Abstract
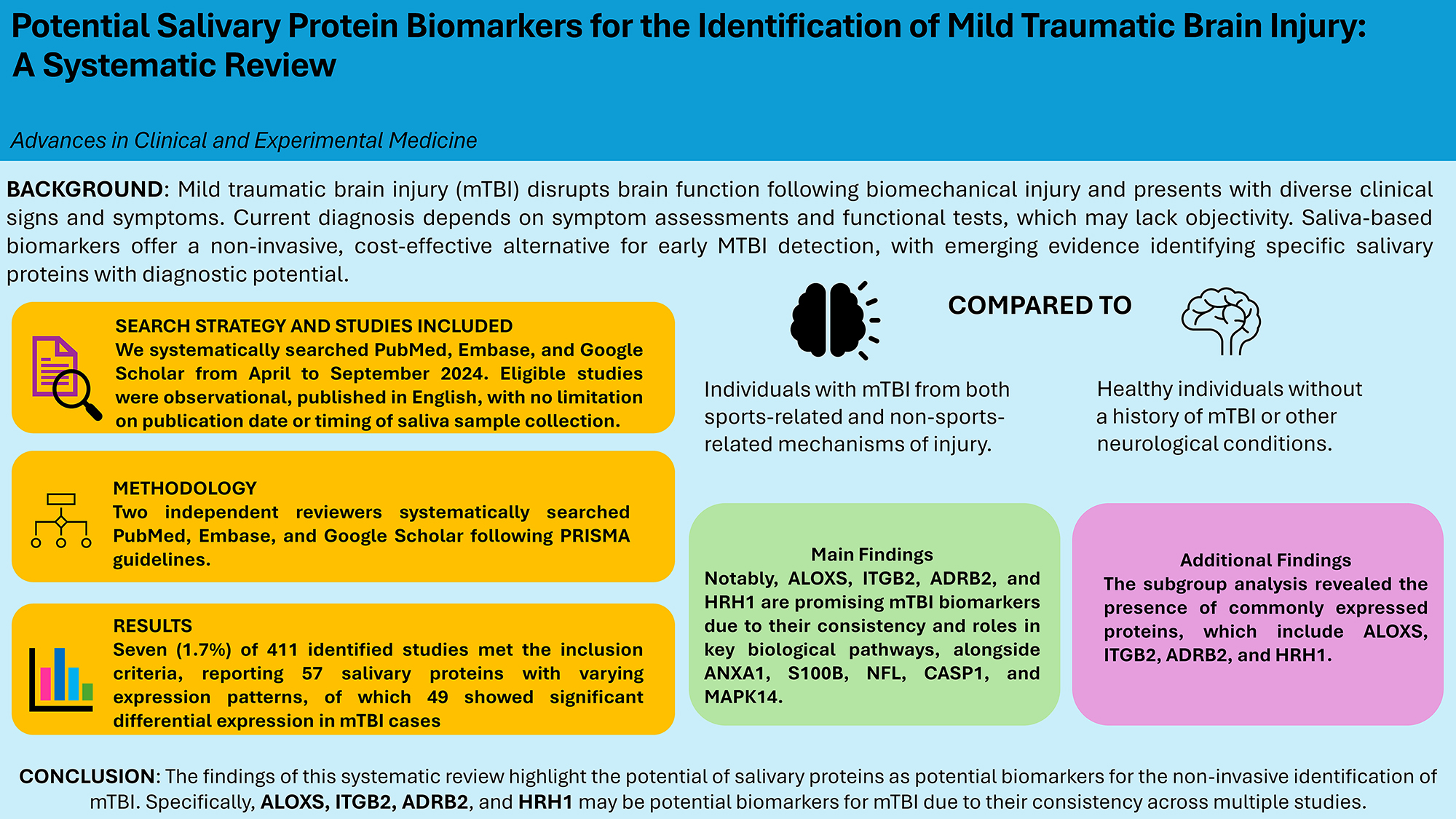
Uncovering objective biomarkers of mild traumatic brain injury (mTBI) may lead to earlier and more accurate identification of this injury, improved prognostic assessment and earlier targeted treatments. Emerging evidence highlights the potential of salivary biomarkers, particularly proteins, as a noninvasive, inexpensive and objective method for diagnosing mTBI, due to their presence in saliva following blood–brain barrier disruption; however, consensus on their utility remains limited. Our aim was to explore the diagnostic potential of salivary protein biomarkers in the identification of mTBI. A comprehensive search was conducted from April to September 2024 across the PubMed, Embase and Google Scholar databases. Original observational studies examining the diagnostic utility of salivary protein biomarkers for mTBI in human populations were included. Seven (1.7%) studies out of 411 identified reports met the inclusion criteria. A total of 49 salivary proteins demonstrated significant differential expression. Subgroup analysis based on the mechanism of injury (sports-related and non-sports-related) identified 4 overlapping proteins: ALOX5, ITGB2, ADRB2, and HRH1. ALOX5 and ITGB2 were significantly upregulated in both subgroups, while ADRB2 and HRH1 were downregulated in sports-related cases and upregulated in non-sports-related cases, which may reflect differential expression related to the region of impact and progression of injury pathophysiology. These findings suggest that salivary proteins such as ALOX5, ITGB2, ADRB2, and HRH1 may serve as promising noninvasive biomarkers for the diagnosis of mTBI. Their involvement in key processes in mTBI pathology supports further investigation in larger, rigorously designed clinical studies to validate their diagnostic utility.
Key words: biomarkers, brain concussion, mild traumatic brain injury
Introduction
Mild traumatic brain injury (mTBI) is defined as a physiological disruption of brain function resulting from a biomechanically plausible mechanism of injury and accompanied by 1 or more clinical signs (e.g., loss of consciousness (LOC), acute symptoms (e.g., physical, cognitive or emotional), and clinical or laboratory findings (e.g., cognitive or vestibular/ocular motor impairment).1 If the injury involves LOC lasting longer than 30 min, a Glasgow Coma Scale (GCS) score of less than 13 at 30 min after injury, or post-traumatic amnesia lasting longer than 24 h, the diagnosis is traumatic brain injury (TBI) rather than mTBI.1 The annual global incidence of mTBI is around 56 million,2 with an estimate of 200–300 per 100,000 individuals for hospitalized patients3 and 1.6–3.8 million for sports-related mTBI.4 However, current research suggests that up to 50% of mTBIs are unreported or undiagnosed,5, 6 highlighting the widespread burden of the injury and its significance as a major public health concern.
The current diagnosis of mTBI relies on a multidomain assessment comprising a clinical exam, symptom evaluation, and balance, neurocognitive, and vestibular/ocular motor testing.7, 8 Standard neuroimaging techniques such as computed tomography (CT) and magnetic resonance imaging (MRI) are costly, not ubiquitously available, and come with concerns about the safety of repeated radiation exposure in monitoring recovery.9 Similarly, blood biomarkers such as glial fibrillar acidic protein (GFAP), ubiquitin carboxy-terminal hydrolase L1 (UCH-L1), tau, and light neurofilament protein (NFL) have shown potential in identifying mTBI, yet the invasive nature of sampling (blood collection) reduces their practicality for widespread use, especially at sites of mTBI occurrence (e.g., sideline).10, 11 Thus, there is a need for novel and/or more discerning approaches to detecting mTBI, such as assessing changes in the expression of salivary biomarkers.12
Saliva-based biomarkers offer a noninvasive means of diagnosing mTBI that is economically efficient, patient-friendly and convenient for sample collection. Saliva contains diverse biomolecules, including proteins, microRNAs and extracellular vesicles, that can mirror pathological processes occurring throughout the body, including within the central nervous system (CNS). Several pathways underpin the connection between CNS pathology and the expression of biomarkers in saliva, the most established being blood-to-saliva transfer. In mTBI, disruption of the blood–brain barrier permits CNS-derived proteins, many of which are low molecular weight (<20 kDa), to enter the bloodstream13; these proteins subsequently pass into saliva through transcellular or paracellular diffusion and ultrafiltration across tight junctions.14 Additionally, the salivary glands are densely innervated by parasympathetic and sympathetic autonomic fibers that lie in close proximity to secretory epithelial cells. This intimate neuro-exocrine arrangement may provide a direct neural route through which neuronal-derived vesicles are released locally and subsequently into saliva.15
Salivary biomarkers have gained traction in TBI research in recent years, with evidence suggesting a link between salivary biomarker expression and TBI severity.16 Recent studies have reported changes in salivary microRNAs (miRNAs) such as let7a-5p, miR-143-3p, miR-103a-3p, miR-34b-3p, RNU6-7, RNU6-45, Snora57, snoU13.120, tRNA18Arg-CCT, U6-168, U6-428, U6-1249, Uco22cjg1, and YRNA_255, which showed accurate association with mTBI.17 Extant evidence has also demonstrated a relationship between miRNAs and mTBI assessment tools, such as the Immediate Post-Concussion Assessment and Cognitive Testing (ImPACT).
Saliva also presents several unique advantages over blood for the diagnosis of mTBI. Its noninvasive collection method enables frequent, real-time and point-of-care testing, which is critical in high-risk environments such as sports, military and emergency settings. Unlike blood-based diagnostic methods requiring sterile conditions or trained personnel, saliva sampling can be performed with minimal resources, thereby reducing healthcare costs and biosafety concerns. These practical benefits, coupled with improved patient compliance and the feasibility of longitudinal monitoring, position salivary biomarkers as a promising alternative for mTBI diagnosis and management.
While existing systematic reviews and meta-analyses of potential salivary biomarkers have primarily focused on other diagnostic parameters, such as miRNAs,18, 19 highlighting their potential relevance, the diagnostic capacity of salivary proteins remains underinvestigated. Moreover, researchers have identified differentially expressed salivary proteins with diagnostic potential for mTBI, but currently, the need for a comprehensive biomarker panel limits their practical application in mTBI diagnosis.
Objectives
This systematic review aimed to comprehensively evaluate the clinical utility of salivary protein biomarkers for identifying mTBI, without limiting the studies based on the time of sample collection.
Materials and methods
Protocol registration
To enhance the transparency of the systematic review process, the protocol was registered with the International Prospective Register of Systematic Reviews (PROSPERO) on October 3, 2024, under registration No. CRD42024592200.
Search strategy
This systematic review was conducted between April and September 2024 following the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines.20 The study was based on the Population, Exposure, Comparisons, Outcome, and Study Design (PECOS) framework,21 where the population included individuals with mTBI; the exposure was mTBI; healthy individuals without mTBI served as the comparison group; and the outcome focused on diagnostic identification.
We systematically searched 3 databases – PubMed, Embase and Google Scholar (for grey literature) – using different combinations of key terms and Medical Subject Headings (MeSH terms) from April to September 2024. We included only observational studies, such as case-control, cross-sectional and cohort studies, published in English, with no restriction on the year of publication or the time of sample collection post-injury. For Google Scholar, the search was limited to the first 200 results (approx. 20 pages). Boolean operators (“AND” and “OR”) were used to combine keywords or independent searches, and the final search string used for the systematic search was (“traumatic brain injury” OR “mild TBI” OR “mTBI” OR “concussion” (MeSH) OR “head trauma”) AND (“salivary biomarkers” OR “salivary extracellular vesicles” OR “salivary cytokines” OR “saliva samples”). The bibliographies of the shortlisted articles were also searched to supplement the database findings.
Eligibility criteria
Only studies conducted in human populations meeting the following criteria were included: 1) original observational studies, 2) reported salivary protein biomarkers, 3) examined the diagnostic relevance of salivary protein biomarkers in mTBI, and 4) published in English only. Animal studies, case reports, randomized controlled trials (RCTs), systematic reviews and meta-analyses, abstracts, editorial reviews, and qualitative studies were excluded. Consistent with our predefined exclusion criteria, our search retrieved no RCTs; RCT intervention frameworks can introduce treatment-related alterations in protein expression, thereby confounding the intrinsic differential expression profiles associated with TBI. These changes could obscure the natural differential expression profiles related to TBI that this study seeks to identify.22
Study selection
The articles from the 3 databases meeting the eligibility criteria were saved as CSV files and imported into the Rayyan software,23 where Rayyan’s default algorithm was initially used to identify duplicate studies, which were subsequently manually reviewed for accuracy. Two reviewers (O.T. and D.K.) independently screened the titles and abstracts of all identified studies, as well as the full texts of potentially eligible studies. Any initial differences in judgment were discussed and resolved between the 2 reviewers. In cases where clarification was needed, the 3rd reviewer (S.H.A.) was consulted. However, no major discrepancies were encountered during the screening or selection process. The 3rd reviewer also supervised the search strategy, screening procedures and final study selection to ensure methodological accuracy and consistency.
Critical appraisal
Critical appraisal of the selected studies was conducted using the Joanna Briggs Institute (JBI) tool for cross-sectional studies implemented in the JBI SUMARI tool.24 For this review, the JBI checklist for case-control cross-sectional studies was used, which captured the methodologies represented in the shortlisted studies. Each question was systematically addressed, and the tool automatically calculated the quality score for each study, reflecting its methodological scrutiny.
Data extraction and evidence synthesis
The 1st author extracted essential details, including the authors’ names, publication year, study design, study participants, study sample size, age range, mean age, mechanism of trauma, saliva collection method, detection techniques, the total number of biomarkers assessed, and their pattern of dysregulation, while the 2nd author then verified the extracted data for completeness and accuracy.
We reviewed the methodology of each study to identify the primary mechanism of injury and classified all shortlisted studies into 2 groups: sports-related mTBI and non-sports-related mTBI for subgroup analysis. The sports-related subgroup comprised 3 studies with only 2 studies reporting differential protein expression (n = 80 adults, both sexes; saliva collected ≤1 h post-injury), while the non-sports-related subgroup included 4 studies (n = 253 participants – encompassing both children and adult age groups, both sexes; sampling at <6 h to several days post-injury). From each study, we extracted data on the total number of significantly differentially expressed proteins, the pattern of expression (up- or downregulated), and mean expression values. Both subgroups exhibited balanced sex distributions; however, the sports-related cohort was more homogeneous in age distribution with uniformly early sampling, while the non-sports cohort demonstrated greater age diversity and temporal heterogeneity in sampling. A sample size-weighted mean was calculated for overlapping proteins to provide a descriptive summary of expression changes between subgroups.
Mean fold-change values for proteins reported in multiple studies were combined using a weighted mean approach to facilitate quantitative comparison in the absence of comprehensive statistical metrics.25 The approach was implemented in R v. 4.4.2 (R Foundation for Statistical Computing, Vienna, Austria) using the weighted.mean (mean, n) function (Supplementary Table 1).
In order to understand the neurobiological underpinnings of mTBI and how salivary protein biomarkers may reveal this process, pathway enrichment analysis was carried out using the web-based Enrichr tool,26 with the Kyoto Encyclopedia of Genes and Genomes (KEGG)27 and Gene Ontology (GO)28 databases to identify biological pathways linked to the proteins identified in this review. Only 7 studies met the inclusion criteria, limiting formal assessment of publication bias.29 All authors confirmed the list of selected articles for accuracy.
Results
Study selection and characteristics
The process of screening and selection, as well as the inclusion and exclusion criteria of the studies, is detailed in the PRISMA flowchart (Figure 1). The initial search in PubMed and Embase resulted in 262 studies; title and abstract screening was conducted to exclude 58 duplicate studies. Studies that did not meet the inclusion criteria for the systematic review were also excluded, and 11 studies were retrieved for full-text screening. However, due to the unavailability of full texts, we further excluded 2 studies, and 2 additional studies were excluded during full-text screening as they investigated salivary proteins without evaluating their diagnostic utility. We also searched Google Scholar for grey literature, with the initial search yielding 3,530 studies. Of these, only 149 studies were retrieved, while the remaining studies were excluded because they were previous systematic reviews, meta-analyses, animal studies, or literature reviews. However, none of the 149 studies retrieved were eligible for inclusion, as they included already shortlisted studies or did not examine salivary protein biomarkers for mTBI identification. Finally, 7 (1.7%) of the 411 identified studies were selected for inclusion in our systematic review.10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36. These studies were assessed for risk of bias using the JBI checklist and were found to have a low risk of bias (Supplementary Table 2).
The characteristics and major findings of the 7 studies are summarized in Table 1 and Supplementary Table 3. Out of 7 studies, 4 were case-control31, 32, 33, 36 and 3 were cross-sectional.30, 33, 34 Three studies reported the mechanism of injury to be sports-related,30, 33, 34 while the remaining studies included non-sports-related mechanisms, such as external impacts, accidents, falls, and other head trauma.31, 32, 35, 36 The salivary collection method was stimulated36 and non-stimulated,30, 31, 32, 33, 34, 35 and the frequency of saliva collection was twice daily (pre- and post-events) in 2 studies,33, 34 while other studies only collected saliva samples once.30, 31, 32, 35, 36 The studies predominantly recruited adult participants; only 2 studies included individuals younger than 18 years, raising the possibility that age may confound the biomarker results. In addition, both male and female participants were enrolled, and the interval between injury and saliva collection varied widely – from within 1 h to less than 6 h, up to 24 h in emergency department settings, and even longer in concussion clinics – indicating that sex and sample collection timing constitute further potential confounders that must be considered when interpreting the findings. Collectively, the 7 studies examined differential salivary protein expression for potential diagnostic use in 361 participants, including 299 mTBI cases and 62 controls, providing a sufficient sample size for preliminary findings.37
Pattern of salivary biomarker expression and subgroup analysis
Salivary protein biomarkers
A total of 57 proteins were reported in 7 studies, with individual studies reporting 4,30 27,31 22,32 7,33 2,34 1,35 and 136 protein, respectively. These 57 proteins showed cross-expression and varying expression patterns across the studies, with different biomarkers identified in each study. Only 49 proteins demonstrated significant differential expression with a pattern of dysregulation (Table 2). Notably, only 4 proteins, namely arachidonate 5-lipoxygenase (ALOX5), integrin subunit beta 2 (ITGB2), adrenoceptor beta 2 (ADRB2), and histamine receptor H1 (HRH1), overlapped across 2 studies.32, 33
Differential protein expression based on injury mechanism
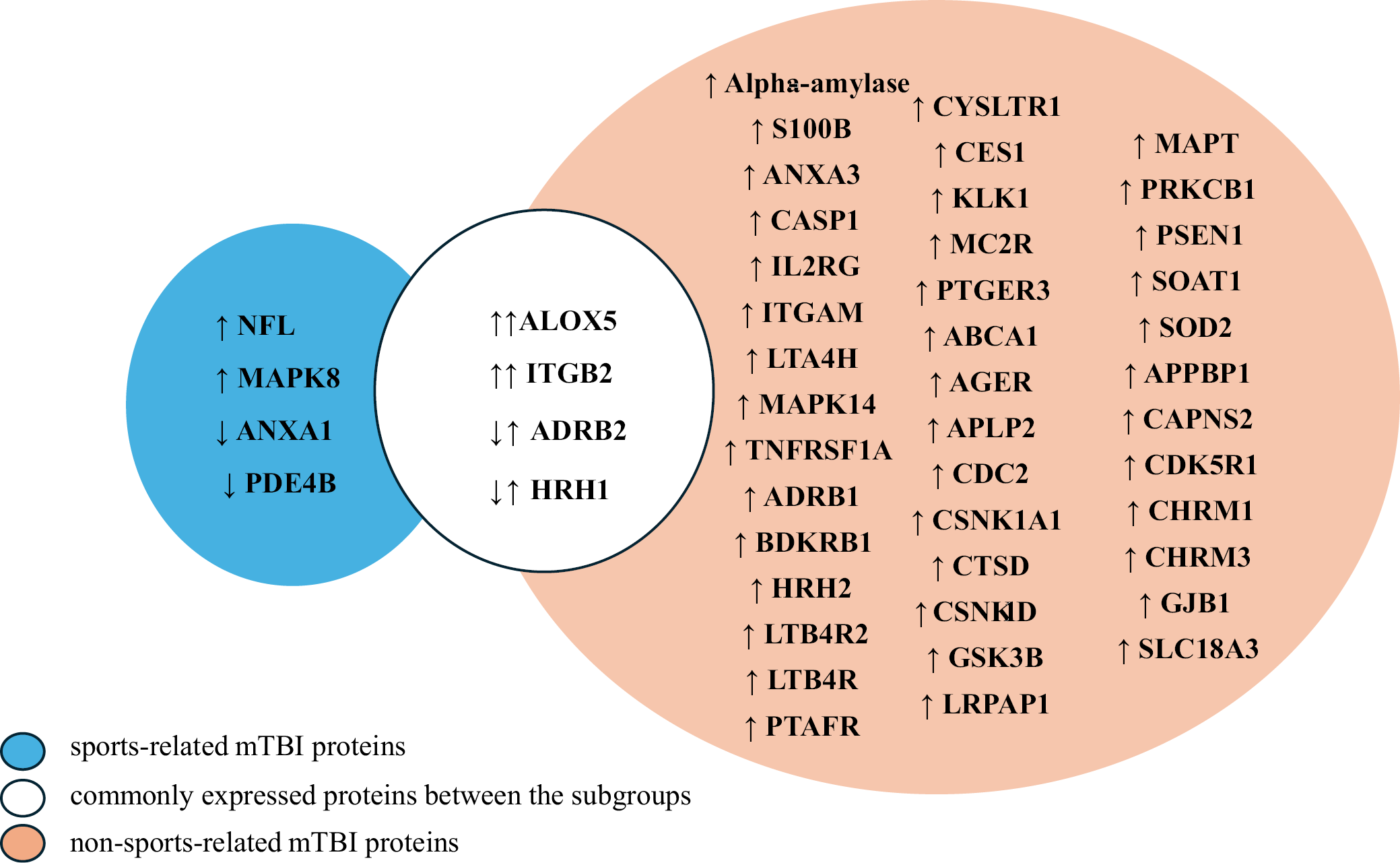
We conducted a subgroup analysis (Figure 2) to explore differences in salivary biomarker expression based on the mechanism of injury (sports-related and non-sports-related cases). Three studies examined sports-related mTBI (collegiate water polo,34 mixed martial arts33 and collision sports30). Out of these 3 studies, 1 study reported no significant differences in salivary protein expression profiles,30 while the other 2 identified 8 differentially expressed proteins: mitoprotein-activated protein kinase 8 (MAPK8), annexin A1 (ANXA1), phosphodiesterase 4B (PDE4B), NFL, ADRB2, ALOX5, ITGB2, and HRH1. The fold changes (FCs) for these proteins ranged from 0.03 to 111, except for NFL, which was not reported. Among these, the highest expression was noted for ALOX5 (FC: 111), followed by ITGB2 (FC: 70) and MAPK8 (FC: 50). Conversely, with the baseline FC set at 1, downregulated proteins included ADRB2 (FC: 0.03), HRH1 (FC: 0.03), ANXA1 (FC: 0.6), and PDE4B (FC: 0.3).
In non-sports-related mTBI, a broader range of biomarkers was reported, with 45 proteins showing differential expression. All these proteins were upregulated in the identified studies, with the most significantly upregulated protein being HRH1 (FC: 253.9 ±140.72), and the least upregulated being amyloid beta precursor protein binding protein 1 (APPBP1) (FC: 0.42 ±0.65).
Comparative analysis of protein expression in 2 groups identified 4 proteins (ALOX5, ITGB2, ADRB2, and HRH1) commonly expressed in both groups. All 4 proteins exhibited differential expression in mTBI patients, with ALOX5 (FC: 253.9 ±140.72), ITGB2 (FC: 14.53 ±6.01), ADRB2 (FC: 25.14 ±10.25), and HRH1 (FC: 253.9 ±140.72) showing significant fold changes in non-sports-related cases. While ALOX5 and ITGB2 maintained upregulation across both groups, ADRB2 and HRH1 showed distinct trends, with significant downregulation observed in sports-related mTBI and upregulation in non-sports-related mTBI, which may suggest differences in protein expression based on the mechanism of injury.
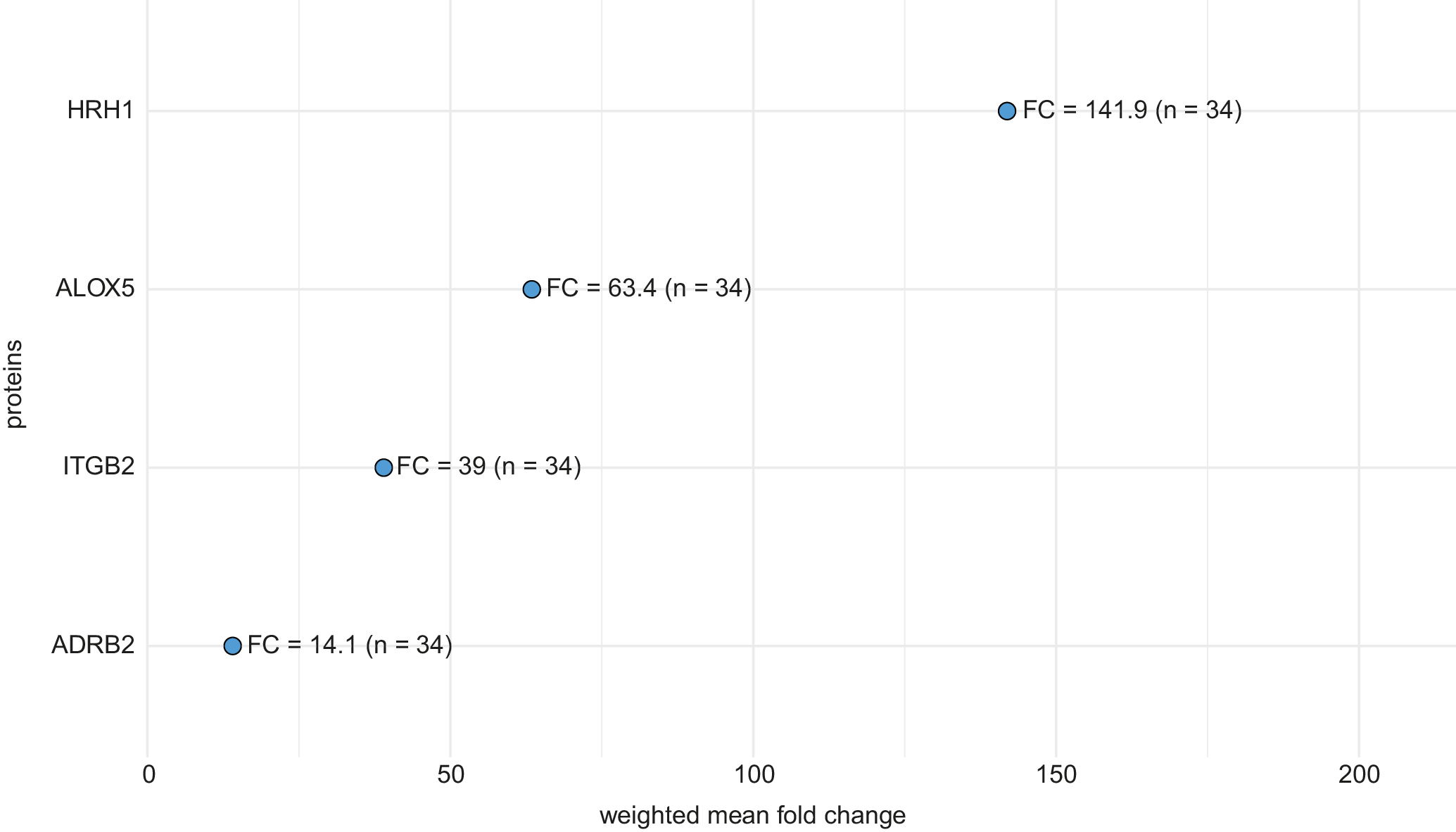
Pooled weighted fold-change analysis of overlapping proteins
Weighted mean analysis of proteins reported in 2 studies32, 33 revealed HRH1 to exhibit the highest pooled upregulation (FC: 141.9), twice the expression level of the next highest protein, ALOX5 (FC: 63.4). ITGB2 also showed substantial upregulation (FC: 39.0), while the least upregulation was observed for ADRB2 (FC: 14.1). These values reflect pooled fold changes based on a total sample size of 34 participants (Figure 3).
However, it is important to note that HRH1 and ADRB2 were downregulated in 1 of the contributing studies, but the significantly higher upregulation observed in the other study shifted the overall pooled direction. Additionally, due to incomplete statistical reporting in the included studies, such as the absence of standard deviations (SDs) or variance estimates, the precision of these pooled fold changes could not be assessed, and confidence intervals (CIs) could not be calculated.
Discussion
This systematic review evaluated the potential of salivary proteins as noninvasive biomarkers for diagnosing mTBI across 7 studies that met the inclusion criteria. Notably, ALOX5, ITGB2, ADRB2, and HRH1 represent potential biomarkers for mTBI due to their consistency across multiple studies with different mechanisms of injury and their roles in inflammation, immune response, and nervous system coordination. Other potential candidate biomarkers include S100 calcium-binding protein B (S100B), caspase 1 (CASP1), mitogen-activated protein kinase 14 (MAPK14), ANXA1, and NFL, which have biological functions similarly implicated in these pathways38 (Supplementary Table 4). Our findings contribute to the growing evidence supporting the utility of salivary biomarkers for mTBI identification and provide a foundation for developing a targeted diagnostic panel based on proteins significantly associated with TBI (differentially expressed).
Due to the inherent heterogeneity of mTBI pathogenesis, employing a panel of biomarkers may provide the most promising strategy to improve diagnostic efficacy.39 This approach has the potential to detect a broader range of pathophysiological changes, including neuronal damage and inflammation.40 Salivary proteins also offer distinct advantages that may enhance diagnostic accuracy, as their levels can reflect the extent of brain injury due to their direct association with neuronal tissue damage and may directly reflect the severity of neural injury.40 Furthermore, proteins are more stable and may provide greater specificity to the injury context, whereas other biomarkers, such as hormones and miRNAs, are easily degraded and affected by physiological factors unrelated to mTBI.41
Our analysis identified 49 salivary proteins that showed significant differential expression (Table 2) across the shortlisted studies. The predominant pattern of expression among these proteins was upregulation, with only a small number of proteins (n = 4) demonstrating downregulation in cases of mTBI. Weighted analysis of the overlapping proteins (Figure 3) yielded pooled estimates of expression levels across the 2 included studies.32, 33 HRH1 exhibited the highest fold change, indicating strong overall upregulation, while ADRB2 showed the lowest, reflecting the aggregate magnitude of change across both studies. Despite downregulation in 1 study,33 the pooled fold changes for HRH1 and ADRB2 remained positive, driven by the stronger upregulation observed in the other study.32
Matuk et al.33 identified 7 differentially expressed proteins in mixed martial artists exposed to head trauma. Three salivary proteins (ALOX5, ITGB2 and MAPK8) were significantly upregulated in post-fight samples. In contrast, ADRB2 and HRH1 exhibited baseline upregulation relative to controls but were significantly downregulated following the fight, while ANXA1 and PDE4B, which did not differ from controls at baseline, also decreased post-fight. These nuanced changes highlight the dynamic nature of protein expression in response to acute head trauma.
In a related study, Monroe et al.34 identified significant upregulation of NFL in a cohort of water polo players, which positively correlated with the number of repeated head impacts, whereas S100B showed no significant changes and was not significantly correlated. This contrasts with the study by Janigro et al.,36 in which mTBI patients exhibited significant upregulation of salivary S100B. This discrepancy may suggest that the mechanism of trauma acts as a confounding factor in biomarker expression. Salivary alpha-amylase was reportedly upregulated in patients with isolated mild head trauma.35 Cheng et al.31 compared differential expression of salivary proteins between controls, concussion clinic patients, and emergency department patients with head trauma and identified 23 upregulated proteins across both comparisons with controls. Of these, 3 proteins – casein kinase 1 alpha 1 (CSNK1A1), cathepsin D (CTSD) and cell division cycle 2 (CDC2) – showed consistent upregulation across both comparisons, but only 2 (CSNK1A1 and CTSD) were statistically significant in both comparisons.31 Furthermore, Cheng et al.,32 in a continuation study with the same participant demographics, identified a total of 22 salivary proteins upregulated in concussion clinic patients and emergency department patients compared with healthy controls, which could offer potential diagnostic relevance in mTBI. Inflammation-related proteins, including brain-derived neurotrophic factor (BDNF), C-reactive protein (CRP), interleukin 1β (IL-1β), and tumor necrosis factor alpha (TNF-α), were examined in a group of collision sports athletes, and no significant changes in their expression patterns were found.30 This indicates that certain inflammatory biomarkers may not be significantly altered in salivary samples from mTBI patients.
Across 2 studies in this systematic review, 4 salivary proteins (ALOX5, ITGB2, ADRB2, and HRH1) were found to overlap.32, 33 ALOX5 and ITGB2 demonstrated consistent upregulation in both studies, reinforcing their potential as biomarkers with high diagnostic value given their direct biological roles in inflammation, cell adhesion and immune response.42 In contrast, ADRB2 and HRH1 exhibited opposing trends: while these proteins were significantly downregulated in sports-related mTBI, as observed by Matuk et al.,33 they were upregulated in non-sports-related cases reported by Cheng et al.32 The downregulation of ADRB2 and HRH1 in the sports-related cohort suggests that specific injury dynamics, such as the region and severity of impact, might influence receptor activity and subsequent protein expression.43 While this hypothesis is plausible, further detailed investigation is necessary to understand how these factors affect biomarker expression in mTBI. Furthermore, some proteins, such as S100B, NFL, ALOX5, ITGB2, ANXA1, PDE4B, HRH1, CASP1, and MAPK14, are involved in key biological processes relevant to mTBI, suggesting that they may be promising biomarkers for further investigation. Specifically, S100B and NFL have also been reported in studies examining potential biomarkers in cerebrospinal fluid and blood from mTBI patients.10, 11, 44
Our subgroup analysis revealed differences in salivary protein expression profiles between sports-related and non-sports-related mTBI, which may contribute to understanding the possible influence of the mechanism of injury on the pathophysiological response. Specifically, 8 differentially expressed proteins were identified in sports-related mTBI and 45 in non-sports-related mTBI (Figure 2). Four protein biomarkers, ALOX5, ADRB2, HRH1, and ITGB2, were found to be shared between both groups; however, a contrasting pattern of dysregulation was observed in ADRB2 and HRH1, which were downregulated in sports-related mTBI but upregulated in non-sports-related mTBI. Such trends suggest that injury-specific factors, such as timing, force of impact and the affected brain region, may modulate protein expression patterns.45 In addition, overlapping expression between the 2 groups suggests that the 2 mechanisms of injury share similar pathophysiological processes. However, more evidence is needed to validate these findings and to explore the basis of the observed differences, which will inform the development of a targeted diagnostic panel.
Several salivary proteins identified in our analysis are known to influence key pathophysiological processes spanning neuroinflammation, neuromodulation, cellular stress, and tissue repair. ALOX5 and ANXA1 are important in regulating inflammation,46 while PDE4B amplifies cyclic AMP-mediated inflammatory cascades47 and ITGB2 mediates immune responses and facilitates cell adhesion.48 NFL serves as a direct indicator of axonal injury and cytoskeletal disruption,49 whereas ADRB2 and HRH1 reflect trauma-induced alterations in neuromodulatory tone.50 Downstream stress-response kinases, including MAPK8, signal apoptotic and inflammatory amplification,51 and the lysosomal enzyme CTSD implicates maladaptive proteolysis linked to neurodegenerative sequelae.52, 53 Concurrently, CSNK1A1 helps recruit cytoskeletal and migratory machinery essential to tissue repair.54
The biological plausibility of detecting these neural proteins in saliva is supported by mechanistic evidence describing how CNS-derived substrates are transported into the salivary environment. This neuroanatomical link has been explored for α-synuclein, a neural protein detectable in saliva and considered a noninvasive biomarker for Parkinson’s disease.15 Neuronal proteins are synthesized in central and peripheral neurons, packaged into vesicles, and transported anterogradely along microtubules via kinesin motor proteins. These vesicles reach the axon terminals of autonomic nerve fibers that innervate the salivary glands, where their contents are released through soluble N-ethylmaleimide-sensitive factor attachment protein receptor (SNARE)-mediated exocytosis.15, 55 Given the proximity of these nerve terminals to the glandular epithelium, neural proteins may diffuse into or be secreted through the salivary ductal system.56 This neurosecretory mechanism provides a credible biological route for the appearance of CNS biomarkers in saliva, reinforcing its value as a noninvasive medium for monitoring brain health.
Building on this mechanistic foundation, our neurobiological pathway enrichment analysis contextualizes the identified salivary proteins within 10 interconnected networks implicated in secondary injury responses following mTBI (Figure 4). The neuroactive ligand–receptor interaction pathway emerges as the most significantly enriched, comprising 13 G protein-coupled receptors (GPCRs) – cholinergic receptor muscarinic 1 (CHRM1), cholinergic receptor muscarinic 3 (CHRM3), platelet-activating factor receptor (PTAFR), prostaglandin E receptor 3 (PTGER3), adrenoceptor beta 1 (ADRB1), leukotriene B4 receptor (LTB4R), leukotriene B4 receptor 2 (LTB4R2), melanocortin 2 receptor (MC2R), histamine receptor H2 (HRH2), cysteinyl leukotriene receptor 1 (CYSLTR1), bradykinin receptor B1 (BDKRB1), ADRB2, and HRH1. These proteins exhibit differential expression that mirrors the dysregulation of glutamate-mediated excitotoxicity, GABAergic disinhibition, and subsequent N-methyl-D-aspartate (NMDA) receptor-driven calcium influx, as observed in mTBI pathophysiology.57, 58 The enrichment of both G protein-coupled receptor (GPCR) and calcium signaling pathways further underscores the role of receptor-driven activation of phospholipase C beta (PLCβ)-mediated calcium release and adenylyl cyclase-mediated cyclic AMP (cAMP) production, which are critical in modulating the intracellular response to injury.59 Additionally, the neuroinflammation module, which includes MC2R, CYSLTR1, tumor necrosis factor receptor superfamily member 1A (TNFRSF1A), HRH2, ALOX5, ADRB1, ADRB2, and MAPK14, integrates both pro- and anti-inflammatory lipid mediators and cytokine responses. The cytokine response pathway, enriched with glycogen synthase kinase 3 beta (GSK3B), integrin subunit alpha M (ITGAM), interleukin 2 receptor subunit gamma (IL2RG), MAPK8, ANXA1, ALOX5, ITGB2, PTAFR, CASP1, and TNFRSF1A, highlights inflammasome activation and NF-κB-dependent gene expression, which are central to the inflammatory cascade following injury. Lastly, the microglial activation network, enriched by microtubule-associated protein tau (MAPT), advanced glycosylation end product-specific receptor (AGER), ITGAM, and ITGB2, captures the engagement of innate immune responses, essential for tissue repair and neuroprotection following mTBI.60
These enriched pathways not only reflect the complex secondary injury processes underlying mTBI but also highlight candidate proteins with mechanistic and diagnostic relevance. By mapping salivary proteins to these interconnected networks, this study underscores their potential utility in developing noninvasive diagnostic panels for early detection, monitoring and therapeutic targeting of mild traumatic brain injury.
Limitations
This systematic review is limited by several factors that may affect the generalizability of the findings. The included studies exhibited substantial variability in injury mechanisms, study design, and participant demographics. Subjects ranged from emergency department and concussion clinic patients to collegiate athletes engaged in high-contact sports, such as water polo and mixed martial arts. While this diversity may enhance the applicability of the findings, it may also introduce variability in biomarker expression levels, which could compromise diagnostic consistency. Additionally, we did not examine differential expression based on biological sex, which may confound the interpretation of the results, given the well-established differences in pathophysiological responses between male and female patients.61 Methodological inconsistencies were also evident in the saliva collection protocols. For example, 1 study utilized a stimulated collection method, compared with other studies employing a non-stimulated approach. In addition, the timing of saliva collection post-injury varied from approx. 1 h post-injury to less than 24 h and even extended to several days in concussion clinic patients, which could potentially introduce pre-analytical variability. The relatively small sample sizes in some studies may increase the risk of false-positive findings. A formal quantitative assessment of publication bias was not feasible due to the limited number of studies.29 However, an assessment of the included studies indicates potential publication bias, as most reported positive findings.
Conclusions
This systematic review highlights the potential of salivary proteins as noninvasive indicators for detecting mild traumatic brain injury. Proteins such as ALOX5, ITGB2, ADRB2, and HRH1 showed consistent differential expression across varied injury contexts, reflecting their involvement in key biological processes, including inflammation, immune signaling, and neuromodulation. Additional candidates such as ANXA1, S100B, NFL, CASP1, and MAPK14 further point to the complex pathophysiology of mTBI, encompassing disruptions in neural integrity and barrier function.
However, despite these promising findings, none of the identified biomarkers are currently suitable for clinical application. The existing evidence remains preliminary, limited by small sample sizes, methodological variability, and a lack of robust validation. Advancing these biomarkers toward clinical relevance will require standardized methodologies, expanded and demographically varied cohorts, and longitudinal studies to track temporal shifts in protein expression. The limited number of studies in this field underscores a significant gap in the current body of knowledge that warrants further investigation. Therefore, future research, particularly well-designed case-control studies, is required to validate existing findings and identify additional salivary proteins associated with mTBI.
Supplementary data
The supplementary materials are available at https://doi.org/10.5281/zenodo.15478937. The package contains the following files:
Supplementary Table 1. R script for weighted means analysis.
Supplementary Table 2. Critical appraisal of the shortlisted studies based on the JBI tool.
Supplementary Table 3. Study characteristics.
Supplementary Table 4. Differentially expressed proteins and their biological functions.
Use of AI and AI-assisted technologies
Not applicable.