EMT process in A549 cells
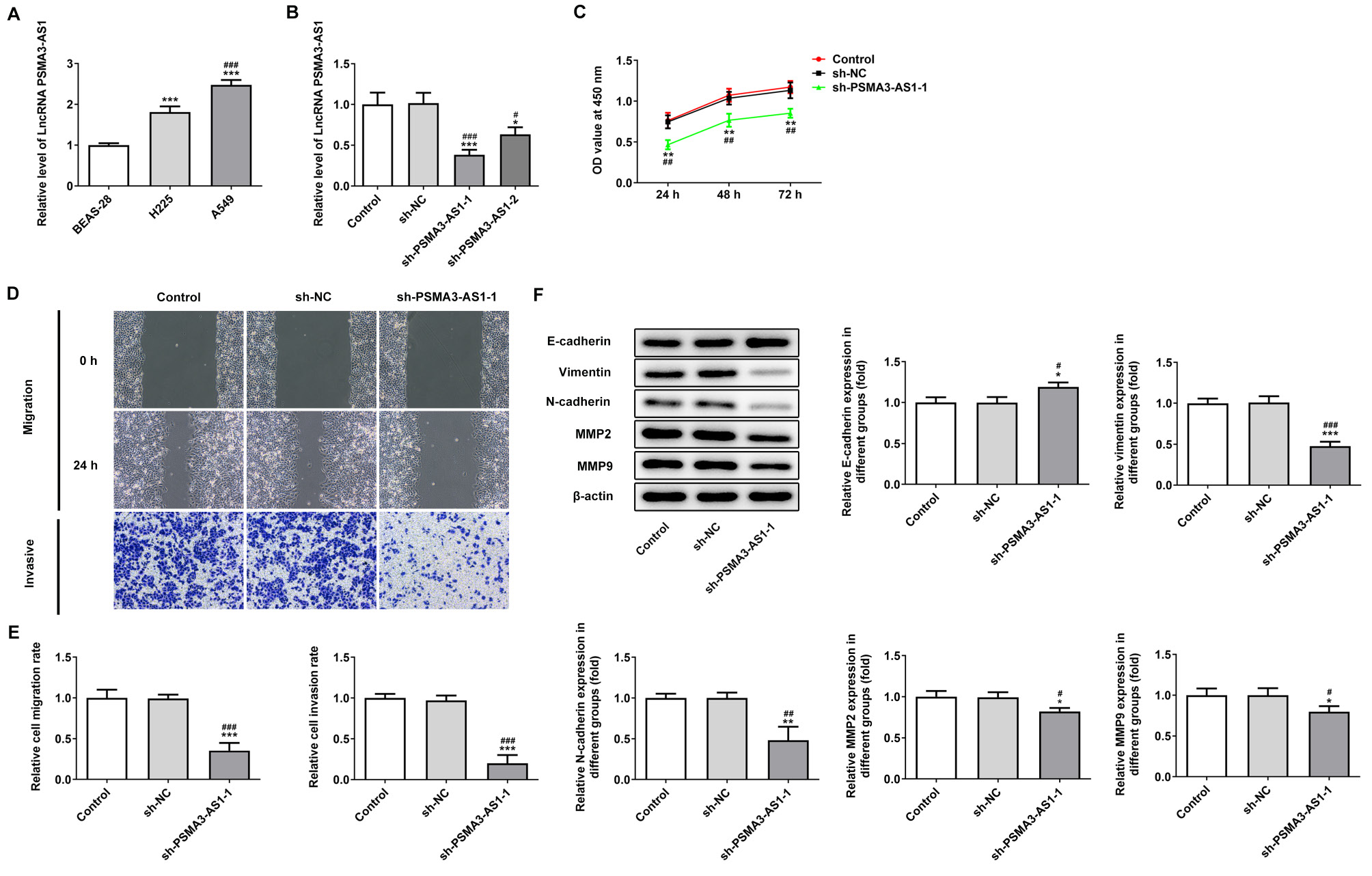
Compared with normal cells, PSMA3-AS1 was upregulated in NSCLC cells, with its highest level of expression in A549 cells (Figure 1A). Therefore, A549 cells were used for the follow-up study. Subsequently, PSMA3-AS1 was found to be decreased in sh-PSMA3-AS1-1 and -2 groups and the expression of PSMA3-AS1 in the sh-PSMA3-AS1-1 group was the lowest (Figure 1B). Knockdown of PSMA3-AS1 significantly inhibited the activity of A549 cells (Figure 1C). As shown in Figure 1D and E, knockdown of PSMA3-AS1 significantly suppressed migration and invasion of A549 cells. In addition, knockdown of PSMA3-AS1 significantly decreased the expression of MMP-2, MMP-9, vimentin, and N-cadherin, whereas it increased the expression of E-cadherin (Figure 1F).
PSMA3-AS1 negatively regulated
miR-17-5p expression
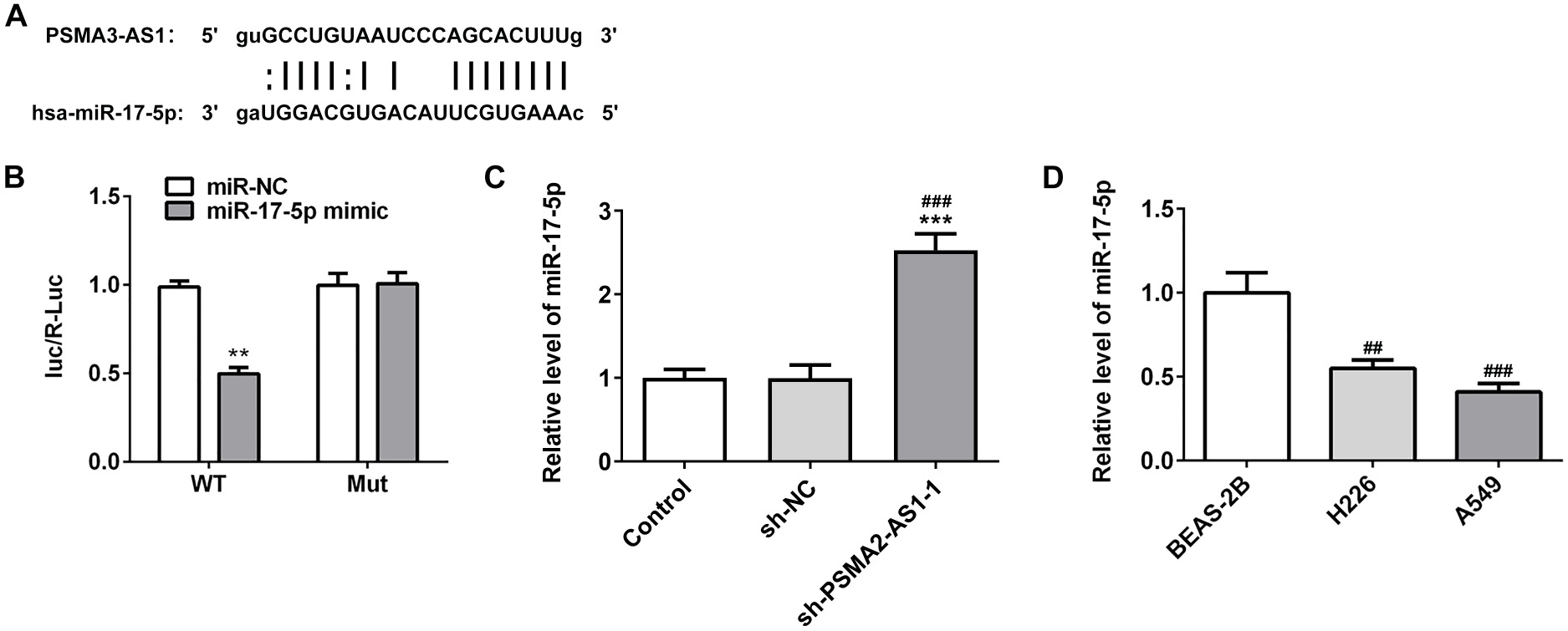
The miR-17-5p and PSMA3-AS1 binding sites were determined (Figure 2A). Relative luciferase activity in the PSMA3-AS1WT+miR-17-5p mimic group was significantly decreased, but there was no change in the Mut group (Figure 2B). miR-17-5p was downregulated in NSCLC cells and knockdown of PSMA3-AS1 significantly enhanced the expression level of miR-17-5p (Figure 2C,D).
miR-17-5p negatively regulated the expression of PD-L1
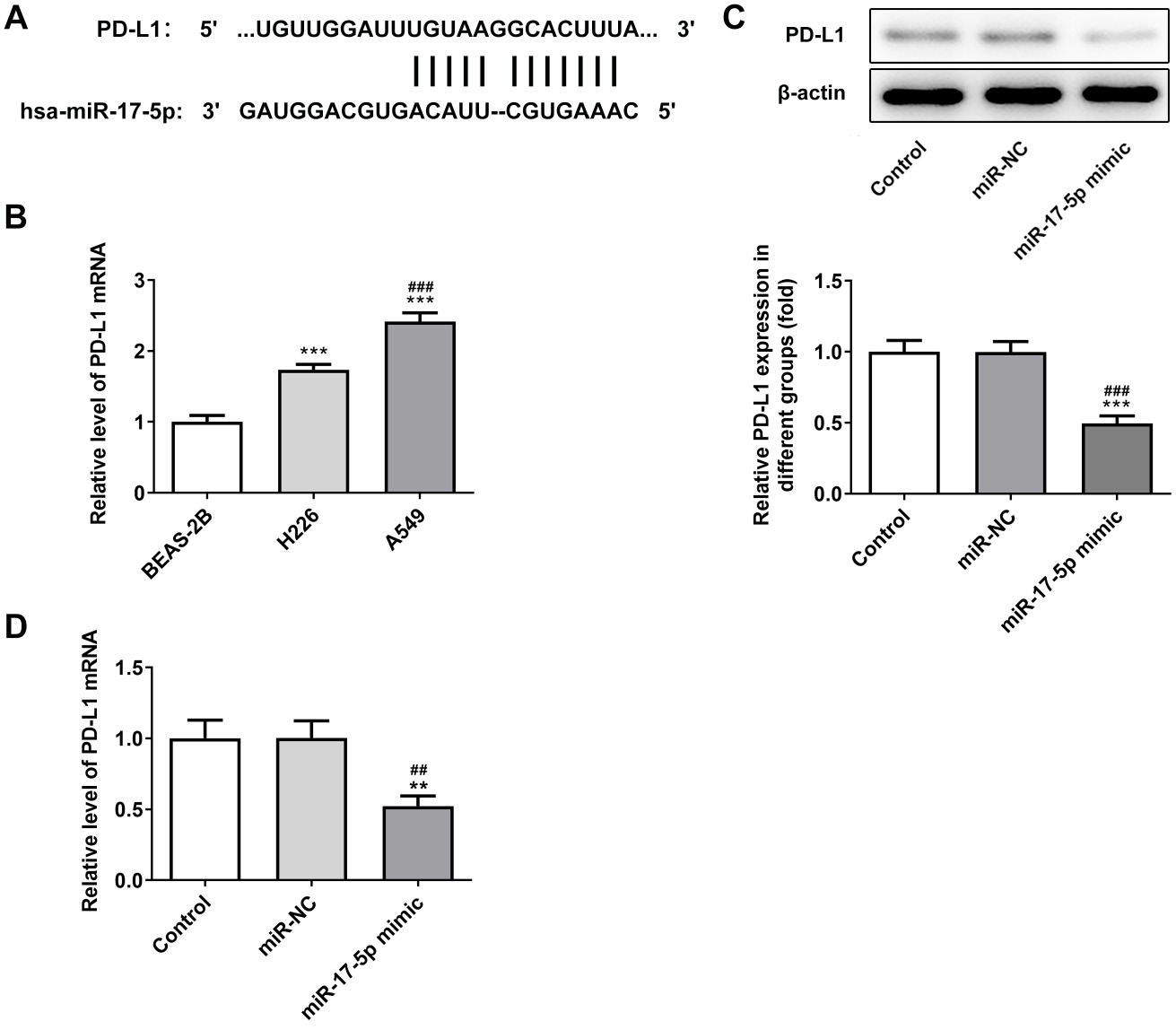
The sites of miR-17-5p and PD-L1 binding are indicated in Figure 3A. PD-L1 was highly expressed in NSCLC cells (Figure 3B). Upregulation of miR-17-5p significantly reduced the expression of PD-L1 in A549 cells (Figure 3C,D).
PSMA3-AS1 regulated the expression of PD-L1 competitively with miR-17-5p
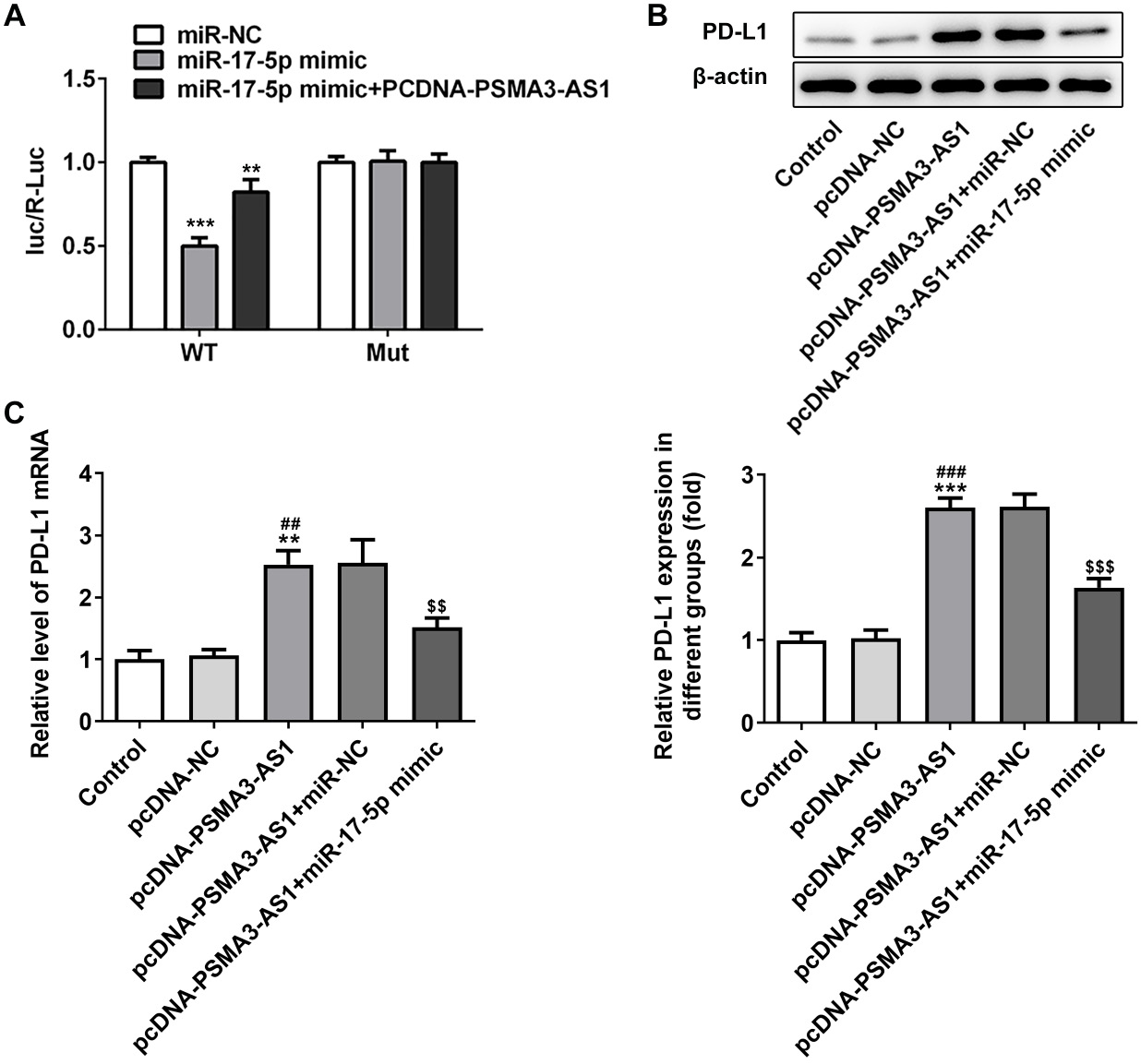
Relative luciferase activity in the PD-L1 WT+ miR-17-5p mimic group was decreased, while it was increased in the PD-L1 WT+ miR-17-5p mimic+PCDNA-PSMA3-AS1 group; this indicated that PSMA3-AS1 and PD-L1 could bind to miR-17-5p competitively (Figure 4A). Upregulation of PSMA3-AS1 significantly promoted the expression of PD-L1, while miR-17-5p overexpression alleviated the enhanced PD-L1 expression caused by overexpression of PSMA3-AS1 (Figure 4B,C).
PD-L1 reversed the inhibitory effect of PSMA3-AS1 on proliferation, invasion, migration, and the EMT process
in A549 cells
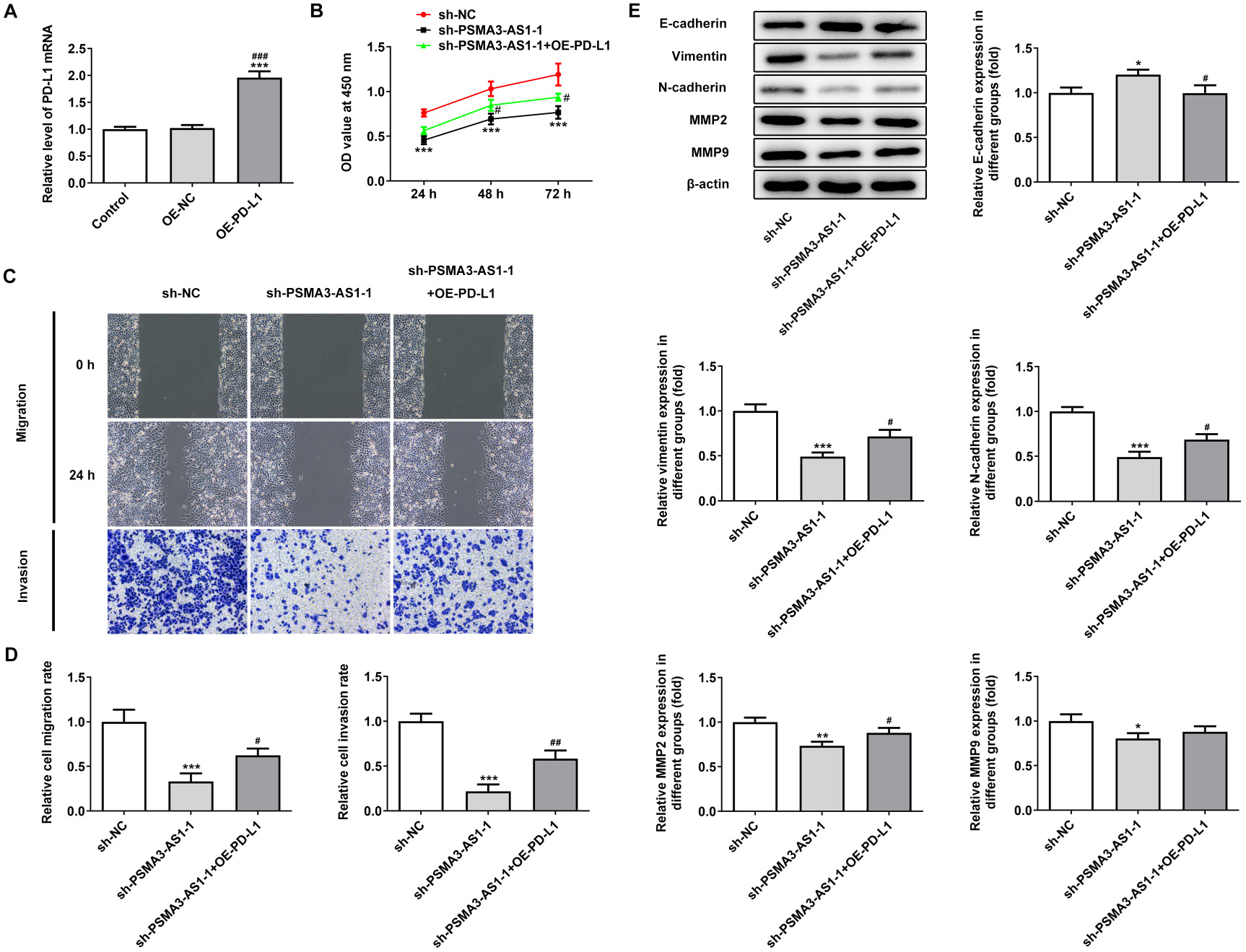
The expression of PD-L1 was increased by transfection with pcDNA-PD-L1 into A549 cells (Figure 5A). Upregulation of PD-L1 significantly reversed the inhibitory effect of PSMA3-AS1 knockdown on cell activity, migration and invasion (Figure 5B–D). In addition, compared with the sh-PSMA3-AS1-1 group, upregulation of PD-L1 significantly promoted the expression of vimentin, N-cadherin, MMP-2, and MMP-9, whereas it decreased the expression of E-cadherin (Figure 5E).
Discussion
The NSCLC is a common respiratory malignancy in adults and is highly susceptible to metastasis.13 Therefore, investigating the potential causes and pathogenesis of NSCLC is critical for improving the clinical curative effects of treatments in NSCLC patients.
Many studies have indicated that lncRNAs are involved in the development of NSCLC.14, 15 Previous studies have shown that PSMA3-AS1 is highly expressed in the majority of tumors.8, 9 Our results confirmed the high expression of PSMA3-AS1 in NSCLC cells. Knockdown of PSMA3-AS1 significantly inhibited the proliferation, migration and invasion of A549 cells. Therefore, upregulation of PSMA3-AS1 in NSCLC may play a role in promoting cancer, but its specific regulatory mechanism remains to be studied.
The lncRNAs not only participate in the activation of transcription and the regulation of the expression of surrounding protein-coding genes but are also relevant to many diseases.16, 17 In addition, lncRNA could affect the role of relevant functional proteins through the adsorption of miRNA.18, 19 For example, the lncRNA SNHG15 accelerates the progression of lung cancer cells by targeting miR-211-3p.20 The lncRNA PVT1-5 facilitates metastasis of lung cancer cells by suppressing the expression of miR-126.21 MiR-17-5p was selected as a target gene of PSMA3-AS1 by screening the starBase (http://starbase.sysu.edu.cn/) database. In thyroid cancer, miR-17-5p aggravates proliferation and invasiveness of thyroid cancer cells.22 In triple-negative breast cancer (TNBC), miR-17-5p can suppress the occurrence of TNBC.23 In this study, miR-17-5p was downregulated in NSCLC and miR-17-5p overexpression significantly reduced the facilitation effect of PSMA3-AS1 on the proliferation, migration and invasion of A549 cells.
For NSCLC treatment, PD-L1 inhibitors could effectively block T-cell proliferation and improve survival of NSCLC patients.24, 25 Meanwhile, PD-L1 was highly expressed in the serum of patients with advanced lung cancer, suggesting that a high level of PD-L1 was correlated with a worse outcome.26 In the present study, we found that miR-17-5p could bind to PD-L1. PD-L1 was upregulated in NSCLC, and the effect of PSM3-AS1 downregulation on cancer progression could be reversed by the upregulation of PD-L1.
Limitations
In our study, we only carried out cellular experiments to investigate the role of PSMA3-AS1 in NSCLC progression by targeting miR-17-5p/PD-L1. However, determining the effect of PSMA3-AS1 inhibition on NSCLC progression in animal experiments and clinical scenarios, as well as the exact mechanism underlying the effect, requires further research.
Conclusions
PSMA3-AS1 was highly expressed in NSCLC. PSMA3-AS1 promoted the proliferation, migration and invasion of NSCLC through upregulation of PD-L1 by targeting miR-17-5p. However, the present study is based on cellular experiments; thus, the detection of PSMA3-AS1 and miR-17-5p in NSCLC tissues and verification with animal experiments will be important to accomplish in future studies.