Abstract
Background. Urinary tract infections (UTIs) are one of the most common infectious diseases. Inappropriate and excessive administration of antibiotics has led to the increased antibiotic resistance in the pathogens that cause UTIs. This work focused on identifying genetic determinants of antibiotic resistance in a clinical isolate of UTI-causing Escherichia coli.
Objectives. A clinical isolate of E. coli resistant to β-lactam, tetracycline and aminoglycoside antibiotics was analyzed using whole-genome sequencing (WGS) to identify genes that contribute to its resistance.
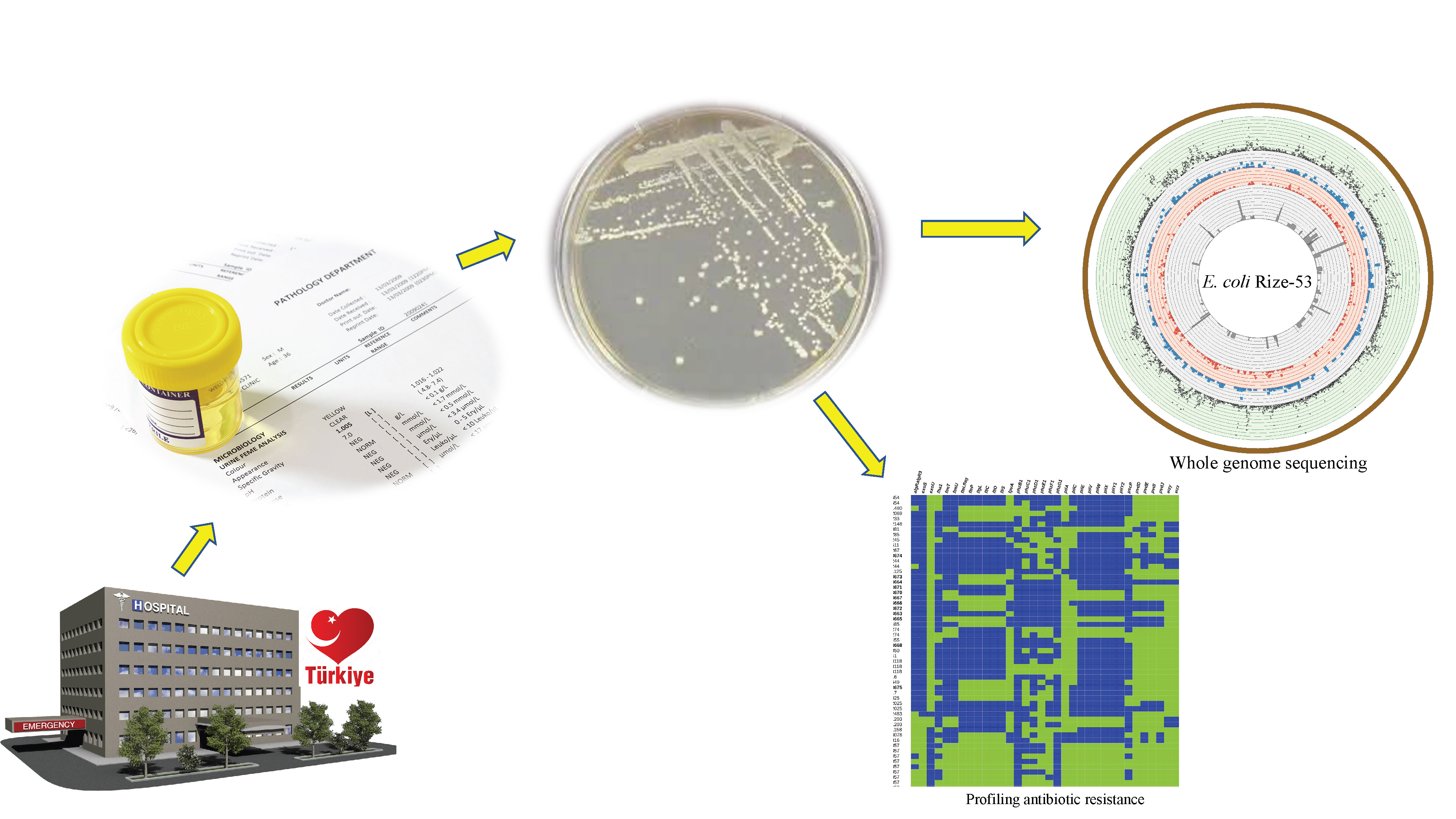
Materials and methods. The clinical isolate was obtained from a urine sample of a UTI patient in Turkey and identified via 16S rDNA sequencing. Antimicrobial susceptibility test was performed for 17 antibiotics using VITEK® 2 and the results were confirmed using minimum inhibitory concentration assay. Whole-genome sequencing of the isolate was performed using Illumina sequencing and analyzed with bioinformatic tools for multilocus sequence typing, replicon types, virulence factors, and antimicrobial resistance genes.
Results. Whole-genome datum was submitted to the National Center for Biotechnology Information (NCBI; accession No. JAKSGM000000000). The isolate was only found to be resistant to piperacillin in the β-lactam class of antibiotics. While the isolate was also resistant to aminoglycoside and tetracycline antibiotics, it was sensitive to other antibiotics tested. Ten antibiotic resistance genes were identified in the genome of the isolate: blaOXA-1, blaOXA-2, aac(6’)-II, aac(6’)-Ib-cr, tetB, catB3, qacE, sitABCD, mdfA, and sul-2. Clonal subtype (ST) and serotype of the isolate were identified as ST2141 and O107/H39, respectively. Plasmid replicon typing was used to identify 5 plasmid types in the genome of E. coli Rize-53 (Col(BS512), IncC, IncIA, IncFIB(AP1918), and IncFII(pRSB107)); however, none of the resistance genes were encoded on the plasmid.
Conclusions. Genetic determinants of resistance to tetracycline, β-lactam and aminoglycoside antibiotics were identified using WGS in a uropathogenic E. coli from ST2141 lineage and O107:H39 serotype, isolated in Turkey.
Key words: whole-genome sequencing, uropathogenic Escherichia coli ST2141, serotype O107:H39, tetracycline resistance (tetB), blaOXA-1/aac6
Background
Extensive antibiotic use to treat urinary tract infections (UTIs) has led to the widespread antibiotic resistance in the pathogens that cause these infections. The primary cause of many UTIs is Gram-negative Enterobacterales, such as uropathogenic Escherichia coli (UPEC) and Klebsiella pneumoniae spp.1 Spread of drug-resistant strains of E. coli is a serious health concern worldwide, as it leads to more persistent infections and higher mortality rates.2 Escherichia coli is an prevalent human pathogen that is responsible for up to 90% of all community-acquired UTIs.3
The production of β-lactamases by uropathogens has led to an increased β-lactam resistance, complicating UTI treatment.4 The β-lactam antibiotics inhibit peptidoglycan synthesis, though they are ineffective against Enterobacteriaceae that produce extended-spectrum β-lactamases (ESBLs). The resistance to β-lactams means carbapenems need to be used to effectively treat infections caused by ESBL-producing Enterobacteriaceae. The β-lactamases are categorized into 4 classes based on amino acid contents, namely class A, B, C, and D.5 Class D β-lactamases include oxacillinase (OXA)-type enzymes that hydrolyze cloxacillin and oxacillin faster than benzylpenicillin.6
Another class of antibiotics are tetracyclines (TEs), which inhibit protein synthesis in bacteria. The tet A–E classes are the most frequently detected TE resistance genes in the Enterobacterales, and they primarily confer resistance via TE efflux or ribosomal protection. Resistance to TE is frequently spread among E. coli isolates through horizontal transfer of genes such as tetA.7, 8 Consequently, TE resistance has led to a decreased use of these antibiotics in humans. However, TEs are still among the most widely used antibiotics in livestock production globally.9
Aminoglycoside antibiotics are used for the single-dose treatment of UTIs.10 They mainly act by disrupting bacterial protein synthesis through binding to prokaryotic ribosomes via 16S ribosomal RNA (16S rRNA) and disrupting bacterial cell membrane integrity.11 Aminoglycoside resistance occurs through several mechanisms; however, enzymatic inactivation is the most prevalent in the clinical setting and is carried out by acetyltransferases, nucleotidyltransferases and phosphotransferases, which perform acetylation, adenylation and phosphorylation, respectively. A large number of enzymes that modify aminoglycosides have been described to date, and they are present in nearly all bacteria that present enzymatic resistance to aminoglycosides.12 Understanding the antimicrobial resistance phenotypes of livestock, wildlife and human pathogens is important for identifying proper treatment plans and addressing the urgent situation of rising antimicrobial resistance.
Objectives
Whole-genome sequencing (WGS) is an important approach to revealing characteristics related to antimicrobial resistance genes in bacteria. This study aimed to perform a whole-genome analysis of one UTI isolate from Turkey in order to characterize resistance genes, multilocus sequence type (MLST) and plasmid profiles.
Materials and methods
Bacterial strain and antimicrobial susceptibility testing
One UTI isolate of E. coli was obtained from glycerol stock stored at Recep Tayyip Erdoğan University in Rize, Turkey, and was characterized using standard microbiological procedures and VITEK® 2 (bioMérieux, Marcy-l’Étoile, France). It was further identified using 16S rDNA Sanger dideoxy sequencing and named E. coli Rize-53 after the Basic Local Alignment Search Tool (BLAST) analysis on the National Center for Biotechnology Information (NCBI) website. Susceptibility testing was performed using the following antibiotics: piperacillin, piperacillin/tazobactam, ceftazidime, cefepime, aztreonam, imipenem, meropenem, amikacin, gentamicin, netilmicin, tobramycin, ciprofloxacin, levofloxacin, tetracycline, tigecycline, colistin, and trimethoprim/sulfamethoxazole, confirmed with E-test (bioMérieux) in accordance with Clinical and Laboratory Standards Institute guidelines.13
Whole-genome sequencing
Given its phenotypic resistance to antibiotics from different groups, such as β-lactams, aminoglycosides and tetracyclines, it was decided to perform WGS in order to identify the resistance gene profile. A bacterial sample was sent to Macrogen Europe (Amsterdam, The Netherlands) for preparation and library construction. The library was sequenced with Illumina sequencing using synthesis technology, and converted into raw data for analysis. Sequence reads were de novo assembled. After the whole-genome was assembled, the location of protein-coding sequences, transfer RNA (tRNA) genes, ribosomal RNA (rRNA) genes, and transfer-messenger RNA (tmRNA) genes were identified and their functions annotated. Prokka software (https://narrative.kbase.us/#catalog/apps/ProkkaAnnotation/annotate_contigs) was used to predict location, while BLAST was used to run the assembled sequences against a nucleotide and protein sequence database in order to identify and predict the function of the coding sequences.
Analysis of genome
The analysis was performed using the website of the Center for Genomic Epidemiology project (https://www.genomicepidemiology.org/). Replicase genes of the plasmids were classified using the PlasmidFinder v. 2.0 online tool (https://cge.food.dtu.dk/services/PlasmidFinder/). The MLST was identified using whole-genome sequences in the MLST database by applying bioinformatics tools available from the Center for Genomic Epidemiology. Serotype of E. coli Rize-53 was determined using SerotypeFinder (https://cge.food.dtu.dk/services/SerotypeFinder/) based on the O-(lipopolysaccharide) and H-(flagellar) antigen processing genes.14 Antimicrobial resistance encoding genes were identified using ResFinder (https://cge.food.dtu.dk/services/ResFinder/) and a cutoff of 80% similarity.
Results
According to the Minimum Inhibitory Concentration (MIC) assay results, only piperacillin resistance (≥64) was observed among the β-lactam antibiotics. Moderate resistance was observed in the piperacillin/tazobactam combination, with a decreased MIC value of 16. No resistance to carbapenems was observed. Out of the 4 different aminoglycoside antibiotics, sensitivity was only observed to broad-spectrum gentamicin, while resistance was observed to the other 3 aminoglycosides (Table 1). Of the other antibiotics tested, only tetracycline resistance was observed. A total of 10 resistance genes were identified in the WGS analysis. These were blaOXA-1 and blaOXA-2 (β-lactam resistance; piperacillin, piperacillin/tazobactam and cefepime), aac(6’)-II and aac(6’)-Ib-cr (aminoglycoside resistance), tetB (tetracycline resistance), catB3 (chloramphenicol resistance), qacE (quaternary ammonium compound-resistance protein E), sitABCD, mdfA (erythromycin and roxithromycin resistance), and sul2 (sulfonamide resistance).
The whole-genome sequence was 5,430,376 nucleotides in length and was submitted to GenBank (https://www.ncbi.nlm.nih.gov/genbank/; the accession No. JAKSGM000000000). It had coding sequence for a total of 5230 genes, 83 tRNAs, 9 rRNAs, and 1 tmRNA. Clonal subtype (ST) of E. coli Rize-53 was determined as ST2141 (adk101-fumC88-gyrB262-icd281-mdh59-purA215-recA196). It was then determined that E. coli 53-Rize had wzx and fliC alleles and its serotype was O107/H39. No virulence genes were detected.
A total of 5 plasmids were detected in the genome of E. coli Rize-53, which included Col(BS512), IncC, IncFIA, IncFIB(AP1918), and IncFII(pRSB107). While Col(BS512), IncC, IncFIA and IncFII(pRSB107) were 100% similar in replicon typing, IncFIB(AP1918) had nucleotide changes and was found to be 98.39% similar. None of the resistance genes detected with WGS were on these plasmids.
Discussion
Escherichia coli isolates are normally found in human flora, though certain strains are the most common cause of UTIs. Antimicrobial therapy is crucial in the treatment of UTIs; however, rising resistance to antimicrobials has made an effective treatment more complicated. Results from a comparative study showed that resistance to trimethoprim-sulfamethoxazole (TMP-SMZ), which is used as a first-line resistance in the treatment of uncomplicated UTIs, did not change significantly over time in UPEC isolates. However, more recently, the increasing resistance to TMP-SMZ has been observed15; thus, TMP-SMZ should not be used in empirical therapy.16 In the case of the E. coli Rize-53 isolate analyzed in the study, sensitivity to TMP-SMZ was demonstrated and no genes related to the resistance to this antibiotic were discovered.
Widespread and common use of quinolones and fluoroquinolones in the treatment of UTIs worldwide has led to the increased resistance in UPEC isolates.17 Moreover, the resistance and level of fluoroquinolones in UPEC isolates reported from different countries are important. In this study, ciprofloxacin and levofloxacin from fluoroquinolones were tested against E. coli Rize-53. Escherichia coli was found to be sensitive to these fluoroquinolones, and no genes related to the resistance to this antibiotic group were found. Additionally, no tigecycline or colistin resistance genes were found in the genome, which explained the sensitivity to these antibiotics.
Escherichia coli Rize-53 was sensitive to all β-lactams tested, with the exception of piperacillin, and was also sensitive to piperacillin/tazobactam. It was previously demonstrated that carbapenem, piperacillin-tazobactam and amikacin antibiotics were highly effective against E. coli isolates identified from UTIs in Canada and the USA between 2010 and 2014 (>95% sensitivity).18 The main cause of resistance to β-lactams is different types of β-lactamases. According to the WGS data, only 2 OXA genes (blaOXA-1/2) are encoded in the E. coli Rize-53 genome, which we predict are responsible for piperacillin resistance. Furthermore, WGS data indicated that it was likely the tetB gene was the cause of the observed resistance to tetracycline in E. coli Rize-53. In an earlier study conducted on urine samples of 121 adult patients from Turkey, the resistance to tetracycline was observed in 82 samples and tetB was shown to be the most abundant tetracycline resistance gene.1 Another study of 268 clinical E. coli isolates obtained from urine samples found that 36.8% were resistant to tetracycline.19 Genetic determinants responsible for tetracycline resistance were not investigated in that study, but it demonstrated that tetracycline resistance is high in UTIs in Turkey.
Escherichia coli Rize-53 is only sensitive to broad-spectrum gentamicin and is resistant to the aminoglycoside antibiotics such as amikacin, netilmicin and tobramycin. Consistent with this phenotype, it has 2 genes, aac(6’)-II and aac(6’)-Ib-cr, which are likely responsible for this resistance. Aminoglycoside-modifying enzyme genes can coexist in some strains, and combinations such as aph(3”)-Ib/aac(3)-Ia/aac(60)-Ib, ant(2”)-Ia/aac(6’)-Ib, aac(3)-Ia/aac(6’)-Ib, aac(6’)-Ib/aph(3”)-Ib and aac(6’)-Ib/aac(3)-IIa, aac(3)-IIa/aph(3’)-Ia, ant(2’)-Ia/aph(3’)-Ia, aac(6’)-Ib/aph(3’)-Ia have been identified in clinical isolates of E. coli.20, 21 Production of enzymes that alter aminoglycosides is one of the most common mechanisms of resistance to aminoglycosides in E. coli isolates. Furthermore, genes encoding aminoglycoside-modifying enzymes are mostly found on plasmids, but here they were identified in the chromosomal genome in E. coli Rize-53.
There may be a strong relationship between antibiotic resistance genes and ST in clonal groups. It is important to understand the geographical relationship of this grouping, as this will impact the direction and success of the therapies used. However, there is no detailed data on the STs of E. coli identified from Turkey in the literature. Indeed, no whole-genome sequence of E. coli from ST2141 type has been submitted to the pathogen watch portal (https://pathogen.watch). Therefore, this work represents the first WGS of ST2141 of E. coli from Turkey. Nonetheless, Escherichia coli ST2141 was isolated from the domestic duck (Anas platyrhynchos) in Bangladesh, though no genome analysis was performed.22 In the current study, E. coli ST2141 was isolated from the urine of a UTI patient. Knowing antimicrobial resistance phenotypes for livestock, wildlife species and humans is important in order to follow changes in antibiotic resistance. In this regard, zoonotic transmission of disease represents a significant threat to human health. However, ST2141 isolated from domestic duck was reported to be CTX-M15-positive, whilst this gene could not be detected in the genome of E. coli Rize-53, although β-lactamase encoding OXA-1 and -2 were detected.
Escherichia coli serotyping is based on O and H antigens. The serotype searching database was based on the O-antigen genes wzx, wzy, wzm, and wzt for in silico O typing (comprising more than 188 O antisera) and the flagellin genes fliC, flkA, flmA, flnA, and fllA for in silico H typing (comprising more than 53 H antisera).23 Determining the serotype via an in silico WGS approach is more reliable; therefore, this approach was used to identify the serotype of E. coli Rize-53 as O107/H39. Although E. coli very different E. coli serotypes based on O and H antigens have been shown in the literature, it has been determined that the first example of O107/H39 serotype is E. coli Rize-53. The O-antigen, an oligosaccharide with many repeats, is an essential component of the lipopolysaccharide on the surface of Gram-negative bacteria and one of the most variable cell constituents. The E. coli Rize-53 O107 O-antigen has a close relationship to E. coli O117 O-antigens, with only 1 substitution of D-GlcNAc for D-Glc. Furthermore, the O-antigen gene clusters of E. coli O107 and O117 share 98.6% overall DNA identity.24 Flagellin is the protein subunit of the flagellum, with an average molecular mass of approx. 50 kDa (from 36 to 60 kDa), that carries H-antigen specificity. There are currently 53 H types for E. coli flagella named from H1 to H56, and this established distinction between a highly variable central region and more conserved flanking regions was upheld.25, 26
The Col(BS512) plasmid of E. coli Rize-53 was similar to a plasmid of Shigella boydii (accession No. NC010656, 2089 bp in length, 100% identity) that does not contain any resistance genes. The IncC plasmid of E. coli Rize-53 was also similar to the plasmid pNDM-KN (accession No. JN157804, 162746 bp in length, 100% identity) of Klebsiella pneumoniae strain Kp7. However, pNDM-KN, unlike E. coli Rize-53 IncC, had genes encoding NDM and AmpC-type β-lactamases.27 These genes made the Kp7 strain resistant to carbapenems, while E. coli Rize-53 was carbapenem-susceptible. The IncFIA (100% identity) and IncFIB(AP1918) (98.38% identity) were similar to E. coli K-12 plasmid F (accession No. AP001918, 99159 bp in length) that did not contain any resistance genes.28 The IncFII(pRSB107) was similar to an uncultured bacterium plasmid pRSB107 (accession No. AJ851089, 120592 bp in length, 100% identity), and harbored several resistance genes such as aph3 (kanamycin), strA and strB (streptomycin), sulII (sulfonamide), blaTEM-1b (β-lactamase), mph(A) (erythromycin and roxithromycin), dhfR (trimethoprim), catA (chloramphenicol), tetA, tetC, and tetR (tetracycline). The fact that the resistance genes of the E. coli Rize-53 isolate are on its genome limits the transmission of these resistance genes to other strains. Limiting transmissibility of these resistance genes is important in terms of preventing their spread.
Limitations
The main limitation of this study was a sample size, as WGS-type studies should be carried out on a whole population and include a variety of bacteria with different resistance patterns. Such efforts require organization at the national level and these limitations will be the focus of subsequent studies.
Conclusions
Antibiotic therapy is important in UTI treatment and there is an increasing resistance of UTIs to routinely applied antibiotics. Identifying the resistance genes encoded in the genome of a pathogen is crucial to understanding the resistance profile and the mechanisms behind the action of these pathogens. In this study, E. coli Rize-53 was resistant to the β-lactam piperacillin, the aminoglycosides netilmicin, tobramycin, and amikacin, as well as tetracycline. Resistance genes were identified using WGS, which revealed that piperacillin resistance was due to the blaOXA1/2 genes, while aac(6’)-Il, aac(6’)-Ib-cr and tetB genes conferred resistance to the aminoglycosides and tetracycline, respectively. However, none of the resistance genes identified in the study were carried by any of the 5 plasmids present in the strain. The genome sequence of E. coli Rize-53 demonstrated that this clinical isolate is equipped with important antibiotic resistance genes. Therefore, WGS offers a unique approach to evaluating antibiotic resistance genes across the entire genome of pathogens. This will allow the emerging and serious public health threat posed by such pathogens in both developed and developing countries around the world to be addressed more readily.